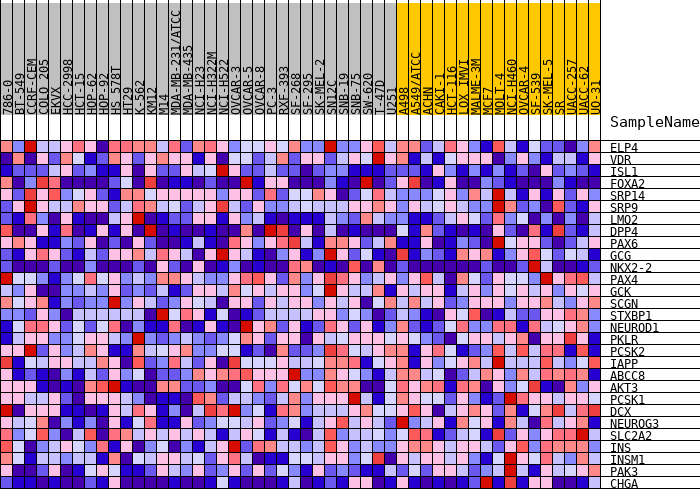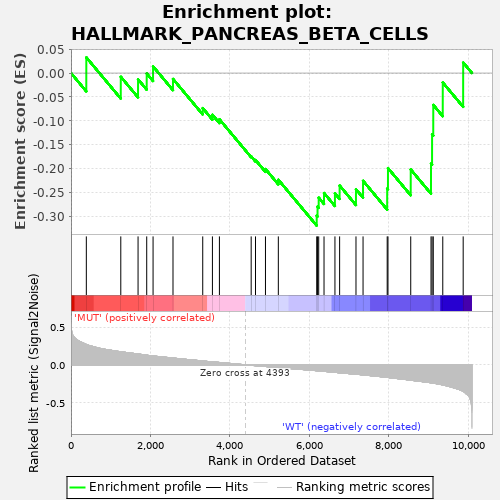
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
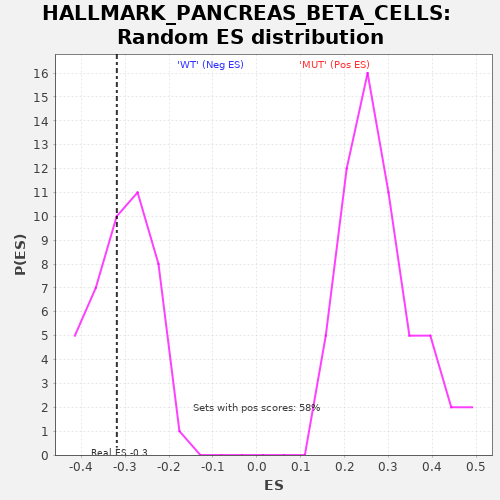
| GeneSet | HALLMARK_PANCREAS_BETA_CELLS |
| Enrichment Score (ES) | -0.31914747 |
| Normalized Enrichment Score (NES) | -1.0450974 |
| Nominal p-value | 0.3809524 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.99 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ELP4 | na | 386 | 0.268 | 0.0327 | No | ||
| 2 | VDR | na | 1254 | 0.172 | -0.0078 | No | ||
| 3 | ISL1 | na | 1689 | 0.143 | -0.0132 | No | ||
| 4 | FOXA2 | na | 1907 | 0.128 | -0.0010 | No | ||
| 5 | SRP14 | na | 2067 | 0.116 | 0.0139 | No | ||
| 6 | SRP9 | na | 2569 | 0.089 | -0.0124 | No | ||
| 7 | LMO2 | na | 3318 | 0.050 | -0.0734 | No | ||
| 8 | DPP4 | na | 3562 | 0.038 | -0.0873 | No | ||
| 9 | PAX6 | na | 3738 | 0.030 | -0.0967 | No | ||
| 10 | GCG | na | 4538 | -0.007 | -0.1743 | No | ||
| 11 | NKX2-2 | na | 4647 | -0.012 | -0.1819 | No | ||
| 12 | PAX4 | na | 4898 | -0.022 | -0.2008 | No | ||
| 13 | GCK | na | 5222 | -0.036 | -0.2232 | No | ||
| 14 | SCGN | na | 6189 | -0.078 | -0.2984 | Yes | ||
| 15 | STXBP1 | na | 6210 | -0.080 | -0.2794 | Yes | ||
| 16 | NEUROD1 | na | 6239 | -0.081 | -0.2608 | Yes | ||
| 17 | PKLR | na | 6373 | -0.087 | -0.2510 | Yes | ||
| 18 | PCSK2 | na | 6648 | -0.100 | -0.2518 | Yes | ||
| 19 | IAPP | na | 6766 | -0.106 | -0.2354 | Yes | ||
| 20 | ABCC8 | na | 7178 | -0.125 | -0.2432 | Yes | ||
| 21 | AKT3 | na | 7356 | -0.133 | -0.2254 | Yes | ||
| 22 | PCSK1 | na | 7964 | -0.167 | -0.2416 | Yes | ||
| 23 | DCX | na | 7984 | -0.168 | -0.1990 | Yes | ||
| 24 | NEUROG3 | na | 8557 | -0.204 | -0.2018 | Yes | ||
| 25 | SLC2A2 | na | 9068 | -0.239 | -0.1893 | Yes | ||
| 26 | INS | na | 9095 | -0.241 | -0.1281 | Yes | ||
| 27 | INSM1 | na | 9125 | -0.243 | -0.0667 | Yes | ||
| 28 | PAK3 | na | 9364 | -0.266 | -0.0198 | Yes | ||
| 29 | CHGA | na | 9878 | -0.350 | 0.0219 | Yes |