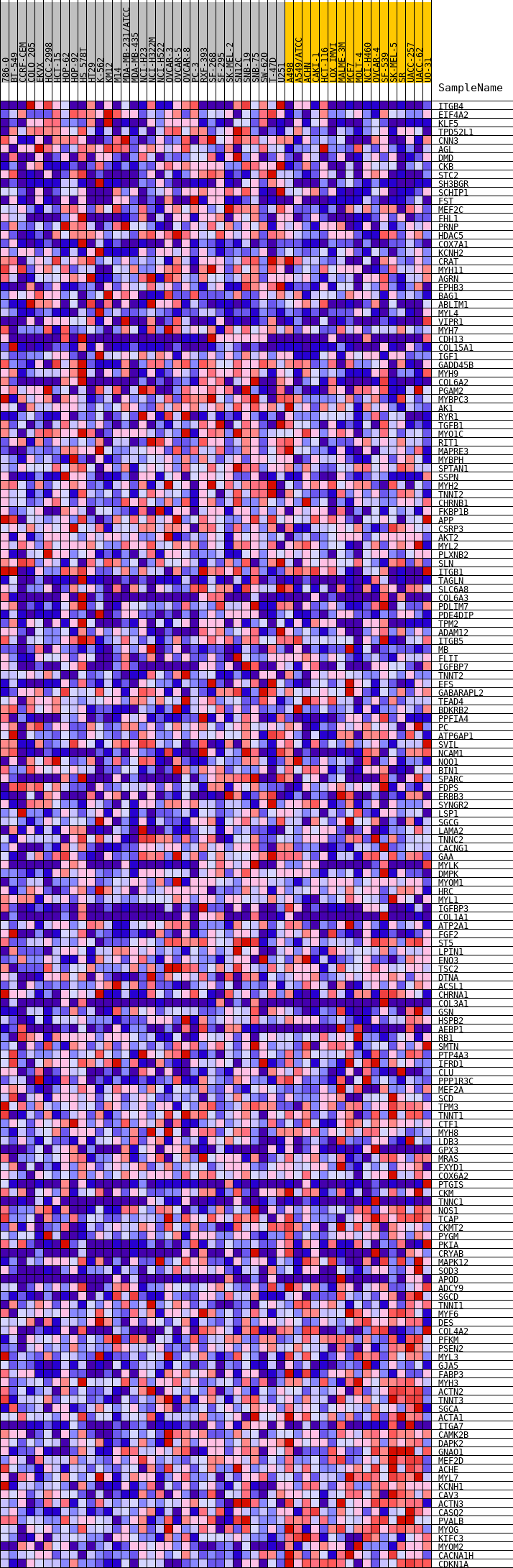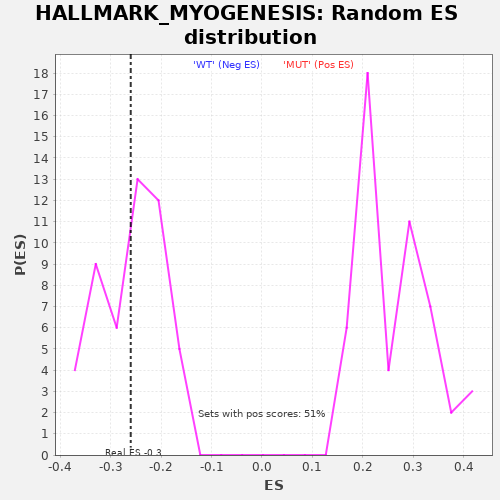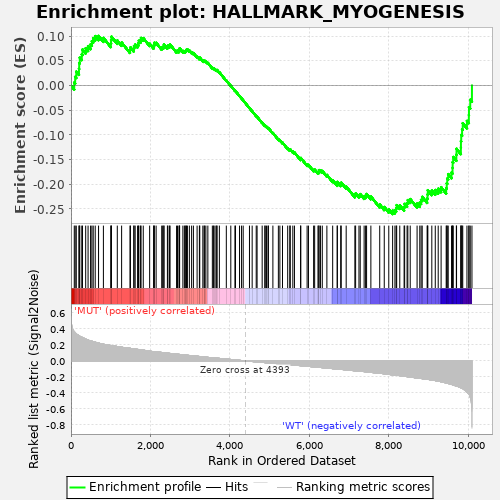
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_MYOGENESIS |
| Enrichment Score (ES) | -0.2604547 |
| Normalized Enrichment Score (NES) | -0.99934936 |
| Nominal p-value | 0.40816328 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ITGB4 | na | 82 | 0.363 | 0.0060 | No | ||
| 2 | EIF4A2 | na | 105 | 0.350 | 0.0176 | No | ||
| 3 | KLF5 | na | 133 | 0.338 | 0.0281 | No | ||
| 4 | TPD52L1 | na | 201 | 0.314 | 0.0337 | No | ||
| 5 | CNN3 | na | 207 | 0.312 | 0.0455 | No | ||
| 6 | AGL | na | 221 | 0.308 | 0.0563 | No | ||
| 7 | DMD | na | 272 | 0.296 | 0.0629 | No | ||
| 8 | CKB | na | 289 | 0.292 | 0.0728 | No | ||
| 9 | STC2 | na | 370 | 0.272 | 0.0754 | No | ||
| 10 | SH3BGR | na | 430 | 0.259 | 0.0796 | No | ||
| 11 | SCHIP1 | na | 493 | 0.250 | 0.0832 | No | ||
| 12 | FST | na | 526 | 0.245 | 0.0896 | No | ||
| 13 | MEF2C | na | 559 | 0.240 | 0.0959 | No | ||
| 14 | FHL1 | na | 610 | 0.232 | 0.1000 | No | ||
| 15 | PRNP | na | 694 | 0.221 | 0.1003 | No | ||
| 16 | HDAC5 | na | 816 | 0.208 | 0.0963 | No | ||
| 17 | COX7A1 | na | 1003 | 0.192 | 0.0851 | No | ||
| 18 | KCNH2 | na | 1010 | 0.192 | 0.0921 | No | ||
| 19 | CRAT | na | 1019 | 0.191 | 0.0988 | No | ||
| 20 | MYH11 | na | 1166 | 0.178 | 0.0911 | No | ||
| 21 | AGRN | na | 1275 | 0.171 | 0.0869 | No | ||
| 22 | EPHB3 | na | 1486 | 0.157 | 0.0719 | No | ||
| 23 | BAG1 | na | 1497 | 0.156 | 0.0771 | No | ||
| 24 | ABLIM1 | na | 1581 | 0.150 | 0.0746 | No | ||
| 25 | MYL4 | na | 1588 | 0.149 | 0.0799 | No | ||
| 26 | VIPR1 | na | 1616 | 0.148 | 0.0830 | No | ||
| 27 | MYH7 | na | 1677 | 0.143 | 0.0825 | No | ||
| 28 | CDH13 | na | 1699 | 0.142 | 0.0860 | No | ||
| 29 | COL15A1 | na | 1709 | 0.141 | 0.0906 | No | ||
| 30 | IGF1 | na | 1753 | 0.138 | 0.0917 | No | ||
