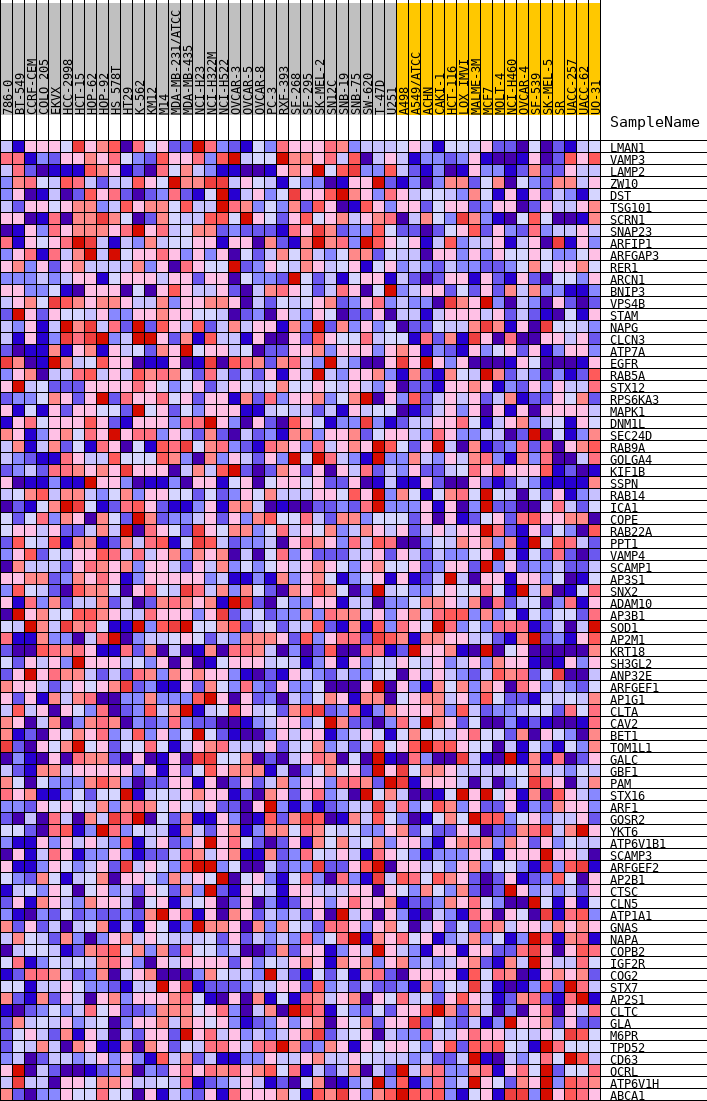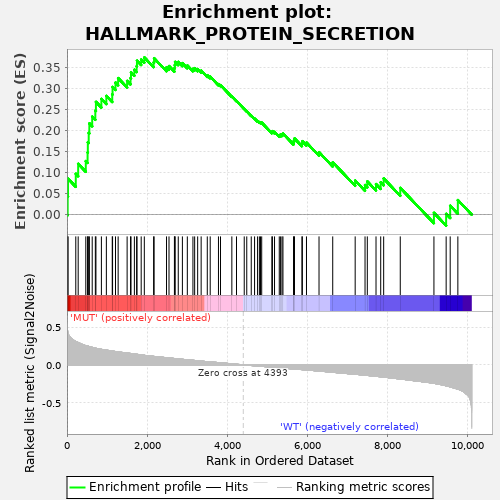
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
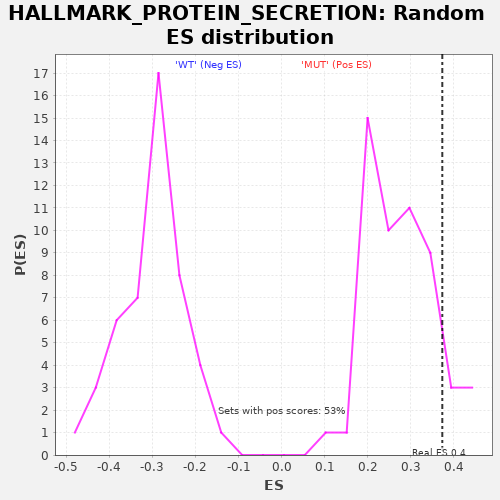
| GeneSet | HALLMARK_PROTEIN_SECRETION |
| Enrichment Score (ES) | 0.37332973 |
| Normalized Enrichment Score (NES) | 1.3577178 |
| Nominal p-value | 0.11320755 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.86 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LMAN1 | na | 17 | 0.451 | 0.0435 | Yes | ||
| 2 | VAMP3 | na | 26 | 0.425 | 0.0854 | Yes | ||
| 3 | LAMP2 | na | 219 | 0.308 | 0.0971 | Yes | ||
| 4 | ZW10 | na | 279 | 0.294 | 0.1207 | Yes | ||
| 5 | DST | na | 469 | 0.253 | 0.1272 | Yes | ||
| 6 | TSG101 | na | 514 | 0.247 | 0.1476 | Yes | ||
| 7 | SCRN1 | na | 520 | 0.246 | 0.1719 | Yes | ||
| 8 | SNAP23 | na | 542 | 0.243 | 0.1941 | Yes | ||
| 9 | ARFIP1 | na | 558 | 0.240 | 0.2168 | Yes | ||
| 10 | ARFGAP3 | na | 629 | 0.231 | 0.2329 | Yes | ||
| 11 | RER1 | na | 707 | 0.220 | 0.2473 | Yes | ||
| 12 | ARCN1 | na | 722 | 0.218 | 0.2678 | Yes | ||
| 13 | BNIP3 | na | 858 | 0.204 | 0.2748 | Yes | ||
| 14 | VPS4B | na | 982 | 0.193 | 0.2820 | Yes | ||
| 15 | STAM | na | 1129 | 0.182 | 0.2857 | Yes | ||
| 16 | NAPG | na | 1137 | 0.182 | 0.3032 | Yes | ||
| 17 | CLCN3 | na | 1210 | 0.175 | 0.3136 | Yes | ||
| 18 | ATP7A | na | 1276 | 0.171 | 0.3242 | Yes | ||
| 19 | EGFR | na | 1500 | 0.156 | 0.3176 | Yes | ||
| 20 | RAB5A | na | 1585 | 0.150 | 0.3242 | Yes | ||
| 21 | STX12 | na | 1597 | 0.149 | 0.3380 | Yes | ||
