Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_TNFA_SIGNALING_VIA_NFKB |
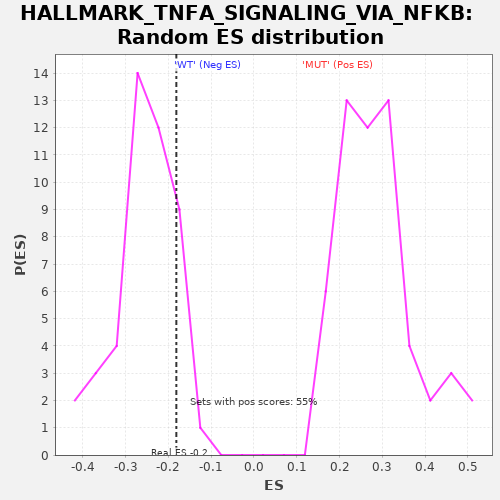
| Enrichment Score (ES) | -0.1814952 |
| Normalized Enrichment Score (NES) | -0.7000643 |
| Nominal p-value | 0.8666667 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | F3 | na | 14 | 0.454 | 0.0187 | Yes | ||
| 2 | RELA | na | 195 | 0.317 | 0.0146 | Yes | ||
| 3 | IL6ST | na | 323 | 0.284 | 0.0144 | Yes | ||
| 4 | BIRC2 | na | 335 | 0.281 | 0.0256 | Yes | ||
| 5 | MAFF | na | 358 | 0.274 | 0.0355 | Yes | ||
| 6 | OLR1 | na | 366 | 0.272 | 0.0468 | Yes | ||
| 7 | RELB | na | 399 | 0.264 | 0.0553 | Yes | ||
| 8 | IL7R | na | 433 | 0.259 | 0.0634 | Yes | ||
| 9 | MARCKS | na | 440 | 0.258 | 0.0742 | Yes | ||
| 10 | KLF4 | na | 509 | 0.247 | 0.0783 | Yes | ||
| 11 | ETS2 | na | 518 | 0.247 | 0.0884 | Yes | ||
| 12 | TIPARP | na | 535 | 0.244 | 0.0975 | Yes | ||
| 13 | NFKB2 | na | 656 | 0.226 | 0.0955 | Yes | ||
| 14 | BCL6 | na | 804 | 0.209 | 0.0899 | Yes | ||
| 15 | IRF1 | na | 820 | 0.208 | 0.0976 | Yes | ||
| 16 | CCRL2 | na | 853 | 0.204 | 0.1034 | Yes | ||
| 17 | IFNGR2 | na | 883 | 0.202 | 0.1094 | Yes | ||
| 18 | IL18 | na | 1100 | 0.185 | 0.0958 | Yes | ||
| 19 | CXCL11 | na | 1294 | 0.169 | 0.0839 | Yes | ||
| 20 | SOD2 | na | 1301 | 0.168 | 0.0907 | Yes | ||
| 21 | IFIT2 | na | 1308 | 0.168 | 0.0975 | Yes | ||
| 22 | IL15RA | na | 1365 | 0.164 | 0.0992 | Yes | ||
| 23 | GEM | na | 1370 | 0.164 | 0.1060 | Yes | ||
| 24 | FOS | na | 1383 | 0.163 | 0.1120 | Yes | ||
| 25 | HES1 | na | 1413 | 0.161 | 0.1162 | Yes | ||
| 26 | DNAJB4 | na | 1461 | 0.158 | 0.1185 | Yes | ||
| 27 | CCL2 | na | 1561 | 0.151 | 0.1152 | Yes | ||
| 28 | GADD45A | na | 1592 | 0.149 | 0.1188 | Yes | ||
| 29 | SDC4 | na | 1619 | 0.147 | 0.1226 | Yes | ||
| 30 | IER2 | na | 1655 | 0.145 | 0.1255 | Yes | ||
| 31 | TNC | na | 1695 | 0.142 | 0.1279 | Yes | ||
| 32 | GADD45B | na | 1763 | 0.137 | 0.1272 | Yes | ||
| 33 | PPP1R15A | na | 1825 | 0.134 | 0.1270 | Yes | ||
| 34 | TRAF1 | na | 1833 | 0.133 | 0.1322 | Yes | ||
| 35 | PTGER4 | na | 1856 | 0.131 | 0.1357 | Yes | ||
| 36 | FOSL1 | na | 1876 | 0.129 | 0.1395 | Yes | ||
