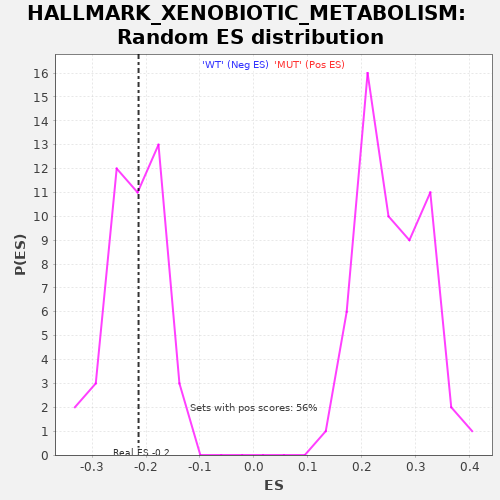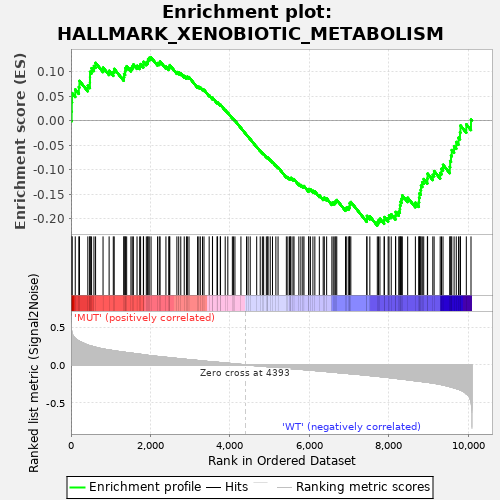
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_XENOBIOTIC_METABOLISM |
| Enrichment Score (ES) | -0.2138443 |
| Normalized Enrichment Score (NES) | -0.9528703 |
| Nominal p-value | 0.6136364 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ACP2 | na | 16 | 0.451 | 0.0184 | No | ||
| 2 | DHRS1 | na | 20 | 0.443 | 0.0378 | No | ||
| 3 | PTS | na | 27 | 0.424 | 0.0560 | No | ||
| 4 | PINK1 | na | 107 | 0.349 | 0.0636 | No | ||
| 5 | LPIN2 | na | 198 | 0.316 | 0.0685 | No | ||
| 6 | EPHA2 | na | 213 | 0.310 | 0.0809 | No | ||
| 7 | CYP2E1 | na | 424 | 0.260 | 0.0713 | No | ||
| 8 | CAT | na | 476 | 0.252 | 0.0774 | No | ||
| 9 | MAN1A1 | na | 478 | 0.252 | 0.0884 | No | ||
| 10 | HSD17B2 | na | 480 | 0.252 | 0.0995 | No | ||
| 11 | ETS2 | na | 518 | 0.247 | 0.1067 | No | ||
| 12 | PMM1 | na | 578 | 0.238 | 0.1114 | No | ||
| 13 | MCCC2 | na | 619 | 0.231 | 0.1176 | No | ||
| 14 | FMO3 | na | 807 | 0.209 | 0.1081 | No | ||
| 15 | CDA | na | 960 | 0.196 | 0.1015 | No | ||
| 16 | ADH1C | na | 1065 | 0.187 | 0.0993 | No | ||
| 17 | SLC1A5 | na | 1088 | 0.186 | 0.1054 | No | ||
| 18 | HPRT1 | na | 1329 | 0.166 | 0.0886 | No | ||
| 19 | DDC | na | 1338 | 0.166 | 0.0952 | No | ||
| 20 | SERPINA6 | na | 1363 | 0.165 | 0.1001 | No | ||
| 21 | JUP | na | 1368 | 0.164 | 0.1069 | No | ||
| 22 | ARG2 | na | 1402 | 0.162 | 0.1108 | No | ||
| 23 | BCAT1 | na | 1510 | 0.155 | 0.1069 | No | ||
| 24 | REG1A | na | 1542 | 0.152 | 0.1106 | No | ||
| 25 | PTGDS | na | 1571 | 0.150 | 0.1144 | No | ||
| 26 | F10 | na | 1663 | 0.145 | 0.1117 | No | ||
| 27 | ENTPD5 | na | 1730 | 0.139 | 0.1113 | No | ||
| 28 | IGF1 | na | 1753 | 0.138 | 0.1152 | No | ||
| 29 | CBR1 | na | 1823 | 0.134 | 0.1142 | No | ||
| 30 | ACOX2 | na | 1827 | 0.133 | 0.1198 | No | ||
| 31 | ACOX1 | na | 1897 | 0.128 | 0.1185 | No | ||
| 32 | ADH5 | na | 1931 | 0.125 | 0.1208 | No | ||
| 33 | PPARD | na | 1942 | 0.124 | 0.1253 | No | ||
| 34 | SPINT2 | na | 1970 | 0.122 | 0.1280 | No | ||
| 35 | MAOA | na | 2016 | 0.119 | 0.1287 | No | ||
| 36 | ETFDH | na | 2180 | 0.110 | 0.1172 | No | ||
| 37 | DDAH2 | na | 2222 | 0.108 | 0.1179 | No | ||