| 22 | RPS6KA3 | na | 1681 | 0.143 | 0.3441 | Yes | ||
| 23 | MAPK1 | na | 1738 | 0.139 | 0.3524 | Yes | ||
| 24 | DNM1L | na | 1746 | 0.138 | 0.3656 | Yes | ||
| 25 | SEC24D | na | 1851 | 0.132 | 0.3684 | Yes | ||
| 26 | RAB9A | na | 1929 | 0.125 | 0.3733 | Yes | ||
| 27 | GOLGA4 | na | 2161 | 0.111 | 0.3614 | No | ||
| 28 | KIF1B | na | 2173 | 0.110 | 0.3714 | No | ||
| 29 | SSPN | na | 2483 | 0.093 | 0.3499 | No | ||
| 30 | RAB14 | na | 2545 | 0.090 | 0.3528 | No | ||
| 31 | ICA1 | na | 2680 | 0.083 | 0.3478 | No | ||
| 32 | COPE | na | 2689 | 0.082 | 0.3552 | No | ||
| 33 | RAB22A | na | 2697 | 0.082 | 0.3627 | No | ||
| 34 | PPT1 | na | 2776 | 0.077 | 0.3627 | No | ||
| 35 | VAMP4 | na | 2879 | 0.072 | 0.3598 | No | ||
| 36 | SCAMP1 | na | 3002 | 0.066 | 0.3542 | No | ||
| 37 | AP3S1 | na | 3142 | 0.059 | 0.3463 | No | ||
| 38 | SNX2 | na | 3185 | 0.057 | 0.3478 | No | ||
| 39 | ADAM10 | na | 3260 | 0.054 | 0.3458 | No | ||
| 40 | AP3B1 | na | 3347 | 0.049 | 0.3421 | No | ||
| 41 | SOD1 | na | 3500 | 0.041 | 0.3310 | No | ||
| 42 | AP2M1 | na | 3573 | 0.038 | 0.3276 | No | ||
| 43 | KRT18 | na | 3779 | 0.028 | 0.3100 | No | ||
| 44 | SH3GL2 | na | 3830 | 0.026 | 0.3076 | No | ||
| 45 | ANP32E | na | 4113 | 0.012 | 0.2807 | No | ||
| 46 | ARFGEF1 | na | 4229 | 0.007 | 0.2699 | No | ||
| 47 | AP1G1 | na | 4422 | -0.002 | 0.2509 | No | ||
| 48 | CLTA | na | 4485 | -0.004 | 0.2451 | No | ||
| 49 | CAV2 | na | 4597 | -0.009 | 0.2350 | No | ||
| 50 | BET1 | na | 4680 | -0.013 | 0.2281 | No | ||
| 51 | TOM1L1 | na | 4755 | -0.017 | 0.2225 | No | ||
