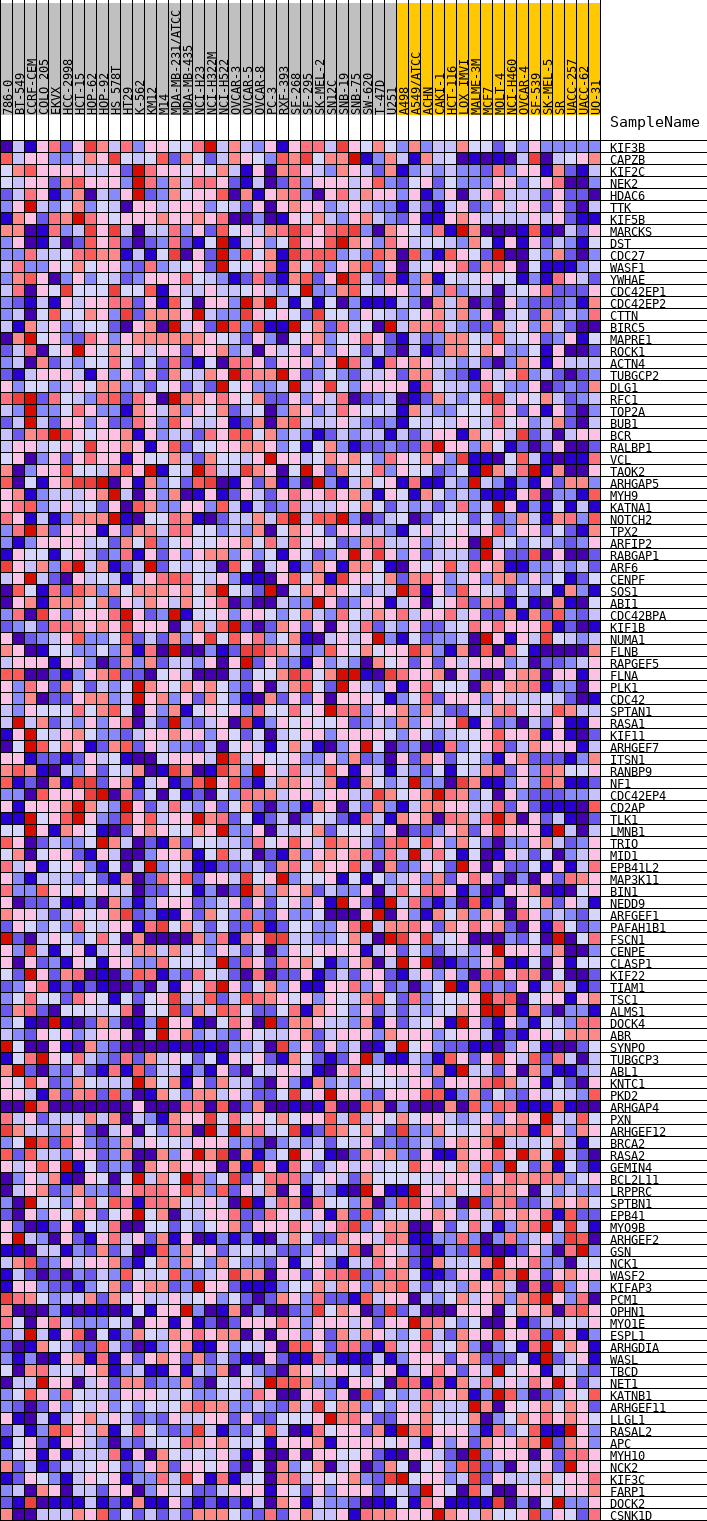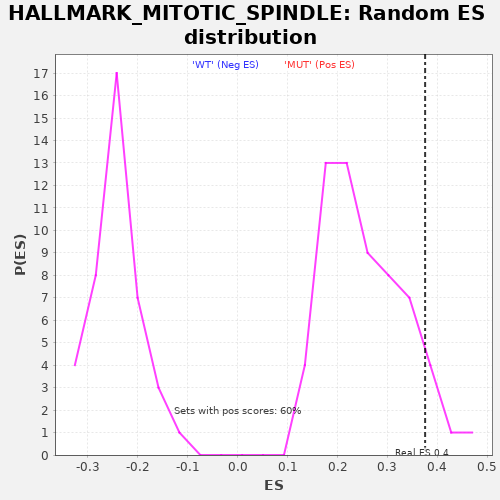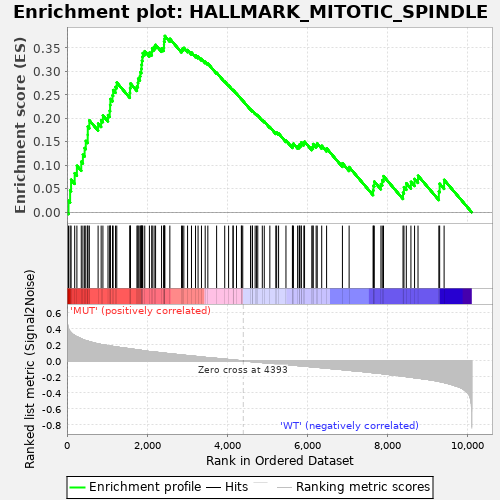
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_MITOTIC_SPINDLE |
| Enrichment Score (ES) | 0.37541014 |
| Normalized Enrichment Score (NES) | 1.4504205 |
| Nominal p-value | 0.083333336 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.69 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KIF3B | na | 38 | 0.400 | 0.0247 | Yes | ||
| 2 | CAPZB | na | 78 | 0.366 | 0.0468 | Yes | ||
| 3 | KIF2C | na | 102 | 0.351 | 0.0695 | Yes | ||
| 4 | NEK2 | na | 192 | 0.318 | 0.0832 | Yes | ||
| 5 | HDAC6 | na | 247 | 0.300 | 0.0991 | Yes | ||
| 6 | TTK | na | 356 | 0.274 | 0.1078 | Yes | ||
| 7 | KIF5B | na | 397 | 0.265 | 0.1227 | Yes | ||
| 8 | MARCKS | na | 440 | 0.258 | 0.1369 | Yes | ||
| 9 | DST | na | 469 | 0.253 | 0.1521 | Yes | ||
| 10 | CDC27 | na | 517 | 0.247 | 0.1649 | Yes | ||
| 11 | WASF1 | na | 519 | 0.246 | 0.1823 | Yes | ||
| 12 | YWHAE | na | 557 | 0.241 | 0.1957 | Yes | ||
| 13 | CDC42EP1 | na | 776 | 0.212 | 0.1890 | Yes | ||
| 14 | CDC42EP2 | na | 852 | 0.205 | 0.1960 | Yes | ||
| 15 | CTTN | na | 896 | 0.201 | 0.2060 | Yes | ||
| 16 | BIRC5 | na | 1023 | 0.191 | 0.2070 | Yes | ||
| 17 | MAPRE1 | na | 1066 | 0.187 | 0.2161 | Yes | ||
| 18 | ROCK1 | na | 1076 | 0.187 | 0.2285 | Yes | ||
| 19 | ACTN4 | na | 1082 | 0.186 | 0.2412 | Yes | ||
| 20 | TUBGCP2 | na | 1135 | 0.182 | 0.2490 | Yes | ||
| 21 | DLG1 | na | 1151 | 0.180 | 0.2603 | Yes | ||
| 22 | RFC1 | na | 1211 | 0.175 | 0.2668 | Yes | ||
| 23 | TOP2A | na | 1242 | 0.173 | 0.2761 | Yes | ||
