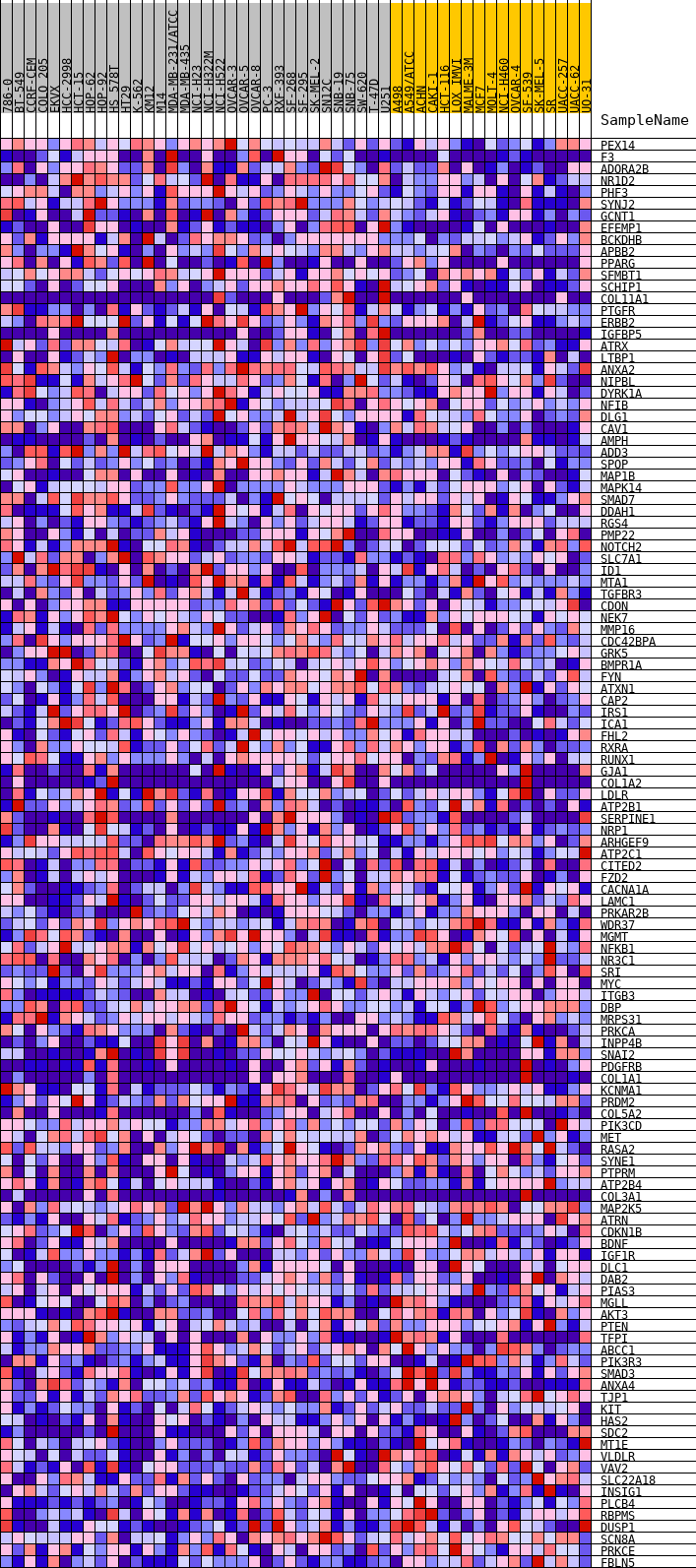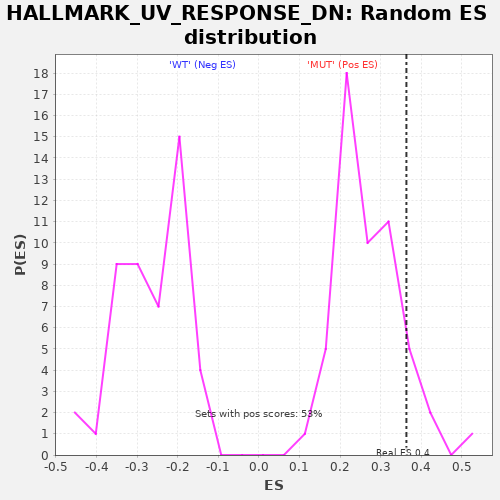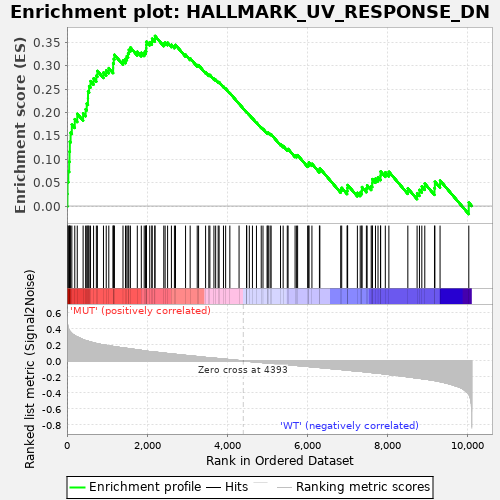
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_UV_RESPONSE_DN |
| Enrichment Score (ES) | 0.3634873 |
| Normalized Enrichment Score (NES) | 1.3415395 |
| Nominal p-value | 0.094339624 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.89 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PEX14 | na | 11 | 0.457 | 0.0258 | Yes | ||
| 2 | F3 | na | 14 | 0.454 | 0.0524 | Yes | ||
| 3 | ADORA2B | na | 33 | 0.409 | 0.0746 | Yes | ||
| 4 | NR1D2 | na | 55 | 0.379 | 0.0949 | Yes | ||
| 5 | PHF3 | na | 63 | 0.376 | 0.1163 | Yes | ||
| 6 | SYNJ2 | na | 71 | 0.371 | 0.1374 | Yes | ||
| 7 | GCNT1 | na | 90 | 0.360 | 0.1568 | Yes | ||
| 8 | EFEMP1 | na | 121 | 0.343 | 0.1740 | Yes | ||
| 9 | BCKDHB | na | 194 | 0.317 | 0.1855 | Yes | ||
| 10 | APBB2 | na | 256 | 0.299 | 0.1970 | Yes | ||
| 11 | PPARG | na | 402 | 0.264 | 0.1980 | Yes | ||
| 12 | SFMBT1 | na | 465 | 0.254 | 0.2068 | Yes | ||
| 13 | SCHIP1 | na | 493 | 0.250 | 0.2188 | Yes | ||
| 14 | COL11A1 | na | 523 | 0.246 | 0.2304 | Yes | ||
| 15 | PTGFR | na | 525 | 0.246 | 0.2447 | Yes | ||
| 16 | ERBB2 | na | 552 | 0.241 | 0.2563 | Yes | ||
| 17 | IGFBP5 | na | 587 | 0.237 | 0.2669 | Yes | ||
| 18 | ATRX | na | 663 | 0.225 | 0.2726 | Yes | ||
| 19 | LTBP1 | na | 732 | 0.217 | 0.2786 | Yes | ||
| 20 | ANXA2 | na | 758 | 0.214 | 0.2887 | Yes | ||
| 21 | NIPBL | na | 910 | 0.200 | 0.2853 | Yes | ||
| 22 | DYRK1A | na | 979 | 0.194 | 0.2899 | Yes | ||
| 23 | NFIB | na | 1043 | 0.189 | 0.2947 | Yes | ||
| 24 | DLG1 | na | 1151 | 0.180 | 0.2946 | Yes | ||
| 25 | CAV1 | na | 1152 | 0.180 | 0.3052 | Yes | ||