| 52 | GALC | na | 4800 | -0.019 | 0.2200 | No | ||
| 53 | GBF1 | na | 4828 | -0.020 | 0.2193 | No | ||
| 54 | PAM | na | 4858 | -0.021 | 0.2185 | No | ||
| 55 | STX16 | na | 5112 | -0.031 | 0.1963 | No | ||
| 56 | ARF1 | na | 5125 | -0.032 | 0.1983 | No | ||
| 57 | GOSR2 | na | 5178 | -0.034 | 0.1965 | No | ||
| 58 | YKT6 | na | 5296 | -0.039 | 0.1888 | No | ||
| 59 | ATP6V1B1 | na | 5325 | -0.041 | 0.1901 | No | ||
| 60 | SCAMP3 | na | 5363 | -0.043 | 0.1907 | No | ||
| 61 | ARFGEF2 | na | 5388 | -0.044 | 0.1926 | No | ||
| 62 | AP2B1 | na | 5652 | -0.055 | 0.1719 | No | ||
| 63 | CTSC | na | 5662 | -0.056 | 0.1766 | No | ||
| 64 | CLN5 | na | 5676 | -0.056 | 0.1810 | No | ||
| 65 | ATP1A1 | na | 5862 | -0.064 | 0.1689 | No | ||
| 66 | GNAS | na | 5872 | -0.064 | 0.1744 | No | ||
| 67 | NAPA | na | 5975 | -0.069 | 0.1711 | No | ||
| 68 | COPB2 | na | 6288 | -0.083 | 0.1483 | No | ||
| 69 | IGF2R | na | 6630 | -0.099 | 0.1243 | No | ||
| 70 | COG2 | na | 7190 | -0.125 | 0.0811 | No | ||
| 71 | STX7 | na | 7436 | -0.138 | 0.0704 | No | ||
| 72 | AP2S1 | na | 7493 | -0.141 | 0.0789 | No | ||
| 73 | CLTC | na | 7710 | -0.152 | 0.0726 | No | ||
| 74 | GLA | na | 7827 | -0.158 | 0.0769 | No | ||
| 75 | M6PR | na | 7901 | -0.163 | 0.0860 | No | ||
| 76 | TPD52 | na | 8315 | -0.188 | 0.0636 | No | ||
| 77 | CD63 | na | 9155 | -0.245 | 0.0044 | No | ||
| 78 | OCRL | na | 9460 | -0.278 | 0.0020 | No | ||
| 79 | ATP6V1H | na | 9561 | -0.292 | 0.0212 | No | ||
| 80 | ABCA1 | na | 9752 | -0.322 | 0.0346 | No |