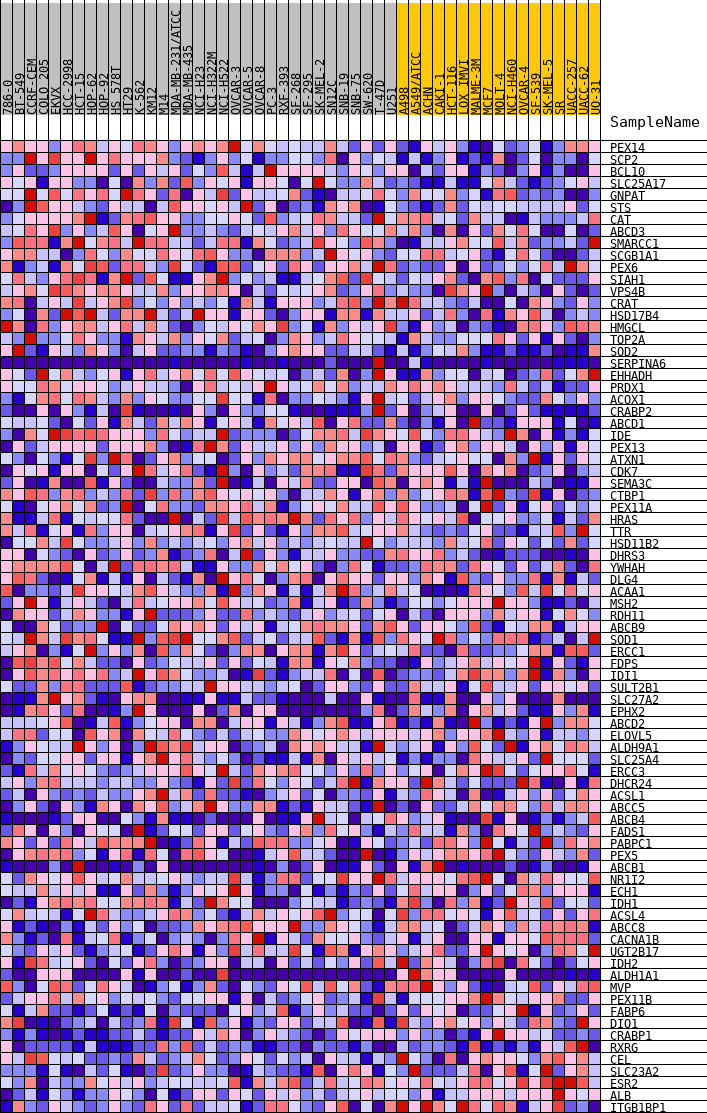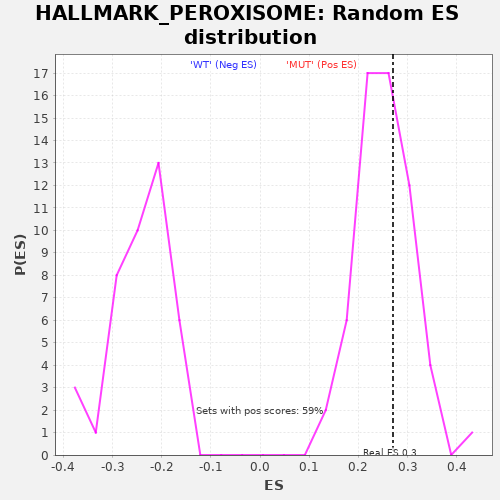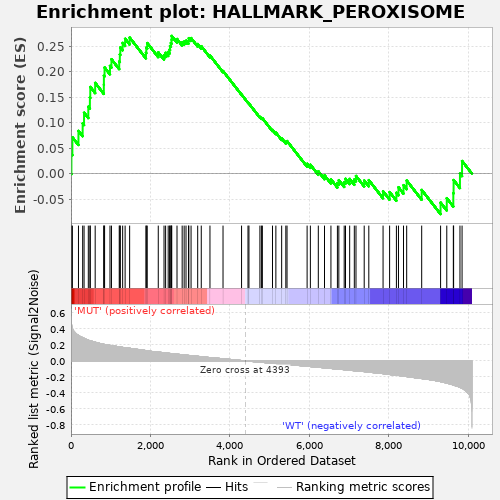
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_PEROXISOME |
| Enrichment Score (ES) | 0.2703663 |
| Normalized Enrichment Score (NES) | 1.0584946 |
| Nominal p-value | 0.37288135 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PEX14 | na | 11 | 0.457 | 0.0384 | Yes | ||
| 2 | SCP2 | na | 35 | 0.408 | 0.0714 | Yes | ||
| 3 | BCL10 | na | 188 | 0.320 | 0.0839 | Yes | ||
| 4 | SLC25A17 | na | 294 | 0.291 | 0.0986 | Yes | ||
| 5 | GNPAT | na | 329 | 0.282 | 0.1197 | Yes | ||
| 6 | STS | na | 436 | 0.258 | 0.1315 | Yes | ||
| 7 | CAT | na | 476 | 0.252 | 0.1494 | Yes | ||
| 8 | ABCD3 | na | 486 | 0.251 | 0.1702 | Yes | ||
| 9 | SMARCC1 | na | 609 | 0.233 | 0.1782 | Yes | ||
| 10 | SCGB1A1 | na | 827 | 0.207 | 0.1744 | Yes | ||
| 11 | PEX6 | na | 829 | 0.207 | 0.1922 | Yes | ||
| 12 | SIAH1 | na | 846 | 0.205 | 0.2084 | Yes | ||
| 13 | VPS4B | na | 982 | 0.193 | 0.2117 | Yes | ||
| 14 | CRAT | na | 1019 | 0.191 | 0.2246 | Yes | ||
| 15 | HSD17B4 | na | 1215 | 0.175 | 0.2203 | Yes | ||
| 16 | HMGCL | na | 1230 | 0.174 | 0.2340 | Yes | ||
| 17 | TOP2A | na | 1242 | 0.173 | 0.2478 | Yes | ||
| 18 | SOD2 | na | 1301 | 0.168 | 0.2566 | Yes | ||
| 19 | SERPINA6 | na | 1363 | 0.165 | 0.2648 | Yes | ||
| 20 | EHHADH | na | 1477 | 0.157 | 0.2671 | Yes | ||
| 21 | PRDX1 | na | 1888 | 0.129 | 0.2373 | Yes | ||
| 22 | ACOX1 | na | 1897 | 0.128 | 0.2476 | Yes | ||
| 23 | CRABP2 | na | 1920 | 0.126 | 0.2563 | Yes | ||
