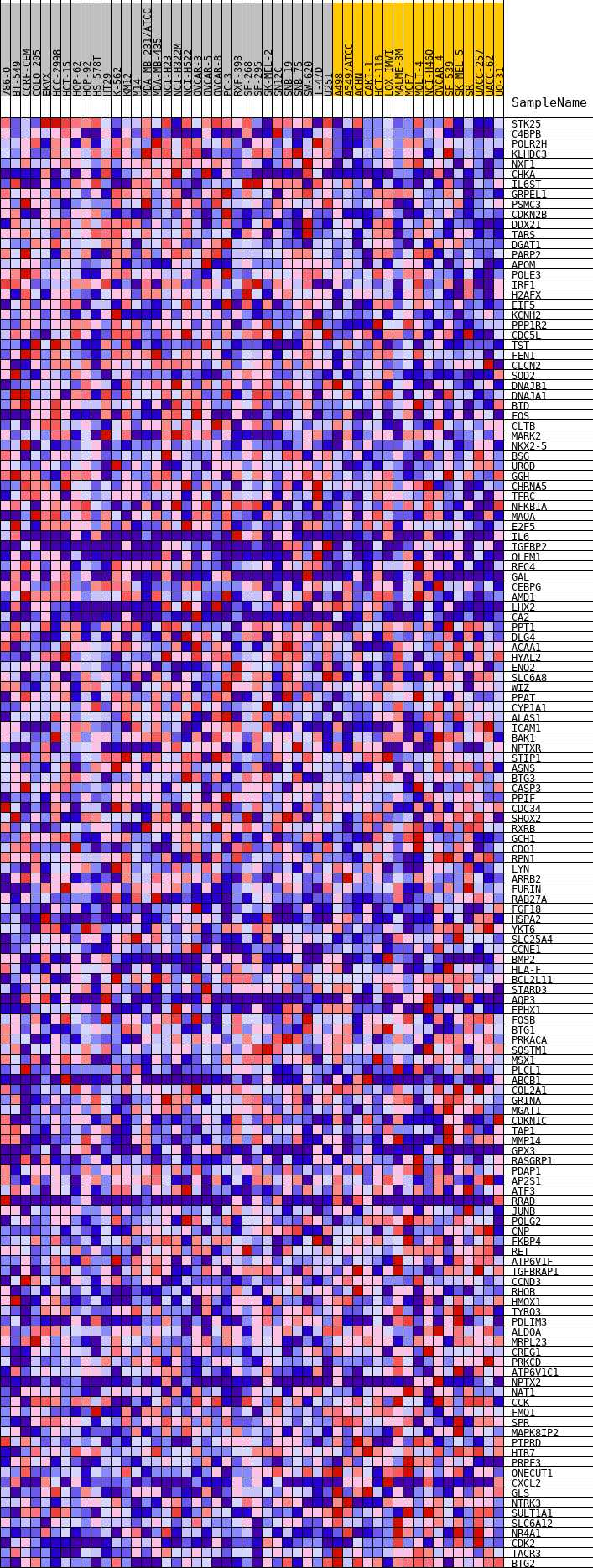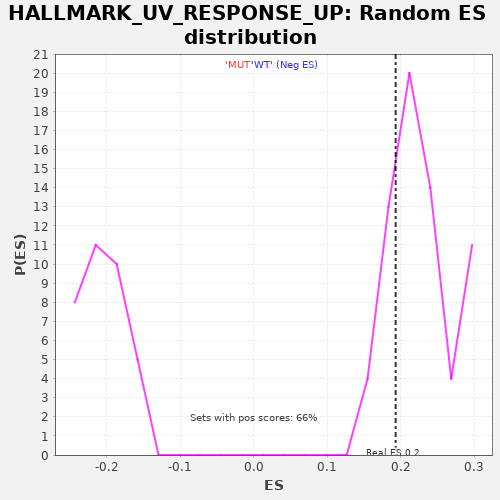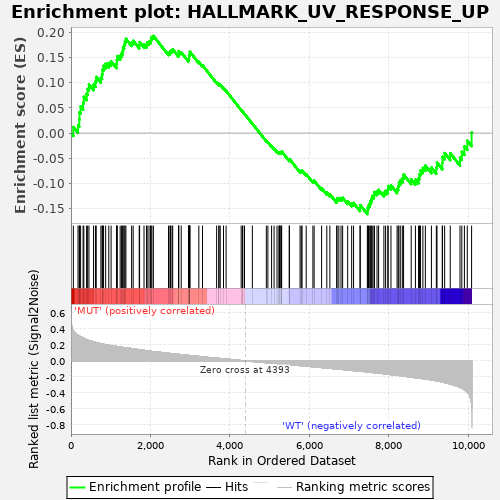
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_UV_RESPONSE_UP |
| Enrichment Score (ES) | 0.1929226 |
| Normalized Enrichment Score (NES) | 0.8495004 |
| Nominal p-value | 0.77272725 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | STK25 | na | 59 | 0.378 | 0.0120 | Yes | ||
| 2 | C4BPB | na | 177 | 0.322 | 0.0154 | Yes | ||
| 3 | POLR2H | na | 208 | 0.312 | 0.0272 | Yes | ||
| 4 | KLHDC3 | na | 214 | 0.310 | 0.0413 | Yes | ||
| 5 | NXF1 | na | 242 | 0.301 | 0.0529 | Yes | ||
| 6 | CHKA | na | 305 | 0.288 | 0.0603 | Yes | ||
| 7 | IL6ST | na | 323 | 0.284 | 0.0721 | Yes | ||
| 8 | GRPEL1 | na | 393 | 0.266 | 0.0777 | Yes | ||
| 9 | PSMC3 | na | 419 | 0.260 | 0.0876 | Yes | ||
| 10 | CDKN2B | na | 452 | 0.256 | 0.0964 | Yes | ||
| 11 | DDX21 | na | 568 | 0.239 | 0.0962 | Yes | ||
| 12 | TARS | na | 616 | 0.232 | 0.1025 | Yes | ||
| 13 | DGAT1 | na | 635 | 0.230 | 0.1115 | Yes | ||
| 14 | PARP2 | na | 753 | 0.215 | 0.1100 | Yes | ||
| 15 | APOM | na | 783 | 0.211 | 0.1171 | Yes | ||
| 16 | POLE3 | na | 794 | 0.210 | 0.1260 | Yes | ||
| 17 | IRF1 | na | 820 | 0.208 | 0.1333 | Yes | ||
| 18 | H2AFX | na | 873 | 0.203 | 0.1377 | Yes | ||
| 19 | EIF5 | na | 952 | 0.197 | 0.1392 | Yes | ||
| 20 | KCNH2 | na | 1010 | 0.192 | 0.1425 | Yes | ||
| 21 | PPP1R2 | na | 1146 | 0.180 | 0.1375 | Yes | ||
| 22 | CDC5L | na | 1155 | 0.179 | 0.1452 | Yes | ||
| 23 | TST | na | 1165 | 0.178 | 0.1528 | Yes | ||
| 24 | FEN1 | na | 1238 | 0.174 | 0.1537 | Yes | ||
| 25 | CLCN2 | na | 1274 | 0.171 | 0.1583 | Yes | ||
| 26 | SOD2 | na | 1301 | 0.168 | 0.1637 | Yes | ||
| 27 | DNAJB1 | na | 1312 | 0.168 | 0.1706 | Yes | ||
| 28 | DNAJA1 | na | 1339 | 0.166 | 0.1758 | Yes | ||
| 29 | BID | na | 1359 | 0.165 | 0.1817 | Yes | ||
| 30 | FOS | na | 1383 | 0.163 | 0.1871 | Yes | ||
| 31 | CLTB | na | 1527 | 0.153 | 0.1800 | Yes | ||
| 32 | MARK2 | na | 1566 | 0.151 | 0.1834 | Yes | ||