| 24 | BUB1 | na | 1568 | 0.151 | 0.2543 | Yes | ||
| 25 | BCR | na | 1574 | 0.150 | 0.2645 | Yes | ||
| 26 | RALBP1 | na | 1584 | 0.150 | 0.2742 | Yes | ||
| 27 | VCL | na | 1745 | 0.138 | 0.2681 | Yes | ||
| 28 | TAOK2 | na | 1769 | 0.137 | 0.2755 | Yes | ||
| 29 | ARHGAP5 | na | 1776 | 0.136 | 0.2846 | Yes | ||
| 30 | MYH9 | na | 1820 | 0.134 | 0.2898 | Yes | ||
| 31 | KATNA1 | na | 1830 | 0.133 | 0.2984 | Yes | ||
| 32 | NOTCH2 | na | 1854 | 0.132 | 0.3054 | Yes | ||
| 33 | TPX2 | na | 1860 | 0.130 | 0.3142 | Yes | ||
| 34 | ARFIP2 | na | 1870 | 0.130 | 0.3225 | Yes | ||
| 35 | RABGAP1 | na | 1882 | 0.129 | 0.3306 | Yes | ||
| 36 | ARF6 | na | 1891 | 0.128 | 0.3389 | Yes | ||
| 37 | CENPF | na | 1940 | 0.125 | 0.3430 | Yes | ||
| 38 | SOS1 | na | 2056 | 0.116 | 0.3398 | Yes | ||
| 39 | ABI1 | na | 2117 | 0.113 | 0.3418 | Yes | ||
| 40 | CDC42BPA | na | 2123 | 0.113 | 0.3493 | Yes | ||
| 41 | KIF1B | na | 2173 | 0.110 | 0.3523 | Yes | ||
| 42 | NUMA1 | na | 2210 | 0.109 | 0.3564 | Yes | ||
| 43 | FLNB | na | 2359 | 0.100 | 0.3487 | Yes | ||
| 44 | RAPGEF5 | na | 2414 | 0.097 | 0.3502 | Yes | ||
| 45 | FLNA | na | 2420 | 0.097 | 0.3566 | Yes | ||
| 46 | PLK1 | na | 2422 | 0.097 | 0.3634 | Yes | ||
| 47 | CDC42 | na | 2434 | 0.096 | 0.3692 | Yes | ||
| 48 | SPTAN1 | na | 2441 | 0.096 | 0.3754 | Yes | ||
| 49 | RASA1 | na | 2566 | 0.089 | 0.3693 | No | ||
| 50 | KIF11 | na | 2858 | 0.073 | 0.3454 | No | ||
| 51 | ARHGEF7 | na | 2880 | 0.072 | 0.3484 | No | ||
| 52 | ITSN1 | na | 2915 | 0.070 | 0.3500 | No | ||
| 53 | RANBP9 | na | 3008 | 0.066 | 0.3455 | No | ||
| 54 | NF1 | na | 3106 | 0.061 | 0.3401 | No | ||