| 38 | SLC12A4 | na | 2244 | 0.107 | 0.1205 | No | ||
| 39 | MTHFD1 | na | 2391 | 0.098 | 0.1102 | No | ||
| 40 | RBP4 | na | 2462 | 0.094 | 0.1073 | No | ||
| 41 | ITIH1 | na | 2478 | 0.093 | 0.1100 | No | ||
| 42 | CYP4F2 | na | 2489 | 0.093 | 0.1131 | No | ||
| 43 | CASP6 | na | 2667 | 0.084 | 0.0990 | No | ||
| 44 | CA2 | na | 2713 | 0.081 | 0.0981 | No | ||
| 45 | GNMT | na | 2765 | 0.078 | 0.0964 | No | ||
| 46 | COMT | na | 2852 | 0.073 | 0.0910 | No | ||
| 47 | SLC6A6 | na | 2908 | 0.071 | 0.0886 | No | ||
| 48 | TGFB2 | na | 2931 | 0.069 | 0.0895 | No | ||
| 49 | PTGES | na | 2972 | 0.067 | 0.0885 | No | ||
| 50 | CYP2C18 | na | 3188 | 0.057 | 0.0694 | No | ||
| 51 | CYP1A1 | na | 3220 | 0.055 | 0.0687 | No | ||
| 52 | GART | na | 3257 | 0.054 | 0.0675 | No | ||
| 53 | ALAS1 | na | 3312 | 0.050 | 0.0643 | No | ||
| 54 | GSS | na | 3350 | 0.048 | 0.0627 | No | ||
| 55 | GSTT2 | na | 3480 | 0.042 | 0.0516 | No | ||
| 56 | SLC35B1 | na | 3563 | 0.038 | 0.0450 | No | ||
| 57 | SERPINE1 | na | 3565 | 0.038 | 0.0466 | No | ||
| 58 | PC | na | 3678 | 0.033 | 0.0368 | No | ||
| 59 | GCLC | na | 3691 | 0.033 | 0.0371 | No | ||
| 60 | CYP26A1 | na | 3760 | 0.029 | 0.0315 | No | ||
| 61 | FABP1 | na | 3766 | 0.029 | 0.0323 | No | ||
| 62 | IGFBP4 | na | 3884 | 0.024 | 0.0216 | No | ||
| 63 | TDO2 | na | 3950 | 0.021 | 0.0159 | No | ||
| 64 | KARS | na | 4064 | 0.014 | 0.0052 | No | ||
| 65 | MBL2 | na | 4091 | 0.013 | 0.0032 | No | ||
| 66 | NQO1 | na | 4129 | 0.011 | -0.0000 | No | ||
| 67 | PLG | na | 4280 | 0.005 | -0.0149 | No | ||
| 68 | IGFBP1 | na | 4421 | -0.002 | -0.0289 | No | ||
| 69 | G6PC | na | 4460 | -0.003 | -0.0326 | No | ||
| 70 | ENPEP | na | 4514 | -0.006 | -0.0377 | No | ||
| 71 | LCAT | na | 4677 | -0.013 | -0.0534 | No | ||
| 72 | PYCR1 | na | 4766 | -0.017 | -0.0615 | No | ||
| 73 | ABCD2 | na | 4820 | -0.020 | -0.0659 | No | ||
| 74 | ACOX3 | na | 4852 | -0.021 | -0.0681 | No | ||
| 75 | GCH1 | na | 4920 | -0.023 | -0.0739 | No | ||
| 76 | CROT | na | 4958 | -0.025 | -0.0765 | No | ||
| 77 | CDO1 | na | 4959 | -0.025 | -0.0754 | No | ||
| 78 | GABARAPL1 | na | 5018 | -0.027 | -0.0800 | No | ||