| 26 | AMPH | na | 1167 | 0.178 | 0.3143 | Yes | ||
| 27 | ADD3 | na | 1183 | 0.177 | 0.3232 | Yes | ||
| 28 | SPOP | na | 1397 | 0.162 | 0.3114 | Yes | ||
| 29 | MAP1B | na | 1460 | 0.158 | 0.3145 | Yes | ||
| 30 | MAPK14 | na | 1494 | 0.156 | 0.3204 | Yes | ||
| 31 | SMAD7 | na | 1524 | 0.153 | 0.3266 | Yes | ||
| 32 | DDAH1 | na | 1543 | 0.152 | 0.3337 | Yes | ||
| 33 | RGS4 | na | 1582 | 0.150 | 0.3388 | Yes | ||
| 34 | PMP22 | na | 1754 | 0.138 | 0.3297 | Yes | ||
| 35 | NOTCH2 | na | 1854 | 0.132 | 0.3276 | Yes | ||
| 36 | SLC7A1 | na | 1926 | 0.126 | 0.3279 | Yes | ||
| 37 | ID1 | na | 1961 | 0.122 | 0.3317 | Yes | ||
| 38 | MTA1 | na | 1976 | 0.122 | 0.3374 | Yes | ||
| 39 | TGFBR3 | na | 1978 | 0.122 | 0.3445 | Yes | ||
| 40 | CDON | na | 1983 | 0.121 | 0.3512 | Yes | ||
| 41 | NEK7 | na | 2064 | 0.116 | 0.3500 | Yes | ||
| 42 | MMP16 | na | 2118 | 0.113 | 0.3514 | Yes | ||
| 43 | CDC42BPA | na | 2123 | 0.113 | 0.3576 | Yes | ||
| 44 | GRK5 | na | 2191 | 0.109 | 0.3574 | Yes | ||
| 45 | BMPR1A | na | 2195 | 0.109 | 0.3635 | Yes | ||
| 46 | FYN | na | 2415 | 0.097 | 0.3473 | No | ||
| 47 | ATXN1 | na | 2448 | 0.095 | 0.3497 | No | ||
| 48 | CAP2 | na | 2510 | 0.092 | 0.3490 | No | ||
| 49 | IRS1 | na | 2605 | 0.087 | 0.3447 | No | ||
| 50 | ICA1 | na | 2680 | 0.083 | 0.3422 | No | ||
| 51 | FHL2 | na | 2707 | 0.081 | 0.3443 | No | ||
| 52 | RXRA | na | 2958 | 0.068 | 0.3233 | No | ||
| 53 | RUNX1 | na | 3073 | 0.063 | 0.3156 | No | ||
| 54 | GJA1 | na | 3249 | 0.054 | 0.3012 | No | ||
| 55 | COL1A2 | na | 3283 | 0.052 | 0.3010 | No | ||
| 56 | LDLR | na | 3458 | 0.043 | 0.2860 | No | ||
| 57 | ATP2B1 | na | 3539 | 0.039 | 0.2803 | No | ||