| 37 | TANK | na | 1941 | 0.124 | 0.1386 | Yes | ||
| 38 | EHD1 | na | 1946 | 0.124 | 0.1436 | Yes | ||
| 39 | BIRC3 | na | 1955 | 0.123 | 0.1482 | Yes | ||
| 40 | PHLDA2 | na | 1973 | 0.122 | 0.1519 | Yes | ||
| 41 | NFKBIA | na | 2002 | 0.120 | 0.1544 | Yes | ||
| 42 | EFNA1 | na | 2034 | 0.118 | 0.1564 | Yes | ||
| 43 | EGR2 | na | 2057 | 0.116 | 0.1594 | Yes | ||
| 44 | IL6 | na | 2075 | 0.115 | 0.1628 | Yes | ||
| 45 | CSF2 | na | 2097 | 0.114 | 0.1657 | Yes | ||
| 46 | SNN | na | 2185 | 0.110 | 0.1618 | Yes | ||
| 47 | PTX3 | na | 2187 | 0.110 | 0.1665 | Yes | ||
| 48 | CCL5 | na | 2213 | 0.109 | 0.1688 | Yes | ||
| 49 | LAMB3 | na | 2219 | 0.108 | 0.1731 | Yes | ||
| 50 | PNRC1 | na | 2253 | 0.106 | 0.1744 | Yes | ||
| 51 | B4GALT5 | na | 2369 | 0.100 | 0.1673 | No | ||
| 52 | FUT4 | na | 2389 | 0.098 | 0.1697 | No | ||
| 53 | TLR2 | na | 2679 | 0.083 | 0.1443 | No | ||
| 54 | PANX1 | na | 2832 | 0.075 | 0.1323 | No | ||
| 55 | TNFAIP6 | na | 2842 | 0.074 | 0.1347 | No | ||
| 56 | NFAT5 | na | 3028 | 0.065 | 0.1189 | No | ||
| 57 | CXCL10 | na | 3211 | 0.056 | 0.1031 | No | ||
| 58 | CXCL3 | na | 3288 | 0.052 | 0.0978 | No | ||
| 59 | TRIP10 | na | 3357 | 0.048 | 0.0930 | No | ||
| 60 | CD44 | na | 3423 | 0.045 | 0.0885 | No | ||
| 61 | B4GALT1 | na | 3427 | 0.044 | 0.0901 | No | ||
| 62 | ZFP36 | na | 3446 | 0.043 | 0.0902 | No | ||
| 63 | LDLR | na | 3458 | 0.043 | 0.0910 | No | ||
| 64 | F2RL1 | na | 3497 | 0.041 | 0.0890 | No | ||
| 65 | ATP2B1 | na | 3539 | 0.039 | 0.0866 | No | ||
| 66 | SERPINE1 | na | 3565 | 0.038 | 0.0858 | No | ||
| 67 | CEBPB | na | 3580 | 0.037 | 0.0860 | No | ||
| 68 | CCND1 | na | 3593 | 0.037 | 0.0864 | No | ||
| 69 | ICAM1 | na | 3668 | 0.034 | 0.0805 | No | ||
| 70 | TNFRSF9 | na | 3702 | 0.032 | 0.0786 | No | ||
| 71 | AREG | na | 3773 | 0.029 | 0.0728 | No | ||
| 72 | TNIP1 | na | 3885 | 0.023 | 0.0627 | No | ||
| 73 | BTG3 | na | 3906 | 0.023 | 0.0617 | No | ||
| 74 | DUSP2 | na | 4024 | 0.017 | 0.0506 | No | ||
| 75 | JAG1 | na | 4027 | 0.017 | 0.0512 | No | ||
| 76 | IL12B | na | 4062 | 0.015 | 0.0484 | No | ||
| 77 | MAP3K8 | na | 4114 | 0.012 | 0.0438 | No | ||
