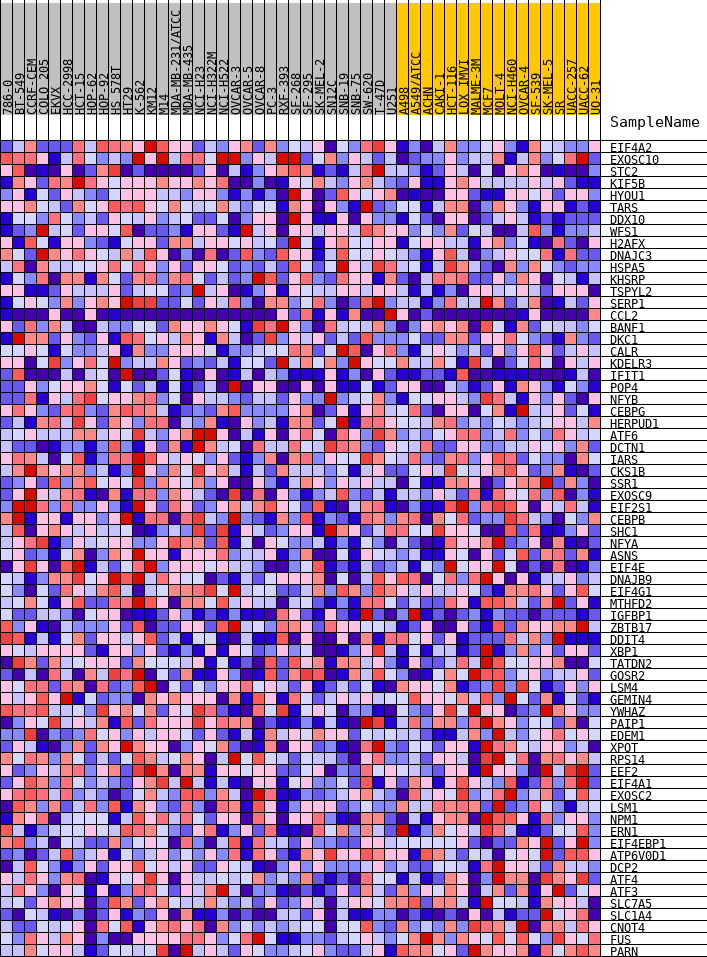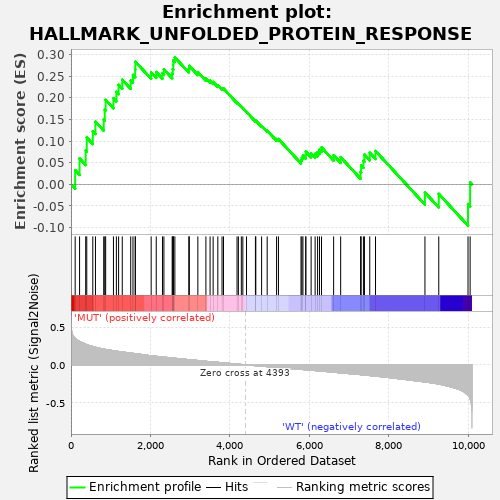
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_UNFOLDED_PROTEIN_RESPONSE |
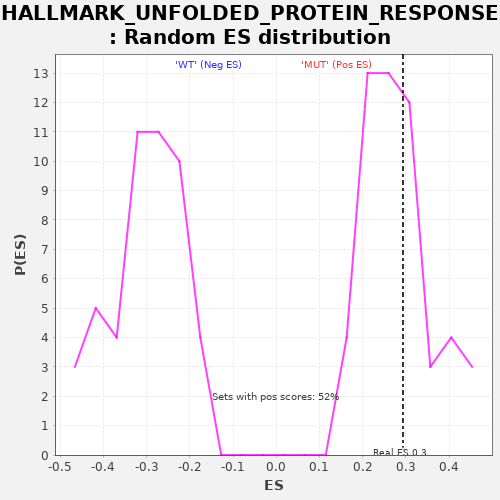
| Enrichment Score (ES) | 0.29334632 |
| Normalized Enrichment Score (NES) | 1.0381806 |
| Nominal p-value | 0.42307693 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EIF4A2 | na | 105 | 0.350 | 0.0326 | Yes | ||
| 2 | EXOSC10 | na | 216 | 0.309 | 0.0597 | Yes | ||
| 3 | STC2 | na | 370 | 0.272 | 0.0780 | Yes | ||
| 4 | KIF5B | na | 397 | 0.265 | 0.1081 | Yes | ||
| 5 | HYOU1 | na | 549 | 0.242 | 0.1228 | Yes | ||
| 6 | TARS | na | 616 | 0.232 | 0.1448 | Yes | ||
| 7 | DDX10 | na | 824 | 0.207 | 0.1497 | Yes | ||
| 8 | WFS1 | na | 850 | 0.205 | 0.1725 | Yes | ||
| 9 | H2AFX | na | 873 | 0.203 | 0.1953 | Yes | ||
| 10 | DNAJC3 | na | 1069 | 0.187 | 0.1989 | Yes | ||
| 11 | HSPA5 | na | 1143 | 0.181 | 0.2139 | Yes | ||
| 12 | KHSRP | na | 1199 | 0.176 | 0.2301 | Yes | ||
| 13 | TSPYL2 | na | 1291 | 0.169 | 0.2419 | Yes | ||
| 14 | SERP1 | na | 1504 | 0.155 | 0.2399 | Yes | ||
| 15 | CCL2 | na | 1561 | 0.151 | 0.2529 | Yes | ||
| 16 | BANF1 | na | 1617 | 0.148 | 0.2656 | Yes | ||
| 17 | DKC1 | na | 1621 | 0.147 | 0.2834 | Yes | ||
| 18 | CALR | na | 2019 | 0.119 | 0.2585 | Yes | ||
| 19 | KDELR3 | na | 2147 | 0.112 | 0.2596 | Yes | ||
| 20 | IFIT1 | na | 2304 | 0.104 | 0.2568 | Yes | ||
| 21 | POP4 | na | 2340 | 0.101 | 0.2658 | Yes | ||
| 22 | NFYB | na | 2548 | 0.090 | 0.2563 | Yes | ||
| 23 | CEBPG | na | 2561 | 0.089 | 0.2661 | Yes | ||
| 24 | HERPUD1 | na | 2576 | 0.088 | 0.2755 | Yes | ||
| 25 | ATF6 | na | 2577 | 0.088 | 0.2864 | Yes | ||
| 26 | DCTN1 | na | 2616 | 0.087 | 0.2933 | Yes | ||
| 27 | IARS | na | 2967 | 0.068 | 0.2668 | No | ||
| 28 | CKS1B | na | 2980 | 0.067 | 0.2739 | No | ||
| 29 | SSR1 | na | 3195 | 0.057 | 0.2595 | No | ||
| 30 | EXOSC9 | na | 3398 | 0.046 | 0.2450 | No | ||
| 31 | EIF2S1 | na | 3503 | 0.041 | 0.2397 | No | ||