| 58 | SERPINE1 | na | 3565 | 0.038 | 0.2801 | No | ||
| 59 | NRP1 | na | 3666 | 0.034 | 0.2720 | No | ||
| 60 | ARHGEF9 | na | 3709 | 0.032 | 0.2697 | No | ||
| 61 | ATP2C1 | na | 3772 | 0.029 | 0.2652 | No | ||
| 62 | CITED2 | na | 3795 | 0.028 | 0.2646 | No | ||
| 63 | FZD2 | na | 3904 | 0.023 | 0.2551 | No | ||
| 64 | CACNA1A | na | 3958 | 0.020 | 0.2510 | No | ||
| 65 | LAMC1 | na | 4063 | 0.014 | 0.2414 | No | ||
| 66 | PRKAR2B | na | 4293 | 0.004 | 0.2187 | No | ||
| 67 | WDR37 | na | 4479 | -0.004 | 0.2004 | No | ||
| 68 | MGMT | na | 4481 | -0.004 | 0.2005 | No | ||
| 69 | NFKB1 | na | 4483 | -0.004 | 0.2006 | No | ||
| 70 | NR3C1 | na | 4549 | -0.007 | 0.1946 | No | ||
| 71 | SRI | na | 4627 | -0.011 | 0.1875 | No | ||
| 72 | MYC | na | 4727 | -0.015 | 0.1785 | No | ||
| 73 | ITGB3 | na | 4846 | -0.021 | 0.1678 | No | ||
| 74 | DBP | na | 4890 | -0.022 | 0.1648 | No | ||
| 75 | MRPS31 | na | 4992 | -0.026 | 0.1563 | No | ||
| 76 | PRKCA | na | 5013 | -0.027 | 0.1558 | No | ||
| 77 | INPP4B | na | 5019 | -0.027 | 0.1569 | No | ||
| 78 | SNAI2 | na | 5067 | -0.029 | 0.1539 | No | ||
| 79 | PDGFRB | na | 5093 | -0.030 | 0.1532 | No | ||
| 80 | COL1A1 | na | 5327 | -0.041 | 0.1322 | No | ||
| 81 | KCNMA1 | na | 5393 | -0.044 | 0.1283 | No | ||
| 82 | PRDM2 | na | 5496 | -0.049 | 0.1210 | No | ||
| 83 | COL5A2 | na | 5516 | -0.049 | 0.1220 | No | ||
| 84 | PIK3CD | na | 5691 | -0.057 | 0.1079 | No | ||
| 85 | MET | na | 5729 | -0.058 | 0.1076 | No | ||
| 86 | RASA2 | na | 5754 | -0.059 | 0.1087 | No | ||
| 87 | SYNE1 | na | 6004 | -0.070 | 0.0879 | No | ||
| 88 | PTPRM | na | 6032 | -0.072 | 0.0894 | No | ||
| 89 | ATP2B4 | na | 6036 | -0.072 | 0.0933 | No | ||