| 78 | NFKB1 | na | 4483 | -0.004 | 0.0070 | No | ||
| 79 | SERPINB8 | na | 4542 | -0.007 | 0.0015 | No | ||
| 80 | MYC | na | 4727 | -0.015 | -0.0164 | No | ||
| 81 | JUN | na | 4816 | -0.019 | -0.0244 | No | ||
| 82 | GCH1 | na | 4920 | -0.023 | -0.0337 | No | ||
| 83 | TNFAIP8 | na | 5043 | -0.028 | -0.0447 | No | ||
| 84 | NFKBIE | na | 5081 | -0.030 | -0.0471 | No | ||
| 85 | MAP2K3 | na | 5101 | -0.031 | -0.0477 | No | ||
| 86 | PLAUR | na | 5152 | -0.033 | -0.0513 | No | ||
| 87 | CEBPD | na | 5187 | -0.035 | -0.0532 | No | ||
| 88 | FOSL2 | na | 5366 | -0.043 | -0.0692 | No | ||
| 89 | NR4A2 | na | 5404 | -0.044 | -0.0709 | No | ||
| 90 | CSF1 | na | 5473 | -0.047 | -0.0757 | No | ||
| 91 | EGR1 | na | 5478 | -0.048 | -0.0740 | No | ||
| 92 | BMP2 | na | 5501 | -0.049 | -0.0740 | No | ||
| 93 | CD69 | na | 5555 | -0.051 | -0.0771 | No | ||
| 94 | PLAU | na | 5583 | -0.052 | -0.0775 | No | ||
| 95 | IER3 | na | 5592 | -0.053 | -0.0760 | No | ||
| 96 | ID2 | na | 5868 | -0.064 | -0.1008 | No | ||
| 97 | REL | na | 5873 | -0.064 | -0.0984 | No | ||
| 98 | LITAF | na | 5961 | -0.068 | -0.1041 | No | ||
| 99 | SERPINB2 | na | 6040 | -0.072 | -0.1088 | No | ||
| 100 | CD80 | na | 6099 | -0.074 | -0.1114 | No | ||
| 101 | GFPT2 | na | 6106 | -0.074 | -0.1087 | No | ||
| 102 | FOSB | na | 6124 | -0.075 | -0.1071 | No | ||
| 103 | DUSP5 | na | 6201 | -0.079 | -0.1112 | No | ||
| 104 | BTG1 | na | 6312 | -0.084 | -0.1186 | No | ||
| 105 | PTPRE | na | 6404 | -0.088 | -0.1238 | No | ||
| 106 | SQSTM1 | na | 6520 | -0.094 | -0.1312 | No | ||
| 107 | BCL3 | na | 6654 | -0.100 | -0.1402 | No | ||
| 108 | G0S2 | na | 6755 | -0.105 | -0.1456 | No | ||
| 109 | DUSP4 | na | 6816 | -0.108 | -0.1469 | No | ||
| 110 | STAT5A | na | 6840 | -0.109 | -0.1444 | No | ||
| 111 | CD83 | na | 6967 | -0.115 | -0.1520 | No | ||
| 112 | NFIL3 | na | 6973 | -0.115 | -0.1474 | No | ||
| 113 | MSC | na | 7023 | -0.118 | -0.1472 | No | ||
| 114 | LIF | na | 7032 | -0.118 | -0.1427 | No | ||
| 115 | PHLDA1 | na | 7045 | -0.119 | -0.1387 | No | ||
| 116 | TAP1 | na | 7117 | -0.122 | -0.1404 | No | ||
| 117 | TNFAIP2 | na | 7183 | -0.125 | -0.1414 | No | ||
