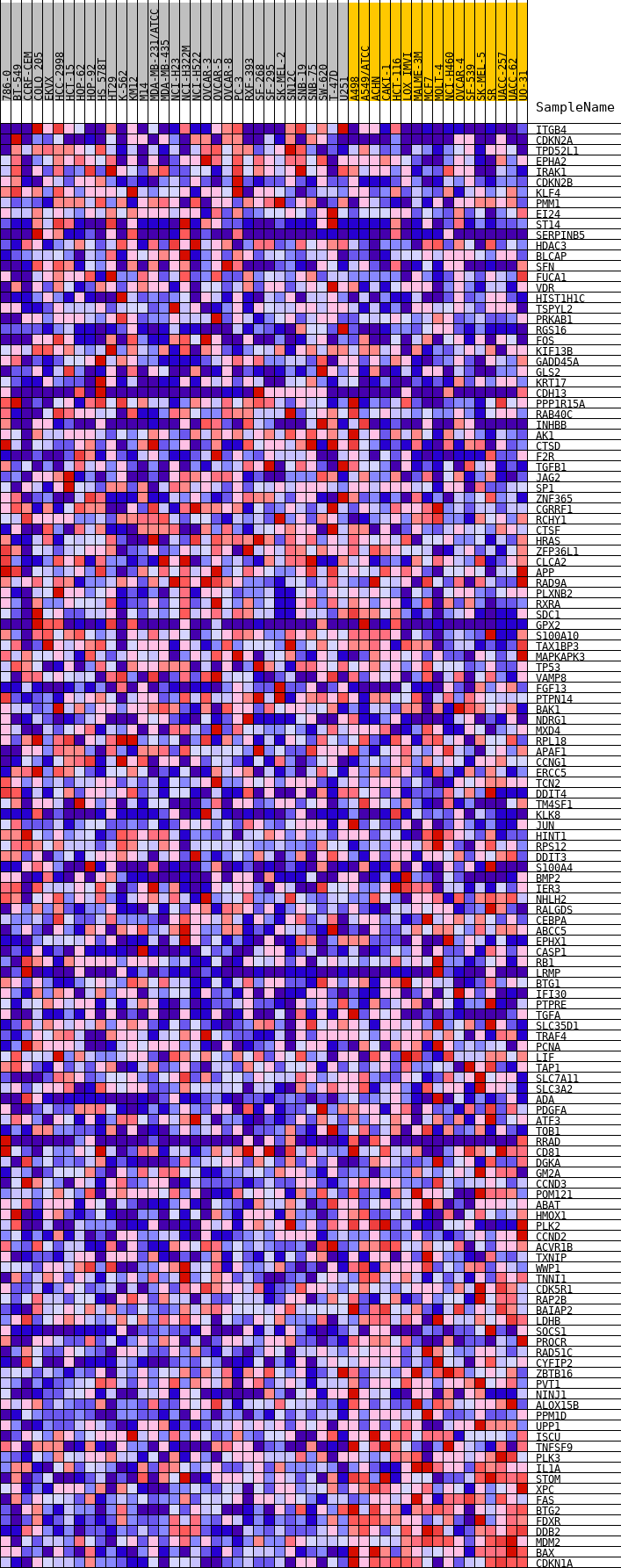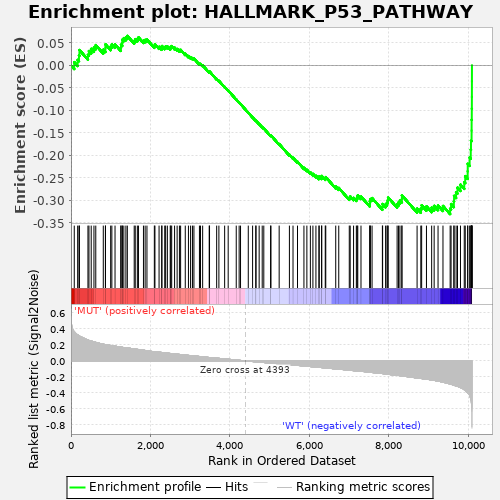
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_P53_PATHWAY |
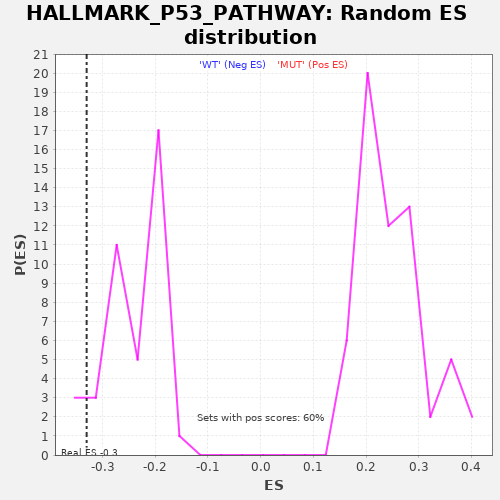
| Enrichment Score (ES) | -0.3303733 |
| Normalized Enrichment Score (NES) | -1.3743259 |
| Nominal p-value | 0.1 |
| FDR q-value | 0.6998731 |
| FWER p-Value | 0.73 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ITGB4 | na | 82 | 0.363 | 0.0070 | No | ||
| 2 | CDKN2A | na | 168 | 0.324 | 0.0121 | No | ||
| 3 | TPD52L1 | na | 201 | 0.314 | 0.0221 | No | ||
| 4 | EPHA2 | na | 213 | 0.310 | 0.0340 | No | ||
| 5 | IRAK1 | na | 427 | 0.260 | 0.0236 | No | ||
| 6 | CDKN2B | na | 452 | 0.256 | 0.0319 | No | ||
| 7 | KLF4 | na | 509 | 0.247 | 0.0367 | No | ||
| 8 | PMM1 | na | 578 | 0.238 | 0.0398 | No | ||
| 9 | EI24 | na | 624 | 0.231 | 0.0450 | No | ||
| 10 | ST14 | na | 813 | 0.208 | 0.0349 | No | ||
| 11 | SERPINB5 | na | 867 | 0.203 | 0.0381 | No | ||
| 12 | HDAC3 | na | 868 | 0.203 | 0.0467 | No | ||
| 13 | BLCAP | na | 999 | 0.192 | 0.0417 | No | ||
| 14 | SFN | na | 1029 | 0.190 | 0.0468 | No | ||
| 15 | FUCA1 | na | 1109 | 0.184 | 0.0466 | No | ||
| 16 | VDR | na | 1254 | 0.172 | 0.0393 | No | ||
| 17 | HIST1H1C | na | 1260 | 0.172 | 0.0460 | No | ||
| 18 | TSPYL2 | na | 1291 | 0.169 | 0.0501 | No | ||
| 19 | PRKAB1 | na | 1293 | 0.169 | 0.0572 | No | ||
| 20 | RGS16 | na | 1331 | 0.166 | 0.0604 | No | ||
| 21 | FOS | na | 1383 | 0.163 | 0.0622 | No | ||
| 22 | KIF13B | na | 1419 | 0.161 | 0.0654 | No | ||
| 23 | GADD45A | na | 1592 | 0.149 | 0.0544 | No | ||
| 24 | GLS2 | na | 1618 | 0.147 | 0.0581 | No | ||
| 25 | KRT17 | na | 1671 | 0.144 | 0.0589 | No | ||
| 26 | CDH13 | na | 1699 | 0.142 | 0.0622 | No | ||
| 27 | PPP1R15A | na | 1825 | 0.134 | 0.0552 | No | ||
| 28 | RAB40C | na | 1867 | 0.130 | 0.0566 | No | ||
| 29 | INHBB | na | 1913 | 0.127 | 0.0574 | No | ||
| 30 | AK1 | na | 2099 | 0.114 | 0.0436 | No | ||