| 79 | ELOVL5 | na | 5075 | -0.029 | -0.0843 | No | ||
| 80 | ALDH9A1 | na | 5160 | -0.033 | -0.0913 | No | ||
| 81 | PGRMC1 | na | 5215 | -0.036 | -0.0951 | No | ||
| 82 | GCNT2 | na | 5424 | -0.045 | -0.1140 | No | ||
| 83 | PSMB10 | na | 5462 | -0.047 | -0.1156 | No | ||
| 84 | ACO2 | na | 5505 | -0.049 | -0.1177 | No | ||
| 85 | NMT1 | na | 5525 | -0.050 | -0.1174 | No | ||
| 86 | FBLN1 | na | 5540 | -0.050 | -0.1166 | No | ||
| 87 | ABCC3 | na | 5590 | -0.053 | -0.1192 | No | ||
| 88 | ALDH2 | na | 5617 | -0.054 | -0.1194 | No | ||
| 89 | GAD1 | na | 5739 | -0.059 | -0.1290 | No | ||
| 90 | PGD | na | 5790 | -0.061 | -0.1313 | No | ||
| 91 | ADH7 | na | 5838 | -0.063 | -0.1333 | No | ||
| 92 | ID2 | na | 5868 | -0.064 | -0.1333 | No | ||
| 93 | VTN | na | 5986 | -0.069 | -0.1420 | No | ||
| 94 | CRP | na | 5990 | -0.069 | -0.1393 | No | ||
| 95 | HSD11B1 | na | 6030 | -0.071 | -0.1400 | No | ||
| 96 | EPHX1 | na | 6093 | -0.074 | -0.1430 | No | ||
| 97 | XDH | na | 6142 | -0.076 | -0.1444 | No | ||
| 98 | ALDH3A1 | na | 6255 | -0.082 | -0.1521 | No | ||
| 99 | AQP9 | na | 6352 | -0.086 | -0.1579 | No | ||
| 100 | TNFRSF1A | na | 6380 | -0.087 | -0.1568 | No | ||
| 101 | CYP2J2 | na | 6439 | -0.090 | -0.1586 | No | ||
| 102 | ASL | na | 6571 | -0.096 | -0.1675 | No | ||
| 103 | PAPSS2 | na | 6605 | -0.098 | -0.1665 | No | ||
| 104 | ACP1 | na | 6645 | -0.100 | -0.1660 | No | ||
| 105 | SLC35D1 | na | 6668 | -0.101 | -0.1637 | No | ||
| 106 | BLVRB | na | 6694 | -0.102 | -0.1617 | No | ||
| 107 | ECH1 | na | 6914 | -0.112 | -0.1787 | No | ||
| 108 | DHRS7 | na | 6945 | -0.114 | -0.1767 | No | ||
| 109 | TPST1 | na | 6995 | -0.116 | -0.1765 | No | ||
| 110 | ATP2A2 | na | 7018 | -0.117 | -0.1735 | No | ||
| 111 | IDH1 | na | 7019 | -0.117 | -0.1683 | No | ||
| 112 | FETUB | na | 7048 | -0.119 | -0.1658 | No | ||
| 113 | PROS1 | na | 7450 | -0.138 | -0.2000 | No | ||
| 114 | FAH | na | 7453 | -0.138 | -0.1940 | No | ||
| 115 | FBP1 | na | 7527 | -0.143 | -0.1950 | No | ||
| 116 | SHMT2 | na | 7715 | -0.152 | -0.2071 | Yes | ||
| 117 | AHCY | na | 7744 | -0.154 | -0.2031 | Yes | ||
| 118 | SMOX | na | 7781 | -0.156 | -0.1998 | Yes | ||