| 90 | COL3A1 | na | 6111 | -0.075 | 0.0903 | No | ||
| 91 | MAP2K5 | na | 6298 | -0.084 | 0.0766 | No | ||
| 92 | ATRN | na | 6309 | -0.084 | 0.0805 | No | ||
| 93 | CDKN1B | na | 6829 | -0.108 | 0.0349 | No | ||
| 94 | BDNF | na | 6854 | -0.109 | 0.0389 | No | ||
| 95 | IGF1R | na | 6989 | -0.116 | 0.0323 | No | ||
| 96 | DLC1 | na | 6997 | -0.117 | 0.0385 | No | ||
| 97 | DAB2 | na | 7002 | -0.117 | 0.0449 | No | ||
| 98 | PIAS3 | na | 7245 | -0.128 | 0.0282 | No | ||
| 99 | MGLL | na | 7316 | -0.132 | 0.0289 | No | ||
| 100 | AKT3 | na | 7356 | -0.133 | 0.0329 | No | ||
| 101 | PTEN | na | 7359 | -0.133 | 0.0405 | No | ||
| 102 | TFPI | na | 7470 | -0.139 | 0.0377 | No | ||
| 103 | ABCC1 | na | 7488 | -0.140 | 0.0443 | No | ||
| 104 | PIK3R3 | na | 7590 | -0.146 | 0.0428 | No | ||
| 105 | SMAD3 | na | 7616 | -0.147 | 0.0489 | No | ||
| 106 | ANXA4 | na | 7618 | -0.147 | 0.0575 | No | ||
| 107 | TJP1 | na | 7697 | -0.151 | 0.0586 | No | ||
| 108 | KIT | na | 7760 | -0.155 | 0.0615 | No | ||
| 109 | HAS2 | na | 7823 | -0.158 | 0.0646 | No | ||
| 110 | SDC2 | na | 7825 | -0.158 | 0.0738 | No | ||
| 111 | MT1E | na | 7942 | -0.165 | 0.0720 | No | ||
| 112 | VLDLR | na | 8031 | -0.171 | 0.0732 | No | ||
| 113 | VAV2 | na | 8504 | -0.200 | 0.0377 | No | ||
| 114 | SLC22A18 | na | 8735 | -0.216 | 0.0274 | No | ||
| 115 | INSIG1 | na | 8794 | -0.220 | 0.0345 | No | ||
| 116 | PLCB4 | na | 8853 | -0.223 | 0.0418 | No | ||
| 117 | RBPMS | na | 8927 | -0.228 | 0.0480 | No | ||
| 118 | DUSP1 | na | 9169 | -0.246 | 0.0383 | No | ||
| 119 | SCN8A | na | 9179 | -0.248 | 0.0520 | No | ||
| 120 | PRKCE | na | 9308 | -0.260 | 0.0545 | No | ||
| 121 | FBLN5 | na | 10025 | -0.419 | 0.0074 | No |