| 31 | GADD45B | na | 1763 | 0.137 | 0.0962 | No | ||
| 32 | MYH9 | na | 1820 | 0.134 | 0.0958 | No | ||
| 33 | COL6A2 | na | 1981 | 0.121 | 0.0845 | No | ||
| 34 | PGAM2 | na | 2081 | 0.115 | 0.0791 | No | ||
| 35 | MYBPC3 | na | 2091 | 0.115 | 0.0827 | No | ||
| 36 | AK1 | na | 2099 | 0.114 | 0.0864 | No | ||
| 37 | RYR1 | na | 2144 | 0.112 | 0.0864 | No | ||
| 38 | TGFB1 | na | 2285 | 0.105 | 0.0764 | No | ||
| 39 | MYO1C | na | 2309 | 0.103 | 0.0782 | No | ||
| 40 | RIT1 | na | 2339 | 0.101 | 0.0792 | No | ||
| 41 | MAPRE3 | na | 2341 | 0.101 | 0.0831 | No | ||
| 42 | MYBPH | na | 2428 | 0.097 | 0.0783 | No | ||
| 43 | SPTAN1 | na | 2441 | 0.096 | 0.0808 | No | ||
| 44 | SSPN | na | 2483 | 0.093 | 0.0804 | No | ||
| 45 | MYH2 | na | 2494 | 0.092 | 0.0830 | No | ||
| 46 | TNNI2 | na | 2659 | 0.084 | 0.0698 | No | ||
| 47 | CHRNB1 | na | 2682 | 0.083 | 0.0708 | No | ||
| 48 | FKBP1B | na | 2720 | 0.081 | 0.0703 | No | ||
| 49 | APP | na | 2731 | 0.080 | 0.0724 | No | ||
| 50 | CSRP3 | na | 2736 | 0.080 | 0.0752 | No | ||
| 51 | AKT2 | na | 2818 | 0.075 | 0.0700 | No | ||
| 52 | MYL2 | na | 2856 | 0.073 | 0.0691 | No | ||
| 53 | PLXNB2 | na | 2878 | 0.072 | 0.0698 | No | ||
| 54 | SLN | na | 2896 | 0.071 | 0.0709 | No | ||
| 55 | ITGB1 | na | 2909 | 0.071 | 0.0725 | No | ||
| 56 | TAGLN | na | 2932 | 0.069 | 0.0730 | No | ||
| 57 | SLC6A8 | na | 2981 | 0.067 | 0.0708 | No | ||
| 58 | COL6A3 | na | 3038 | 0.064 | 0.0677 | No | ||
| 59 | PDLIM7 | na | 3083 | 0.062 | 0.0657 | No | ||
| 60 | PDE4DIP | na | 3176 | 0.057 | 0.0587 | No | ||
| 61 | TPM2 | na | 3234 | 0.054 | 0.0551 | No | ||
| 62 | ADAM12 | na | 3243 | 0.054 | 0.0564 | No | ||
| 63 | ITGB5 | na | 3321 | 0.050 | 0.0506 | No | ||
| 64 | MB | na | 3352 | 0.048 | 0.0495 | No | ||
| 65 | FLII | na | 3366 | 0.048 | 0.0501 | No | ||
| 66 | IGFBP7 | na | 3397 | 0.046 | 0.0489 | No | ||
