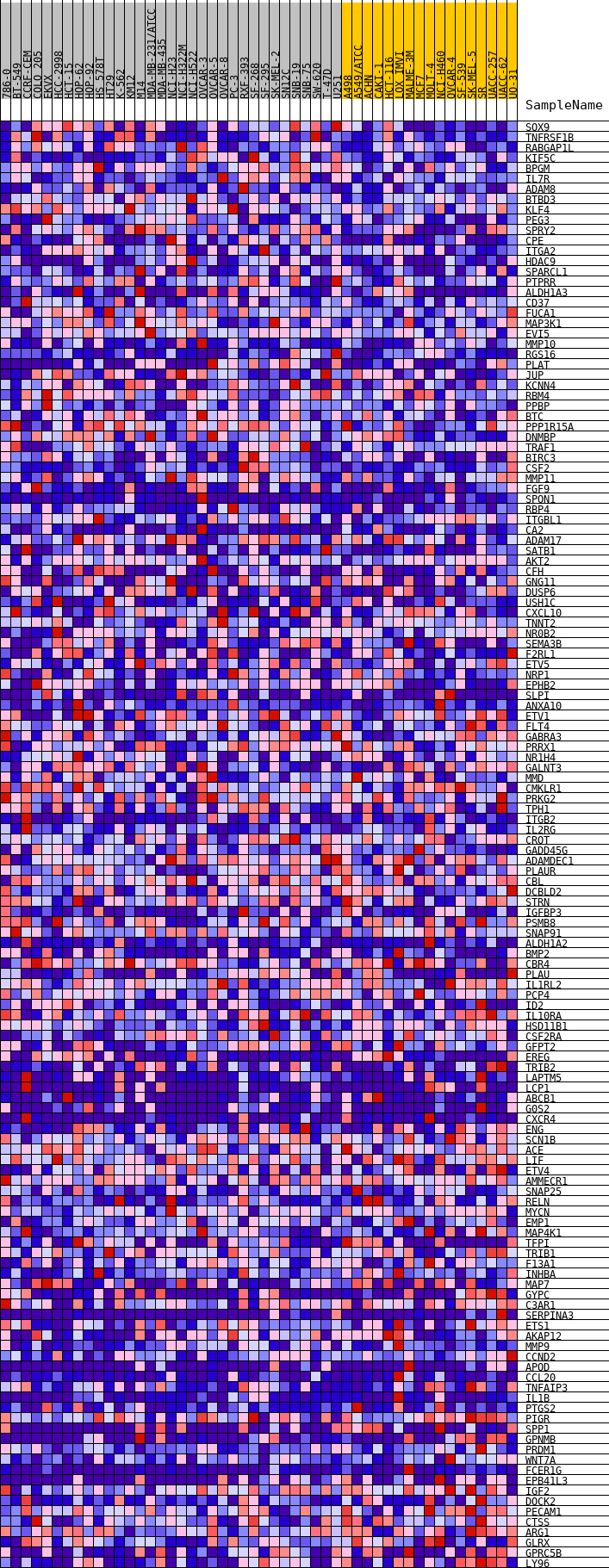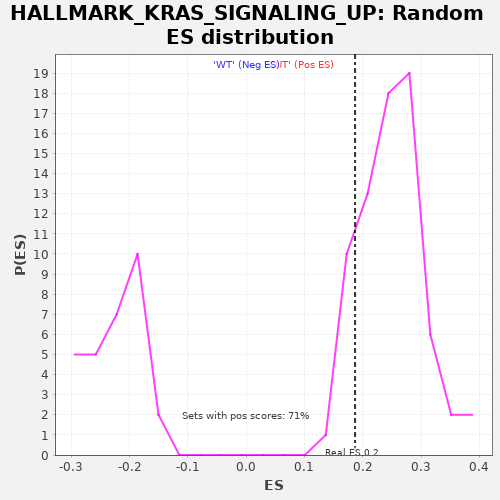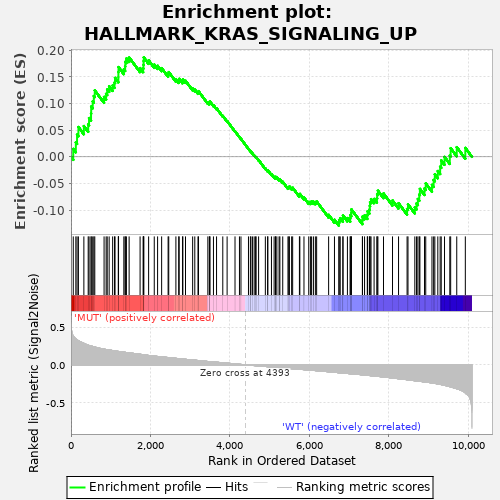
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_KRAS_SIGNALING_UP |
| Enrichment Score (ES) | 0.18679701 |
| Normalized Enrichment Score (NES) | 0.7532251 |
| Nominal p-value | 0.87323946 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SOX9 | na | 57 | 0.378 | 0.0149 | Yes | ||
| 2 | TNFRSF1B | na | 122 | 0.342 | 0.0271 | Yes | ||
| 3 | RABGAP1L | na | 153 | 0.328 | 0.0419 | Yes | ||
| 4 | KIF5C | na | 187 | 0.320 | 0.0560 | Yes | ||
| 5 | BPGM | na | 324 | 0.284 | 0.0578 | Yes | ||
| 6 | IL7R | na | 433 | 0.259 | 0.0611 | Yes | ||
| 7 | ADAM8 | na | 455 | 0.255 | 0.0729 | Yes | ||
| 8 | BTBD3 | na | 505 | 0.248 | 0.0814 | Yes | ||
| 9 | KLF4 | na | 509 | 0.247 | 0.0946 | Yes | ||
| 10 | PEG3 | na | 547 | 0.242 | 0.1041 | Yes | ||
| 11 | SPRY2 | na | 575 | 0.238 | 0.1143 | Yes | ||
| 12 | CPE | na | 599 | 0.234 | 0.1248 | Yes | ||
| 13 | ITGA2 | na | 833 | 0.206 | 0.1126 | Yes | ||
| 14 | HDAC9 | na | 885 | 0.202 | 0.1185 | Yes | ||
| 15 | SPARCL1 | na | 912 | 0.200 | 0.1267 | Yes | ||
| 16 | PTPRR | na | 961 | 0.196 | 0.1326 | Yes | ||
| 17 | ALDH1A3 | na | 1049 | 0.188 | 0.1341 | Yes | ||
| 18 | CD37 | na | 1096 | 0.185 | 0.1395 | Yes | ||
| 19 | FUCA1 | na | 1109 | 0.184 | 0.1484 | Yes | ||
| 20 | MAP3K1 | na | 1193 | 0.177 | 0.1496 | Yes | ||
| 21 | EVI5 | na | 1194 | 0.177 | 0.1593 | Yes | ||
| 22 | MMP10 | na | 1197 | 0.176 | 0.1687 | Yes | ||
| 23 | RGS16 | na | 1331 | 0.166 | 0.1643 | Yes | ||
| 24 | PLAT | na | 1364 | 0.165 | 0.1701 | Yes | ||
| 25 | JUP | na | 1368 | 0.164 | 0.1787 | Yes | ||
| 26 | KCNN4 | na | 1396 | 0.162 | 0.1848 | Yes | ||
| 27 | RBM4 | na | 1464 | 0.158 | 0.1867 | Yes | ||