| 33 | NKX2-5 | na | 1718 | 0.140 | 0.1748 | Yes | ||
| 34 | BSG | na | 1728 | 0.140 | 0.1805 | Yes | ||
| 35 | UROD | na | 1839 | 0.133 | 0.1758 | Yes | ||
| 36 | GGH | na | 1902 | 0.128 | 0.1756 | Yes | ||
| 37 | CHRNA5 | na | 1918 | 0.127 | 0.1801 | Yes | ||
| 38 | TFRC | na | 1964 | 0.122 | 0.1813 | Yes | ||
| 39 | NFKBIA | na | 2002 | 0.120 | 0.1833 | Yes | ||
| 40 | MAOA | na | 2016 | 0.119 | 0.1876 | Yes | ||
| 41 | E2F5 | na | 2037 | 0.118 | 0.1912 | Yes | ||
| 42 | IL6 | na | 2075 | 0.115 | 0.1929 | Yes | ||
| 43 | IGFBP2 | na | 2456 | 0.095 | 0.1592 | No | ||
| 44 | OLFM1 | na | 2492 | 0.093 | 0.1601 | No | ||
| 45 | RFC4 | na | 2506 | 0.092 | 0.1632 | No | ||
| 46 | GAL | na | 2544 | 0.090 | 0.1637 | No | ||
| 47 | CEBPG | na | 2561 | 0.089 | 0.1663 | No | ||
| 48 | AMD1 | na | 2708 | 0.081 | 0.1555 | No | ||
| 49 | LHX2 | na | 2712 | 0.081 | 0.1590 | No | ||
| 50 | CA2 | na | 2713 | 0.081 | 0.1629 | No | ||
| 51 | PPT1 | na | 2776 | 0.077 | 0.1603 | No | ||
| 52 | DLG4 | na | 2959 | 0.068 | 0.1452 | No | ||
| 53 | ACAA1 | na | 2968 | 0.068 | 0.1476 | No | ||
| 54 | HYAL2 | na | 2974 | 0.067 | 0.1503 | No | ||
| 55 | ENO2 | na | 2975 | 0.067 | 0.1535 | No | ||
| 56 | SLC6A8 | na | 2981 | 0.067 | 0.1562 | No | ||
| 57 | WIZ | na | 2993 | 0.066 | 0.1582 | No | ||
| 58 | PPAT | na | 2994 | 0.066 | 0.1614 | No | ||
| 59 | CYP1A1 | na | 3220 | 0.055 | 0.1414 | No | ||
| 60 | ALAS1 | na | 3312 | 0.050 | 0.1346 | No | ||
| 61 | ICAM1 | na | 3668 | 0.034 | 0.1005 | No | ||
| 62 | BAK1 | na | 3725 | 0.031 | 0.0964 | No | ||
| 63 | NPTXR | na | 3726 | 0.031 | 0.0979 | No | ||
| 64 | STIP1 | na | 3758 | 0.029 | 0.0961 | No | ||
| 65 | ASNS | na | 3841 | 0.026 | 0.0891 | No | ||
| 66 | BTG3 | na | 3906 | 0.023 | 0.0837 | No | ||
| 67 | CASP3 | na | 4286 | 0.004 | 0.0459 | No | ||
| 68 | PPIF | na | 4322 | 0.003 | 0.0425 | No | ||
| 69 | CDC34 | na | 4326 | 0.003 | 0.0424 | No | ||
| 70 | SHOX2 | na | 4370 | 0.001 | 0.0381 | No | ||