| 67 | TNNT2 | na | 3447 | 0.043 | 0.0456 | No | ||
| 68 | EFS | na | 3560 | 0.038 | 0.0359 | No | ||
| 69 | GABARAPL2 | na | 3587 | 0.037 | 0.0347 | No | ||
| 70 | TEAD4 | na | 3605 | 0.036 | 0.0344 | No | ||
| 71 | BDKRB2 | na | 3650 | 0.034 | 0.0313 | No | ||
| 72 | PPFIA4 | na | 3671 | 0.033 | 0.0306 | No | ||
| 73 | PC | na | 3678 | 0.033 | 0.0313 | No | ||
| 74 | ATP6AP1 | na | 3737 | 0.030 | 0.0267 | No | ||
| 75 | SVIL | na | 3912 | 0.022 | 0.0100 | No | ||
| 76 | NCAM1 | na | 4025 | 0.017 | -0.0006 | No | ||
| 77 | NQO1 | na | 4129 | 0.011 | -0.0105 | No | ||
| 78 | BIN1 | na | 4143 | 0.011 | -0.0114 | No | ||
| 79 | SPARC | na | 4245 | 0.006 | -0.0213 | No | ||
| 80 | FDPS | na | 4296 | 0.004 | -0.0262 | No | ||
| 81 | ERBB3 | na | 4337 | 0.002 | -0.0301 | No | ||
| 82 | SYNGR2 | na | 4495 | -0.005 | -0.0457 | No | ||
| 83 | LSP1 | na | 4563 | -0.008 | -0.0522 | No | ||
| 84 | SGCG | na | 4665 | -0.012 | -0.0619 | No | ||
| 85 | LAMA2 | na | 4687 | -0.013 | -0.0634 | No | ||
| 86 | TNNC2 | na | 4815 | -0.019 | -0.0755 | No | ||
| 87 | CACNG1 | na | 4881 | -0.022 | -0.0812 | No | ||
| 88 | GAA | na | 4892 | -0.022 | -0.0813 | No | ||
| 89 | MYLK | na | 4925 | -0.023 | -0.0836 | No | ||
| 90 | DMPK | na | 4962 | -0.025 | -0.0862 | No | ||
| 91 | MYOM1 | na | 4972 | -0.025 | -0.0861 | No | ||
| 92 | HRC | na | 5085 | -0.030 | -0.0962 | No | ||
| 93 | MYL1 | na | 5223 | -0.036 | -0.1086 | No | ||
| 94 | IGFBP3 | na | 5261 | -0.038 | -0.1108 | No | ||
| 95 | COL1A1 | na | 5327 | -0.041 | -0.1158 | No | ||
| 96 | ATP2A1 | na | 5458 | -0.047 | -0.1270 | No | ||
| 97 | FGF2 | na | 5511 | -0.049 | -0.1303 | No | ||
| 98 | ST5 | na | 5522 | -0.050 | -0.1294 | No | ||
| 99 | LPIN1 | na | 5587 | -0.053 | -0.1338 | No | ||
| 100 | ENO3 | na | 5628 | -0.054 | -0.1357 | No | ||
| 101 | TSC2 | na | 5783 | -0.060 | -0.1488 | No | ||
| 102 | DTNA | na | 5787 | -0.060 | -0.1468 | No | ||