| 31 | CTSD | na | 2114 | 0.113 | 0.0470 | No | ||
| 32 | F2R | na | 2218 | 0.108 | 0.0412 | No | ||
| 33 | TGFB1 | na | 2285 | 0.105 | 0.0390 | No | ||
| 34 | JAG2 | na | 2290 | 0.104 | 0.0430 | No | ||
| 35 | SP1 | na | 2361 | 0.100 | 0.0401 | No | ||
| 36 | ZNF365 | na | 2384 | 0.098 | 0.0421 | No | ||
| 37 | CGRRF1 | na | 2427 | 0.097 | 0.0419 | No | ||
| 38 | RCHY1 | na | 2499 | 0.092 | 0.0387 | No | ||
| 39 | CTSF | na | 2512 | 0.091 | 0.0413 | No | ||
| 40 | HRAS | na | 2536 | 0.090 | 0.0428 | No | ||
| 41 | ZFP36L1 | na | 2609 | 0.087 | 0.0392 | No | ||
| 42 | CLCA2 | na | 2673 | 0.083 | 0.0364 | No | ||
| 43 | APP | na | 2731 | 0.080 | 0.0340 | No | ||
| 44 | RAD9A | na | 2761 | 0.079 | 0.0344 | No | ||
| 45 | PLXNB2 | na | 2878 | 0.072 | 0.0258 | No | ||
| 46 | RXRA | na | 2958 | 0.068 | 0.0207 | No | ||
| 47 | SDC1 | na | 3012 | 0.066 | 0.0182 | No | ||
| 48 | GPX2 | na | 3059 | 0.064 | 0.0162 | No | ||
| 49 | S100A10 | na | 3096 | 0.061 | 0.0152 | No | ||
| 50 | TAX1BP3 | na | 3235 | 0.054 | 0.0036 | No | ||
| 51 | MAPKAPK3 | na | 3262 | 0.053 | 0.0033 | No | ||
| 52 | TP53 | na | 3316 | 0.050 | 0.0001 | No | ||
| 53 | VAMP8 | na | 3481 | 0.042 | -0.0147 | No | ||
| 54 | FGF13 | na | 3489 | 0.041 | -0.0136 | No | ||
| 55 | PTPN14 | na | 3669 | 0.033 | -0.0302 | No | ||
| 56 | BAK1 | na | 3725 | 0.031 | -0.0344 | No | ||
| 57 | NDRG1 | na | 3870 | 0.024 | -0.0478 | No | ||
| 58 | MXD4 | na | 3959 | 0.020 | -0.0558 | No | ||
| 59 | RPL18 | na | 4162 | 0.010 | -0.0757 | No | ||
| 60 | APAF1 | na | 4235 | 0.007 | -0.0826 | No | ||
| 61 | CCNG1 | na | 4270 | 0.005 | -0.0858 | No | ||
| 62 | ERCC5 | na | 4465 | -0.003 | -0.1052 | No | ||
| 63 | TCN2 | na | 4574 | -0.009 | -0.1156 | No | ||
| 64 | DDIT4 | na | 4655 | -0.012 | -0.1232 | No | ||
| 65 | TM4SF1 | na | 4659 | -0.012 | -0.1230 | No | ||
| 66 | KLK8 | na | 4739 | -0.016 | -0.1302 | No | ||