| 71 | RXRB | na | 4568 | -0.008 | 0.0187 | No | ||
| 72 | GCH1 | na | 4920 | -0.023 | -0.0155 | No | ||
| 73 | CDO1 | na | 4959 | -0.025 | -0.0181 | No | ||
| 74 | RPN1 | na | 5052 | -0.028 | -0.0260 | No | ||
| 75 | LYN | na | 5115 | -0.031 | -0.0307 | No | ||
| 76 | ARRB2 | na | 5189 | -0.035 | -0.0364 | No | ||
| 77 | FURIN | na | 5236 | -0.037 | -0.0393 | No | ||
| 78 | RAB27A | na | 5252 | -0.038 | -0.0390 | No | ||
| 79 | FGF18 | na | 5264 | -0.038 | -0.0383 | No | ||
| 80 | HSPA2 | na | 5290 | -0.039 | -0.0390 | No | ||
| 81 | YKT6 | na | 5296 | -0.039 | -0.0376 | No | ||
| 82 | SLC25A4 | na | 5306 | -0.040 | -0.0367 | No | ||
| 83 | CCNE1 | na | 5497 | -0.049 | -0.0534 | No | ||
| 84 | BMP2 | na | 5501 | -0.049 | -0.0514 | No | ||
| 85 | HLA-F | na | 5769 | -0.060 | -0.0754 | No | ||
| 86 | BCL2L11 | na | 5800 | -0.061 | -0.0755 | No | ||
| 87 | STARD3 | na | 5820 | -0.062 | -0.0745 | No | ||
| 88 | AQP3 | na | 5924 | -0.066 | -0.0817 | No | ||
| 89 | EPHX1 | na | 6093 | -0.074 | -0.0951 | No | ||
| 90 | FOSB | na | 6124 | -0.075 | -0.0946 | No | ||
| 91 | BTG1 | na | 6312 | -0.084 | -0.1094 | No | ||
| 92 | PRKACA | na | 6443 | -0.090 | -0.1181 | No | ||
| 93 | SQSTM1 | na | 6520 | -0.094 | -0.1214 | No | ||
| 94 | MSX1 | na | 6692 | -0.102 | -0.1337 | No | ||
| 95 | PLCL1 | na | 6701 | -0.103 | -0.1297 | No | ||
| 96 | ABCB1 | na | 6746 | -0.105 | -0.1291 | No | ||
| 97 | COL2A1 | na | 6801 | -0.107 | -0.1295 | No | ||
| 98 | GRINA | na | 6842 | -0.109 | -0.1283 | No | ||
| 99 | MGAT1 | na | 6969 | -0.115 | -0.1356 | No | ||
| 100 | CDKN1C | na | 7069 | -0.120 | -0.1398 | No | ||
| 101 | TAP1 | na | 7117 | -0.122 | -0.1387 | No | ||
| 102 | MMP14 | na | 7280 | -0.130 | -0.1489 | No | ||
| 103 | GPX3 | na | 7285 | -0.130 | -0.1431 | No | ||
| 104 | RASGRP1 | na | 7464 | -0.139 | -0.1544 | No | ||
| 105 | PDAP1 | na | 7475 | -0.139 | -0.1488 | No | ||
| 106 | AP2S1 | na | 7493 | -0.141 | -0.1439 | No | ||
| 107 | ATF3 | na | 7525 | -0.142 | -0.1402 | No | ||
| 108 | RRAD | na | 7541 | -0.143 | -0.1350 | No | ||