| 103 | ACSL1 | na | 5947 | -0.067 | -0.1601 | No | ||
| 104 | CHRNA1 | na | 5981 | -0.069 | -0.1607 | No | ||
| 105 | COL3A1 | na | 6111 | -0.075 | -0.1708 | No | ||
| 106 | GSN | na | 6137 | -0.076 | -0.1703 | No | ||
| 107 | HSPB2 | na | 6221 | -0.080 | -0.1756 | No | ||
| 108 | AEBP1 | na | 6244 | -0.081 | -0.1746 | No | ||
| 109 | RB1 | na | 6245 | -0.081 | -0.1714 | No | ||
| 110 | SMTN | na | 6283 | -0.083 | -0.1719 | No | ||
| 111 | PTP4A3 | na | 6329 | -0.085 | -0.1730 | No | ||
| 112 | IFRD1 | na | 6445 | -0.090 | -0.1811 | No | ||
| 113 | CLU | na | 6591 | -0.097 | -0.1919 | No | ||
| 114 | PPP1R3C | na | 6706 | -0.103 | -0.1993 | No | ||
| 115 | MEF2A | na | 6707 | -0.103 | -0.1953 | No | ||
| 116 | SCD | na | 6793 | -0.107 | -0.1996 | No | ||
| 117 | TPM3 | na | 6806 | -0.107 | -0.1966 | No | ||
| 118 | TNNT1 | na | 6931 | -0.113 | -0.2047 | No | ||
| 119 | CTF1 | na | 7150 | -0.124 | -0.2217 | No | ||
| 120 | MYH8 | na | 7169 | -0.125 | -0.2187 | No | ||
| 121 | LDB3 | na | 7253 | -0.128 | -0.2220 | No | ||
| 122 | GPX3 | na | 7285 | -0.130 | -0.2200 | No | ||
| 123 | MRAS | na | 7383 | -0.135 | -0.2245 | No | ||
| 124 | FXYD1 | na | 7420 | -0.136 | -0.2227 | No | ||
| 125 | COX6A2 | na | 7445 | -0.138 | -0.2197 | No | ||
| 126 | PTGIS | na | 7553 | -0.144 | -0.2248 | No | ||
| 127 | CKM | na | 7776 | -0.156 | -0.2411 | No | ||
| 128 | TNNC1 | na | 7890 | -0.162 | -0.2461 | No | ||
| 129 | NOS1 | na | 8004 | -0.169 | -0.2508 | No | ||
| 130 | TCAP | na | 8101 | -0.175 | -0.2536 | Yes | ||
| 131 | CKMT2 | na | 8159 | -0.178 | -0.2523 | Yes | ||
| 132 | PYGM | na | 8191 | -0.180 | -0.2483 | Yes | ||
| 133 | PKIA | na | 8203 | -0.181 | -0.2423 | Yes | ||
| 134 | CRYAB | na | 8281 | -0.185 | -0.2428 | Yes | ||
| 135 | MAPK12 | na | 8390 | -0.192 | -0.2461 | Yes | ||
| 136 | SOD3 | na | 8404 | -0.193 | -0.2398 | Yes | ||