| 55 | CDC42EP4 | na | 3207 | 0.056 | 0.3340 | No | ||
| 56 | CD2AP | na | 3270 | 0.053 | 0.3316 | No | ||
| 57 | TLK1 | na | 3355 | 0.048 | 0.3266 | No | ||
| 58 | LMNB1 | na | 3448 | 0.043 | 0.3205 | No | ||
| 59 | TRIO | na | 3514 | 0.040 | 0.3168 | No | ||
| 60 | MID1 | na | 3733 | 0.031 | 0.2972 | No | ||
| 61 | EPB41L2 | na | 3935 | 0.021 | 0.2785 | No | ||
| 62 | MAP3K11 | na | 4032 | 0.016 | 0.2701 | No | ||
| 63 | BIN1 | na | 4143 | 0.011 | 0.2599 | No | ||
| 64 | NEDD9 | na | 4145 | 0.011 | 0.2605 | No | ||
| 65 | ARFGEF1 | na | 4229 | 0.007 | 0.2527 | No | ||
| 66 | PAFAH1B1 | na | 4350 | 0.002 | 0.2408 | No | ||
| 67 | FSCN1 | na | 4359 | 0.002 | 0.2401 | No | ||
| 68 | CENPE | na | 4385 | 0.000 | 0.2377 | No | ||
| 69 | CLASP1 | na | 4579 | -0.009 | 0.2190 | No | ||
| 70 | KIF22 | na | 4624 | -0.010 | 0.2153 | No | ||
| 71 | TIAM1 | na | 4625 | -0.011 | 0.2160 | No | ||
| 72 | TSC1 | na | 4694 | -0.014 | 0.2102 | No | ||
| 73 | ALMS1 | na | 4730 | -0.016 | 0.2078 | No | ||
| 74 | DOCK4 | na | 4760 | -0.017 | 0.2061 | No | ||
| 75 | ABR | na | 4873 | -0.021 | 0.1964 | No | ||
| 76 | SYNPO | na | 4922 | -0.023 | 0.1933 | No | ||
| 77 | TUBGCP3 | na | 5061 | -0.029 | 0.1815 | No | ||
| 78 | ABL1 | na | 5210 | -0.036 | 0.1693 | No | ||
| 79 | KNTC1 | na | 5231 | -0.037 | 0.1699 | No | ||
| 80 | PKD2 | na | 5279 | -0.039 | 0.1679 | No | ||
| 81 | ARHGAP4 | na | 5463 | -0.047 | 0.1529 | No | ||
| 82 | PXN | na | 5624 | -0.054 | 0.1408 | No | ||
| 83 | ARHGEF12 | na | 5631 | -0.054 | 0.1440 | No | ||
| 84 | BRCA2 | na | 5649 | -0.055 | 0.1462 | No | ||
| 85 | RASA2 | na | 5754 | -0.059 | 0.1400 | No | ||