| 119 | F11 | na | 7887 | -0.162 | -0.2031 | Yes | ||
| 120 | PEMT | na | 7893 | -0.162 | -0.1964 | Yes | ||
| 121 | HMOX1 | na | 7988 | -0.168 | -0.1984 | Yes | ||
| 122 | DDT | na | 8010 | -0.169 | -0.1930 | Yes | ||
| 123 | GCKR | na | 8066 | -0.173 | -0.1908 | Yes | ||
| 124 | HGFAC | na | 8174 | -0.179 | -0.1936 | Yes | ||
| 125 | IL1R1 | na | 8178 | -0.179 | -0.1860 | Yes | ||
| 126 | DHPS | na | 8256 | -0.184 | -0.1856 | Yes | ||
| 127 | BPHL | na | 8283 | -0.185 | -0.1799 | Yes | ||
| 128 | PDK4 | na | 8288 | -0.186 | -0.1721 | Yes | ||
| 129 | CES1 | na | 8302 | -0.187 | -0.1651 | Yes | ||
| 130 | HRG | na | 8327 | -0.188 | -0.1591 | Yes | ||
| 131 | VNN1 | na | 8344 | -0.190 | -0.1523 | Yes | ||
| 132 | UGDH | na | 8480 | -0.198 | -0.1571 | Yes | ||
| 133 | ITIH4 | na | 8674 | -0.212 | -0.1671 | Yes | ||
| 134 | GSR | na | 8761 | -0.217 | -0.1661 | Yes | ||
| 135 | CYP1A2 | na | 8768 | -0.218 | -0.1570 | Yes | ||
| 136 | FMO1 | na | 8777 | -0.218 | -0.1481 | Yes | ||
| 137 | SLC22A1 | na | 8808 | -0.220 | -0.1414 | Yes | ||
| 138 | MPP2 | na | 8816 | -0.221 | -0.1323 | Yes | ||
| 139 | POR | na | 8848 | -0.223 | -0.1255 | Yes | ||
| 140 | AKR1C3 | na | 8884 | -0.226 | -0.1190 | Yes | ||
| 141 | NPC1 | na | 8976 | -0.232 | -0.1179 | Yes | ||
| 142 | GSTO1 | na | 8982 | -0.232 | -0.1080 | Yes | ||
| 143 | CYP17A1 | na | 9113 | -0.242 | -0.1104 | Yes | ||
| 144 | TTPA | na | 9146 | -0.244 | -0.1027 | Yes | ||
| 145 | HNF4A | na | 9299 | -0.259 | -0.1065 | Yes | ||
| 146 | APOE | na | 9332 | -0.263 | -0.0981 | Yes | ||
| 147 | ESR1 | na | 9370 | -0.267 | -0.0899 | Yes | ||
| 148 | ARG1 | na | 9541 | -0.289 | -0.0942 | Yes | ||
| 149 | CYFIP2 | na | 9550 | -0.290 | -0.0821 | Yes | ||
| 150 | TYR | na | 9572 | -0.293 | -0.0712 | Yes | ||
| 151 | CCL25 | na | 9591 | -0.296 | -0.0599 | Yes | ||
| 152 | NINJ1 | na | 9649 | -0.306 | -0.0520 | Yes | ||
| 153 | KYNU | na | 9704 | -0.315 | -0.0435 | Yes | ||
| 154 | CYP27A1 | na | 9761 | -0.324 | -0.0347 | Yes | ||
| 155 | SLC6A12 | na | 9798 | -0.331 | -0.0236 | Yes | ||
| 156 | UPP1 | na | 9810 | -0.334 | -0.0099 | Yes | ||
| 157 | TAT | na | 9957 | -0.385 | -0.0075 | Yes | ||
| 158 | FAS | na | 10075 | -0.489 | 0.0024 | Yes |