| 137 | APOD | na | 8465 | -0.198 | -0.2381 | Yes | ||
| 138 | ADCY9 | na | 8484 | -0.199 | -0.2321 | Yes | ||
| 139 | SGCD | na | 8544 | -0.203 | -0.2301 | Yes | ||
| 140 | TNNI1 | na | 8717 | -0.215 | -0.2389 | Yes | ||
| 141 | MYF6 | na | 8785 | -0.219 | -0.2371 | Yes | ||
| 142 | DES | na | 8818 | -0.221 | -0.2316 | Yes | ||
| 143 | COL4A2 | na | 8846 | -0.223 | -0.2255 | Yes | ||
| 144 | PFKM | na | 8970 | -0.231 | -0.2288 | Yes | ||
| 145 | PSEN2 | na | 8981 | -0.232 | -0.2207 | Yes | ||
| 146 | MYL3 | na | 8987 | -0.232 | -0.2121 | Yes | ||
| 147 | GJA5 | na | 9088 | -0.241 | -0.2127 | Yes | ||
| 148 | FABP3 | na | 9175 | -0.247 | -0.2116 | Yes | ||
| 149 | MYH3 | na | 9249 | -0.253 | -0.2090 | Yes | ||
| 150 | ACTN2 | na | 9322 | -0.262 | -0.2060 | Yes | ||
| 151 | TNNT3 | na | 9450 | -0.277 | -0.2079 | Yes | ||
| 152 | SGCA | na | 9464 | -0.279 | -0.1982 | Yes | ||
| 153 | ACTA1 | na | 9477 | -0.281 | -0.1884 | Yes | ||
| 154 | ITGA7 | na | 9501 | -0.284 | -0.1795 | Yes | ||
| 155 | CAMK2B | na | 9585 | -0.295 | -0.1763 | Yes | ||
| 156 | DAPK2 | na | 9610 | -0.299 | -0.1669 | Yes | ||
| 157 | GNAO1 | na | 9613 | -0.299 | -0.1554 | Yes | ||
| 158 | MEF2D | na | 9627 | -0.303 | -0.1448 | Yes | ||
| 159 | ACHE | na | 9706 | -0.315 | -0.1402 | Yes | ||
| 160 | MYL7 | na | 9711 | -0.316 | -0.1282 | Yes | ||
| 161 | KCNH1 | na | 9820 | -0.336 | -0.1259 | Yes | ||
| 162 | CAV3 | na | 9822 | -0.337 | -0.1127 | Yes | ||
| 163 | ACTN3 | na | 9829 | -0.338 | -0.1000 | Yes | ||
| 164 | CASQ2 | na | 9851 | -0.343 | -0.0887 | Yes | ||
| 165 | PVALB | na | 9867 | -0.348 | -0.0765 | Yes | ||
| 166 | MYOG | na | 9973 | -0.390 | -0.0717 | Yes | ||
| 167 | KIFC3 | na | 10021 | -0.417 | -0.0600 | Yes | ||
| 168 | MYOM2 | na | 10024 | -0.419 | -0.0438 | Yes | ||
| 169 | CACNA1H | na | 10056 | -0.458 | -0.0289 | Yes | ||
| 170 | CDKN1A | na | 10099 | -0.843 | 0.0000 | Yes |