| 118 | PDE4B | na | 7347 | -0.133 | -0.1520 | No | ||
| 119 | SLC2A3 | na | 7357 | -0.133 | -0.1470 | No | ||
| 120 | PLEK | na | 7425 | -0.137 | -0.1477 | No | ||
| 121 | PFKFB3 | na | 7433 | -0.137 | -0.1423 | No | ||
| 122 | CFLAR | na | 7509 | -0.142 | -0.1436 | No | ||
| 123 | TRIB1 | na | 7516 | -0.142 | -0.1379 | No | ||
| 124 | ATF3 | na | 7525 | -0.142 | -0.1325 | No | ||
| 125 | INHBA | na | 7533 | -0.143 | -0.1269 | No | ||
| 126 | NFE2L2 | na | 7539 | -0.143 | -0.1210 | No | ||
| 127 | JUNB | na | 7572 | -0.145 | -0.1178 | No | ||
| 128 | SMAD3 | na | 7616 | -0.147 | -0.1157 | No | ||
| 129 | MCL1 | na | 7762 | -0.155 | -0.1234 | No | ||
| 130 | RHOB | na | 7981 | -0.168 | -0.1379 | No | ||
| 131 | BCL2A1 | na | 8042 | -0.172 | -0.1363 | No | ||
| 132 | RIPK2 | na | 8138 | -0.177 | -0.1381 | No | ||
| 133 | PLK2 | na | 8214 | -0.182 | -0.1376 | No | ||
| 134 | CCL20 | na | 8488 | -0.199 | -0.1562 | No | ||
| 135 | SLC16A6 | na | 8641 | -0.209 | -0.1623 | No | ||
| 136 | TNFAIP3 | na | 8660 | -0.211 | -0.1548 | No | ||
| 137 | IL1B | na | 8703 | -0.214 | -0.1495 | No | ||
| 138 | EGR3 | na | 8733 | -0.216 | -0.1429 | No | ||
| 139 | PTGS2 | na | 8737 | -0.216 | -0.1337 | No | ||
| 140 | DUSP1 | na | 9169 | -0.246 | -0.1662 | No | ||
| 141 | IFIH1 | na | 9176 | -0.247 | -0.1559 | No | ||
| 142 | CXCL2 | na | 9352 | -0.264 | -0.1618 | No | ||
| 143 | IRS2 | na | 9549 | -0.290 | -0.1687 | No | ||
| 144 | CXCL1 | na | 9567 | -0.292 | -0.1575 | No | ||
| 145 | CXCL6 | na | 9619 | -0.301 | -0.1493 | No | ||
| 146 | NINJ1 | na | 9649 | -0.306 | -0.1387 | No | ||
| 147 | NR4A3 | na | 9689 | -0.312 | -0.1288 | No | ||
| 148 | KYNU | na | 9704 | -0.315 | -0.1163 | No | ||
| 149 | ABCA1 | na | 9752 | -0.322 | -0.1068 | No | ||
| 150 | SOCS3 | na | 9795 | -0.330 | -0.0965 | No | ||
| 151 | TNF | na | 9835 | -0.340 | -0.0854 | No | ||
| 152 | NR4A1 | na | 9844 | -0.342 | -0.0711 | No | ||
| 153 | TNFSF9 | na | 9930 | -0.372 | -0.0632 | No | ||
| 154 | IL1A | na | 9993 | -0.400 | -0.0518 | No | ||
| 155 | BTG2 | na | 10090 | -0.567 | -0.0364 | No | ||
| 156 | CDKN1A | na | 10099 | -0.843 | -0.0000 | No |