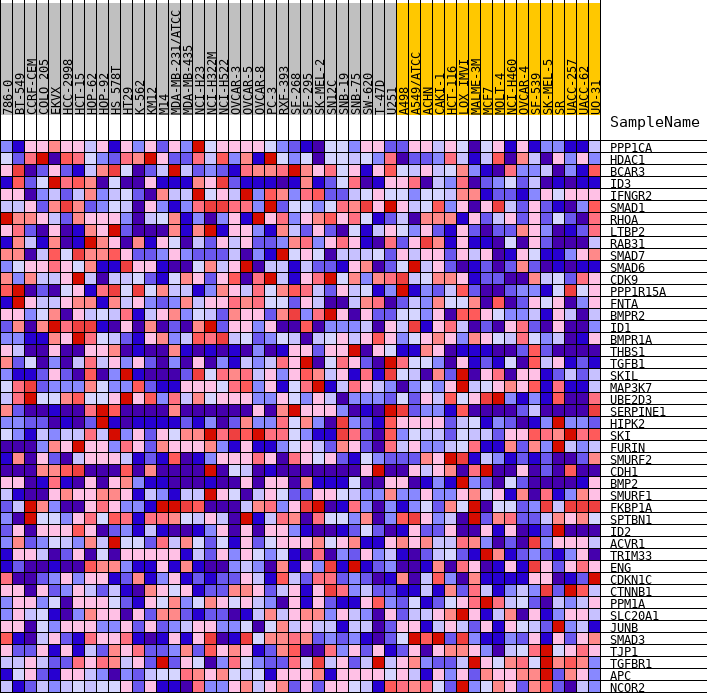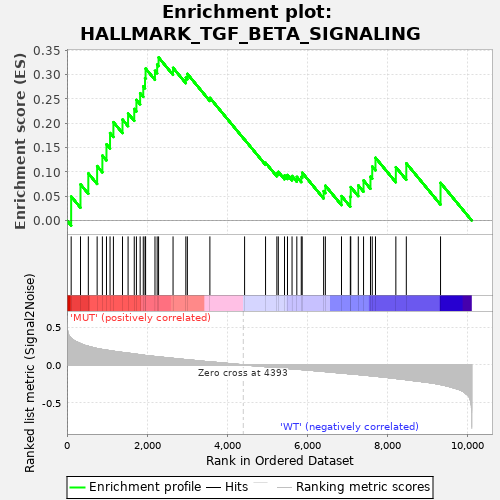
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
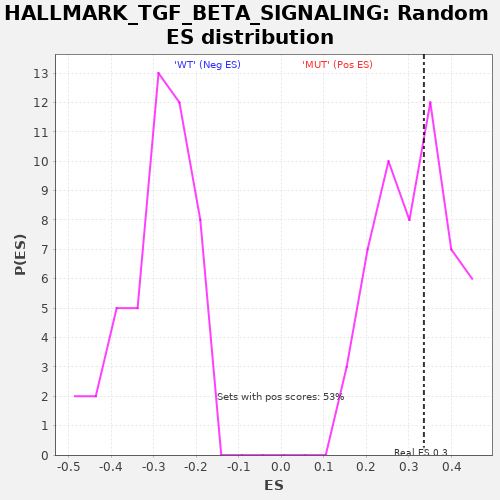
| GeneSet | HALLMARK_TGF_BETA_SIGNALING |
| Enrichment Score (ES) | 0.3350988 |
| Normalized Enrichment Score (NES) | 1.0798762 |
| Nominal p-value | 0.3773585 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PPP1CA | na | 104 | 0.350 | 0.0495 | Yes | ||
| 2 | HDAC1 | na | 337 | 0.280 | 0.0744 | Yes | ||
| 3 | BCAR3 | na | 532 | 0.245 | 0.0969 | Yes | ||
| 4 | ID3 | na | 752 | 0.215 | 0.1119 | Yes | ||
| 5 | IFNGR2 | na | 883 | 0.202 | 0.1336 | Yes | ||
| 6 | SMAD1 | na | 985 | 0.193 | 0.1566 | Yes | ||
| 7 | RHOA | na | 1074 | 0.187 | 0.1798 | Yes | ||
| 8 | LTBP2 | na | 1158 | 0.179 | 0.2022 | Yes | ||
| 9 | RAB31 | na | 1385 | 0.163 | 0.2075 | Yes | ||
| 10 | SMAD7 | na | 1524 | 0.153 | 0.2201 | Yes | ||
| 11 | SMAD6 | na | 1678 | 0.143 | 0.2293 | Yes | ||
| 12 | CDK9 | na | 1732 | 0.139 | 0.2479 | Yes | ||
| 13 | PPP1R15A | na | 1825 | 0.134 | 0.2616 | Yes | ||
| 14 | FNTA | na | 1903 | 0.128 | 0.2758 | Yes | ||
| 15 | BMPR2 | na | 1947 | 0.123 | 0.2926 | Yes | ||
| 16 | ID1 | na | 1961 | 0.122 | 0.3123 | Yes | ||
| 17 | BMPR1A | na | 2195 | 0.109 | 0.3078 | Yes | ||
| 18 | THBS1 | na | 2250 | 0.106 | 0.3206 | Yes | ||
| 19 | TGFB1 | na | 2285 | 0.105 | 0.3351 | Yes | ||