| 24 | ABCD1 | na | 2198 | 0.109 | 0.2381 | Yes | ||
| 25 | IDE | na | 2343 | 0.101 | 0.2325 | Yes | ||
| 26 | PEX13 | na | 2383 | 0.099 | 0.2372 | Yes | ||
| 27 | ATXN1 | na | 2448 | 0.095 | 0.2390 | Yes | ||
| 28 | CDK7 | na | 2484 | 0.093 | 0.2436 | Yes | ||
| 29 | SEMA3C | na | 2495 | 0.092 | 0.2506 | Yes | ||
| 30 | CTBP1 | na | 2516 | 0.091 | 0.2565 | Yes | ||
| 31 | PEX11A | na | 2533 | 0.091 | 0.2627 | Yes | ||
| 32 | HRAS | na | 2536 | 0.090 | 0.2704 | Yes | ||
| 33 | TTR | na | 2670 | 0.084 | 0.2643 | No | ||
| 34 | HSD11B2 | na | 2800 | 0.076 | 0.2580 | No | ||
| 35 | DHRS3 | na | 2850 | 0.074 | 0.2595 | No | ||
| 36 | YWHAH | na | 2897 | 0.071 | 0.2611 | No | ||
| 37 | DLG4 | na | 2959 | 0.068 | 0.2609 | No | ||
| 38 | ACAA1 | na | 2968 | 0.068 | 0.2660 | No | ||
| 39 | MSH2 | na | 3024 | 0.065 | 0.2661 | No | ||
| 40 | RDH11 | na | 3191 | 0.057 | 0.2545 | No | ||
| 41 | ABCB9 | na | 3281 | 0.052 | 0.2501 | No | ||
| 42 | SOD1 | na | 3500 | 0.041 | 0.2319 | No | ||
| 43 | ERCC1 | na | 3829 | 0.026 | 0.2014 | No | ||
| 44 | FDPS | na | 4296 | 0.004 | 0.1552 | No | ||
| 45 | IDI1 | na | 4458 | -0.003 | 0.1394 | No | ||
| 46 | SULT2B1 | na | 4482 | -0.004 | 0.1375 | No | ||
| 47 | SLC27A2 | na | 4758 | -0.017 | 0.1115 | No | ||
| 48 | EPHX2 | na | 4796 | -0.019 | 0.1094 | No | ||
| 49 | ABCD2 | na | 4820 | -0.020 | 0.1088 | No | ||
| 50 | ELOVL5 | na | 5075 | -0.029 | 0.0860 | No | ||
| 51 | ALDH9A1 | na | 5160 | -0.033 | 0.0805 | No | ||
| 52 | SLC25A4 | na | 5306 | -0.040 | 0.0695 | No | ||