| 86 | GEMIN4 | na | 5793 | -0.061 | 0.1405 | No | ||
| 87 | BCL2L11 | na | 5800 | -0.061 | 0.1443 | No | ||
| 88 | LRPPRC | na | 5841 | -0.063 | 0.1448 | No | ||
| 89 | SPTBN1 | na | 5845 | -0.063 | 0.1490 | No | ||
| 90 | EPB41 | na | 5910 | -0.066 | 0.1472 | No | ||
| 91 | MYO9B | na | 5927 | -0.066 | 0.1503 | No | ||
| 92 | ARHGEF2 | na | 6108 | -0.074 | 0.1376 | No | ||
| 93 | GSN | na | 6137 | -0.076 | 0.1402 | No | ||
| 94 | NCK1 | na | 6140 | -0.076 | 0.1454 | No | ||
| 95 | WASF2 | na | 6216 | -0.080 | 0.1435 | No | ||
| 96 | KIFAP3 | na | 6243 | -0.081 | 0.1467 | No | ||
| 97 | PCM1 | na | 6355 | -0.086 | 0.1417 | No | ||
| 98 | OPHN1 | na | 6477 | -0.092 | 0.1361 | No | ||
| 99 | MYO1E | na | 6872 | -0.110 | 0.1045 | No | ||
| 100 | ESPL1 | na | 7039 | -0.119 | 0.0963 | No | ||
| 101 | ARHGDIA | na | 7637 | -0.148 | 0.0470 | No | ||
| 102 | WASL | na | 7647 | -0.149 | 0.0567 | No | ||
| 103 | TBCD | na | 7667 | -0.150 | 0.0655 | No | ||
| 104 | NET1 | na | 7833 | -0.158 | 0.0602 | No | ||
| 105 | KATNB1 | na | 7866 | -0.161 | 0.0685 | No | ||
| 106 | ARHGEF11 | na | 7897 | -0.162 | 0.0770 | No | ||
| 107 | LLGL1 | na | 8384 | -0.192 | 0.0420 | No | ||
| 108 | RASAL2 | na | 8407 | -0.193 | 0.0536 | No | ||
| 109 | APC | na | 8466 | -0.198 | 0.0618 | No | ||
| 110 | MYH10 | na | 8581 | -0.205 | 0.0650 | No | ||
| 111 | NCK2 | na | 8672 | -0.212 | 0.0710 | No | ||
| 112 | KIF3C | na | 8758 | -0.217 | 0.0780 | No | ||
| 113 | FARP1 | na | 9278 | -0.257 | 0.0443 | No | ||
| 114 | DOCK2 | na | 9298 | -0.259 | 0.0608 | No | ||
| 115 | CSNK1D | na | 9409 | -0.272 | 0.0691 | No |