| 20 | SKIL | na | 2646 | 0.085 | 0.3138 | No | ||
| 21 | MAP3K7 | na | 2965 | 0.068 | 0.2938 | No | ||
| 22 | UBE2D3 | na | 3003 | 0.066 | 0.3013 | No | ||
| 23 | SERPINE1 | na | 3565 | 0.038 | 0.2521 | No | ||
| 24 | HIPK2 | na | 4431 | -0.002 | 0.1663 | No | ||
| 25 | SKI | na | 4954 | -0.025 | 0.1187 | No | ||
| 26 | FURIN | na | 5236 | -0.037 | 0.0971 | No | ||
| 27 | SMURF2 | na | 5272 | -0.038 | 0.1001 | No | ||
| 28 | CDH1 | na | 5427 | -0.045 | 0.0926 | No | ||
| 29 | BMP2 | na | 5501 | -0.049 | 0.0936 | No | ||
| 30 | SMURF1 | na | 5615 | -0.054 | 0.0916 | No | ||
| 31 | FKBP1A | na | 5734 | -0.058 | 0.0899 | No | ||
| 32 | SPTBN1 | na | 5845 | -0.063 | 0.0897 | No | ||
| 33 | ID2 | na | 5868 | -0.064 | 0.0984 | No | ||
| 34 | ACVR1 | na | 6403 | -0.088 | 0.0605 | No | ||
| 35 | TRIM33 | na | 6446 | -0.090 | 0.0717 | No | ||
| 36 | ENG | na | 6849 | -0.109 | 0.0503 | No | ||
| 37 | CDKN1C | na | 7069 | -0.120 | 0.0491 | No | ||
| 38 | CTNNB1 | na | 7080 | -0.121 | 0.0688 | No | ||
| 39 | PPM1A | na | 7268 | -0.129 | 0.0723 | No | ||
| 40 | SLC20A1 | na | 7399 | -0.135 | 0.0825 | No | ||
| 41 | JUNB | na | 7572 | -0.145 | 0.0902 | No | ||
| 42 | SMAD3 | na | 7616 | -0.147 | 0.1111 | No | ||
| 43 | TJP1 | na | 7697 | -0.151 | 0.1290 | No | ||
| 44 | TGFBR1 | na | 8205 | -0.182 | 0.1096 | No | ||
| 45 | APC | na | 8466 | -0.198 | 0.1176 | No | ||
| 46 | NCOR2 | na | 9320 | -0.262 | 0.0775 | No |