| 53 | ERCC3 | na | 5411 | -0.045 | 0.0629 | No | ||
| 54 | DHCR24 | na | 5441 | -0.046 | 0.0640 | No | ||
| 55 | ACSL1 | na | 5947 | -0.067 | 0.0194 | No | ||
| 56 | ABCC5 | na | 6029 | -0.071 | 0.0175 | No | ||
| 57 | ABCB4 | na | 6229 | -0.080 | 0.0046 | No | ||
| 58 | FADS1 | na | 6385 | -0.088 | -0.0033 | No | ||
| 59 | PABPC1 | na | 6547 | -0.095 | -0.0111 | No | ||
| 60 | PEX5 | na | 6710 | -0.103 | -0.0184 | No | ||
| 61 | ABCB1 | na | 6746 | -0.105 | -0.0128 | No | ||
| 62 | NR1I2 | na | 6881 | -0.111 | -0.0166 | No | ||
| 63 | ECH1 | na | 6914 | -0.112 | -0.0101 | No | ||
| 64 | IDH1 | na | 7019 | -0.117 | -0.0103 | No | ||
| 65 | ACSL4 | na | 7136 | -0.123 | -0.0112 | No | ||
| 66 | ABCC8 | na | 7178 | -0.125 | -0.0045 | No | ||
| 67 | CACNA1B | na | 7385 | -0.135 | -0.0134 | No | ||
| 68 | UGT2B17 | na | 7501 | -0.141 | -0.0127 | No | ||
| 69 | IDH2 | na | 7860 | -0.160 | -0.0346 | No | ||
| 70 | ALDH1A1 | na | 8025 | -0.171 | -0.0361 | No | ||
| 71 | MVP | na | 8196 | -0.181 | -0.0375 | No | ||
| 72 | PEX11B | na | 8248 | -0.184 | -0.0267 | No | ||
| 73 | FABP6 | na | 8375 | -0.192 | -0.0226 | No | ||
| 74 | DIO1 | na | 8455 | -0.197 | -0.0135 | No | ||
| 75 | CRABP1 | na | 8833 | -0.222 | -0.0320 | No | ||
| 76 | RXRG | na | 9309 | -0.261 | -0.0568 | No | ||
| 77 | CEL | na | 9465 | -0.279 | -0.0481 | No | ||
| 78 | SLC23A2 | na | 9630 | -0.303 | -0.0383 | No | ||
| 79 | ESR2 | na | 9637 | -0.305 | -0.0125 | No | ||
| 80 | ALB | na | 9797 | -0.331 | 0.0003 | No | ||
| 81 | ITGB1BP1 | na | 9850 | -0.343 | 0.0249 | No |