| 109 | JUNB | na | 7572 | -0.145 | -0.1311 | No | ||
| 110 | POLG2 | na | 7587 | -0.146 | -0.1256 | No | ||
| 111 | CNP | na | 7630 | -0.148 | -0.1228 | No | ||
| 112 | FKBP4 | na | 7643 | -0.149 | -0.1169 | No | ||
| 113 | RET | na | 7708 | -0.152 | -0.1162 | No | ||
| 114 | ATP6V1F | na | 7748 | -0.154 | -0.1128 | No | ||
| 115 | TGFBRAP1 | na | 7881 | -0.162 | -0.1184 | No | ||
| 116 | CCND3 | na | 7924 | -0.164 | -0.1148 | No | ||
| 117 | RHOB | na | 7981 | -0.168 | -0.1125 | No | ||
| 118 | HMOX1 | na | 7988 | -0.168 | -0.1052 | No | ||
| 119 | TYRO3 | na | 8059 | -0.173 | -0.1040 | No | ||
| 120 | PDLIM3 | na | 8213 | -0.182 | -0.1108 | No | ||
| 121 | ALDOA | na | 8243 | -0.184 | -0.1050 | No | ||
| 122 | MRPL23 | na | 8261 | -0.184 | -0.0980 | No | ||
| 123 | CREG1 | na | 8305 | -0.187 | -0.0934 | No | ||
| 124 | PRKCD | na | 8354 | -0.190 | -0.0892 | No | ||
| 125 | ATP6V1C1 | na | 8377 | -0.192 | -0.0824 | No | ||
| 126 | NPTX2 | na | 8569 | -0.205 | -0.0918 | No | ||
| 127 | NAT1 | na | 8675 | -0.212 | -0.0923 | No | ||
| 128 | CCK | na | 8754 | -0.217 | -0.0899 | No | ||
| 129 | FMO1 | na | 8777 | -0.218 | -0.0818 | No | ||
| 130 | SPR | na | 8804 | -0.220 | -0.0740 | No | ||
| 131 | MAPK8IP2 | na | 8863 | -0.224 | -0.0692 | No | ||
| 132 | PTPRD | na | 8925 | -0.228 | -0.0645 | No | ||
| 133 | HTR7 | na | 9078 | -0.240 | -0.0684 | No | ||
| 134 | PRPF3 | na | 9202 | -0.250 | -0.0689 | No | ||
| 135 | ONECUT1 | na | 9218 | -0.251 | -0.0586 | No | ||
| 136 | CXCL2 | na | 9352 | -0.264 | -0.0594 | No | ||
| 137 | GLS | na | 9355 | -0.265 | -0.0471 | No | ||
| 138 | NTRK3 | na | 9411 | -0.272 | -0.0397 | No | ||
| 139 | SULT1A1 | na | 9552 | -0.290 | -0.0400 | No | ||
| 140 | SLC6A12 | na | 9798 | -0.331 | -0.0490 | No | ||
| 141 | NR4A1 | na | 9844 | -0.342 | -0.0373 | No | ||
| 142 | CDK2 | na | 9909 | -0.362 | -0.0266 | No | ||
| 143 | TACR3 | na | 9981 | -0.393 | -0.0151 | No | ||
| 144 | BTG2 | na | 10090 | -0.567 | 0.0009 | No |