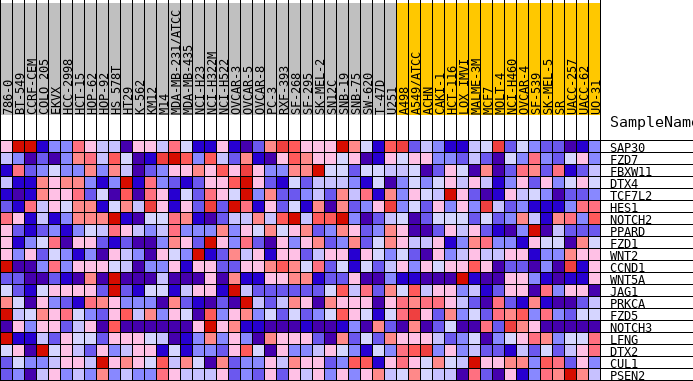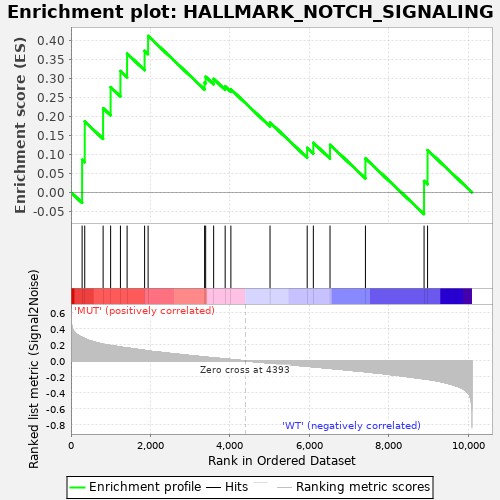
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_NOTCH_SIGNALING |
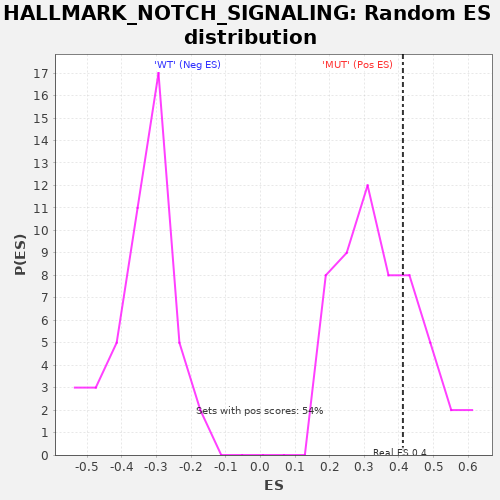
| Enrichment Score (ES) | 0.41131657 |
| Normalized Enrichment Score (NES) | 1.1902404 |
| Nominal p-value | 0.2777778 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.99 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SAP30 | na | 280 | 0.294 | 0.0856 | Yes | ||
| 2 | FZD7 | na | 346 | 0.277 | 0.1863 | Yes | ||
| 3 | FBXW11 | na | 809 | 0.209 | 0.2211 | Yes | ||
| 4 | DTX4 | na | 998 | 0.192 | 0.2767 | Yes | ||
| 5 | TCF7L2 | na | 1245 | 0.173 | 0.3191 | Yes | ||
| 6 | HES1 | na | 1413 | 0.161 | 0.3648 | Yes | ||
| 7 | NOTCH2 | na | 1854 | 0.132 | 0.3720 | Yes | ||
| 8 | PPARD | na | 1942 | 0.124 | 0.4113 | Yes | ||
| 9 | FZD1 | na | 3365 | 0.048 | 0.2886 | No | ||
| 10 | WNT2 | na | 3392 | 0.046 | 0.3038 | No | ||
| 11 | CCND1 | na | 3593 | 0.037 | 0.2982 | No | ||
| 12 | WNT5A | na | 3883 | 0.024 | 0.2787 | No | ||
| 13 | JAG1 | na | 4027 | 0.017 | 0.2709 | No | ||
| 14 | PRKCA | na | 5013 | -0.027 | 0.1836 | No | ||
| 15 | FZD5 | na | 5949 | -0.067 | 0.1169 | No | ||
| 16 | NOTCH3 | na | 6104 | -0.074 | 0.1303 | No | ||
| 17 | LFNG | na | 6524 | -0.094 | 0.1249 | No | ||
| 18 | DTX2 | na | 7416 | -0.136 | 0.0891 | No | ||
| 19 | CUL1 | na | 8893 | -0.226 | 0.0299 | No | ||
| 20 | PSEN2 | na | 8981 | -0.232 | 0.1109 | No |