| 67 | JUN | na | 4816 | -0.019 | -0.1370 | No | ||
| 68 | HINT1 | na | 4856 | -0.021 | -0.1401 | No | ||
| 69 | RPS12 | na | 5027 | -0.028 | -0.1560 | No | ||
| 70 | DDIT3 | na | 5036 | -0.028 | -0.1556 | No | ||
| 71 | S100A4 | na | 5243 | -0.037 | -0.1747 | No | ||
| 72 | BMP2 | na | 5501 | -0.049 | -0.1984 | No | ||
| 73 | IER3 | na | 5592 | -0.053 | -0.2053 | No | ||
| 74 | NHLH2 | na | 5705 | -0.057 | -0.2141 | No | ||
| 75 | RALGDS | na | 5867 | -0.064 | -0.2276 | No | ||
| 76 | CEBPA | na | 5941 | -0.067 | -0.2321 | No | ||
| 77 | ABCC5 | na | 6029 | -0.071 | -0.2378 | No | ||
| 78 | EPHX1 | na | 6093 | -0.074 | -0.2411 | No | ||
| 79 | CASP1 | na | 6169 | -0.077 | -0.2453 | No | ||
| 80 | RB1 | na | 6245 | -0.081 | -0.2494 | No | ||
| 81 | LRMP | na | 6251 | -0.082 | -0.2465 | No | ||
| 82 | BTG1 | na | 6312 | -0.084 | -0.2490 | No | ||
| 83 | IFI30 | na | 6315 | -0.084 | -0.2457 | No | ||
| 84 | PTPRE | na | 6404 | -0.088 | -0.2508 | No | ||
| 85 | TGFA | na | 6417 | -0.089 | -0.2483 | No | ||
| 86 | SLC35D1 | na | 6668 | -0.101 | -0.2691 | No | ||
| 87 | TRAF4 | na | 6744 | -0.105 | -0.2723 | No | ||
| 88 | PCNA | na | 7009 | -0.117 | -0.2938 | No | ||
| 89 | LIF | na | 7032 | -0.118 | -0.2911 | No | ||
| 90 | TAP1 | na | 7117 | -0.122 | -0.2944 | No | ||
| 91 | SLC7A11 | na | 7187 | -0.125 | -0.2960 | No | ||
| 92 | SLC3A2 | na | 7208 | -0.126 | -0.2927 | No | ||
| 93 | ADA | na | 7223 | -0.127 | -0.2888 | No | ||
| 94 | PDGFA | na | 7304 | -0.131 | -0.2914 | No | ||
| 95 | ATF3 | na | 7525 | -0.142 | -0.3075 | No | ||
| 96 | TOB1 | na | 7528 | -0.143 | -0.3017 | No | ||
| 97 | RRAD | na | 7541 | -0.143 | -0.2968 | No | ||
| 98 | CD81 | na | 7583 | -0.146 | -0.2948 | No | ||
| 99 | DGKA | na | 7843 | -0.159 | -0.3141 | No | ||
| 100 | GM2A | na | 7849 | -0.159 | -0.3079 | No | ||
| 101 | CCND3 | na | 7924 | -0.164 | -0.3085 | No | ||
| 102 | POM121 | na | 7959 | -0.166 | -0.3049 | No | ||