| 32 | CEBPB | na | 3580 | 0.037 | 0.2367 | No | ||
| 33 | SHC1 | na | 3698 | 0.032 | 0.2290 | No | ||
| 34 | NFYA | na | 3804 | 0.027 | 0.2219 | No | ||
| 35 | ASNS | na | 3841 | 0.026 | 0.2214 | No | ||
| 36 | EIF4E | na | 4182 | 0.009 | 0.1887 | No | ||
| 37 | DNAJB9 | na | 4216 | 0.008 | 0.1864 | No | ||
| 38 | EIF4G1 | na | 4290 | 0.004 | 0.1796 | No | ||
| 39 | MTHFD2 | na | 4328 | 0.003 | 0.1763 | No | ||
| 40 | IGFBP1 | na | 4421 | -0.002 | 0.1673 | No | ||
| 41 | ZBTB17 | na | 4645 | -0.012 | 0.1465 | No | ||
| 42 | DDIT4 | na | 4655 | -0.012 | 0.1471 | No | ||
| 43 | XBP1 | na | 4801 | -0.019 | 0.1350 | No | ||
| 44 | TATDN2 | na | 4941 | -0.024 | 0.1241 | No | ||
| 45 | GOSR2 | na | 5178 | -0.034 | 0.1048 | No | ||
| 46 | LSM4 | na | 5225 | -0.036 | 0.1047 | No | ||
| 47 | GEMIN4 | na | 5793 | -0.061 | 0.0557 | No | ||
| 48 | YWHAZ | na | 5815 | -0.062 | 0.0612 | No | ||
| 49 | PAIP1 | na | 5842 | -0.063 | 0.0664 | No | ||
| 50 | EDEM1 | na | 5912 | -0.066 | 0.0676 | No | ||
| 51 | XPOT | na | 5915 | -0.066 | 0.0755 | No | ||
| 52 | RPS14 | na | 6048 | -0.072 | 0.0713 | No | ||
| 53 | EEF2 | na | 6149 | -0.076 | 0.0707 | No | ||
| 54 | EIF4A1 | na | 6211 | -0.080 | 0.0744 | No | ||
| 55 | EXOSC2 | na | 6259 | -0.082 | 0.0799 | No | ||
| 56 | LSM1 | na | 6310 | -0.084 | 0.0852 | No | ||
| 57 | NPM1 | na | 6615 | -0.099 | 0.0671 | No | ||
| 58 | ERN1 | na | 6792 | -0.107 | 0.0627 | No | ||
| 59 | EIF4EBP1 | na | 7294 | -0.130 | 0.0288 | No | ||
| 60 | ATP6V0D1 | na | 7308 | -0.131 | 0.0437 | No | ||
| 61 | DCP2 | na | 7365 | -0.134 | 0.0545 | No | ||
| 62 | ATF4 | na | 7391 | -0.135 | 0.0687 | No | ||
| 63 | ATF3 | na | 7525 | -0.142 | 0.0730 | No | ||
| 64 | SLC7A5 | na | 7670 | -0.150 | 0.0771 | No | ||
| 65 | SLC1A4 | na | 8915 | -0.227 | -0.0189 | No | ||
| 66 | CNOT4 | na | 9263 | -0.255 | -0.0221 | No | ||
| 67 | FUS | na | 10000 | -0.402 | -0.0460 | No | ||
| 68 | PARN | na | 10053 | -0.452 | 0.0046 | No |