| 103 | ABAT | na | 7978 | -0.168 | -0.2997 | No | ||
| 104 | HMOX1 | na | 7988 | -0.168 | -0.2935 | No | ||
| 105 | PLK2 | na | 8214 | -0.182 | -0.3084 | No | ||
| 106 | CCND2 | na | 8250 | -0.184 | -0.3042 | No | ||
| 107 | ACVR1B | na | 8285 | -0.186 | -0.2998 | No | ||
| 108 | TXNIP | na | 8334 | -0.189 | -0.2967 | No | ||
| 109 | WWP1 | na | 8335 | -0.189 | -0.2888 | No | ||
| 110 | TNNI1 | na | 8717 | -0.215 | -0.3180 | No | ||
| 111 | CDK5R1 | na | 8811 | -0.221 | -0.3181 | No | ||
| 112 | RAP2B | na | 8832 | -0.222 | -0.3108 | No | ||
| 113 | BAIAP2 | na | 8953 | -0.230 | -0.3132 | No | ||
| 114 | LDHB | na | 9084 | -0.240 | -0.3161 | No | ||
| 115 | SOCS1 | na | 9147 | -0.244 | -0.3121 | No | ||
| 116 | PROCR | na | 9245 | -0.253 | -0.3112 | No | ||
| 117 | RAD51C | na | 9369 | -0.267 | -0.3123 | No | ||
| 118 | CYFIP2 | na | 9550 | -0.290 | -0.3182 | Yes | ||
| 119 | ZBTB16 | na | 9573 | -0.293 | -0.3081 | Yes | ||
| 120 | PVT1 | na | 9643 | -0.306 | -0.3022 | Yes | ||
| 121 | NINJ1 | na | 9649 | -0.306 | -0.2898 | Yes | ||
| 122 | ALOX15B | na | 9700 | -0.315 | -0.2816 | Yes | ||
| 123 | PPM1D | na | 9735 | -0.319 | -0.2716 | Yes | ||
| 124 | UPP1 | na | 9810 | -0.334 | -0.2650 | Yes | ||
| 125 | ISCU | na | 9905 | -0.361 | -0.2593 | Yes | ||
| 126 | TNFSF9 | na | 9930 | -0.372 | -0.2461 | Yes | ||
| 127 | PLK3 | na | 9991 | -0.400 | -0.2353 | Yes | ||
| 128 | IL1A | na | 9993 | -0.400 | -0.2186 | Yes | ||
| 129 | STOM | na | 10041 | -0.439 | -0.2049 | Yes | ||
| 130 | XPC | na | 10069 | -0.482 | -0.1874 | Yes | ||
| 131 | FAS | na | 10075 | -0.489 | -0.1673 | Yes | ||
| 132 | BTG2 | na | 10090 | -0.567 | -0.1449 | Yes | ||
| 133 | FDXR | na | 10091 | -0.568 | -0.1211 | Yes | ||
| 134 | DDB2 | na | 10095 | -0.595 | -0.0964 | Yes | ||
| 135 | MDM2 | na | 10097 | -0.631 | -0.0700 | Yes | ||
| 136 | BAX | na | 10098 | -0.823 | -0.0354 | Yes | ||
| 137 | CDKN1A | na | 10099 | -0.843 | -0.0000 | Yes |