| 28 | PPBP | na | 1740 | 0.139 | 0.1667 | Yes | ||
| 29 | BTC | na | 1817 | 0.134 | 0.1663 | Yes | ||
| 30 | PPP1R15A | na | 1825 | 0.134 | 0.1729 | Yes | ||
| 31 | DNMBP | na | 1829 | 0.133 | 0.1799 | Yes | ||
| 32 | TRAF1 | na | 1833 | 0.133 | 0.1868 | Yes | ||
| 33 | BIRC3 | na | 1955 | 0.123 | 0.1813 | No | ||
| 34 | CSF2 | na | 2097 | 0.114 | 0.1734 | No | ||
| 35 | MMP11 | na | 2181 | 0.110 | 0.1710 | No | ||
| 36 | FGF9 | na | 2283 | 0.105 | 0.1666 | No | ||
| 37 | SPON1 | na | 2446 | 0.095 | 0.1555 | No | ||
| 38 | RBP4 | na | 2462 | 0.094 | 0.1592 | No | ||
| 39 | ITGBL1 | na | 2642 | 0.085 | 0.1458 | No | ||
| 40 | CA2 | na | 2713 | 0.081 | 0.1432 | No | ||
| 41 | ADAM17 | na | 2724 | 0.080 | 0.1465 | No | ||
| 42 | SATB1 | na | 2808 | 0.076 | 0.1423 | No | ||
| 43 | AKT2 | na | 2818 | 0.075 | 0.1455 | No | ||
| 44 | CFH | na | 2884 | 0.072 | 0.1429 | No | ||
| 45 | GNG11 | na | 3064 | 0.063 | 0.1284 | No | ||
| 46 | DUSP6 | na | 3118 | 0.060 | 0.1263 | No | ||
| 47 | USH1C | na | 3199 | 0.056 | 0.1214 | No | ||
| 48 | CXCL10 | na | 3211 | 0.056 | 0.1233 | No | ||
| 49 | TNNT2 | na | 3447 | 0.043 | 0.1021 | No | ||
| 50 | NR0B2 | na | 3485 | 0.042 | 0.1006 | No | ||
| 51 | SEMA3B | na | 3496 | 0.041 | 0.1019 | No | ||
| 52 | F2RL1 | na | 3497 | 0.041 | 0.1041 | No | ||
| 53 | ETV5 | na | 3588 | 0.037 | 0.0971 | No | ||
| 54 | NRP1 | na | 3666 | 0.034 | 0.0912 | No | ||
| 55 | EPHB2 | na | 3823 | 0.026 | 0.0769 | No | ||
| 56 | SLPI | na | 3932 | 0.022 | 0.0673 | No | ||
| 57 | ANXA10 | na | 4131 | 0.011 | 0.0480 | No | ||
| 58 | ETV1 | na | 4243 | 0.006 | 0.0372 | No | ||
| 59 | FLT4 | na | 4281 | 0.005 | 0.0337 | No | ||
| 60 | GABRA3 | na | 4470 | -0.003 | 0.0150 | No | ||
| 61 | PRRX1 | na | 4518 | -0.006 | 0.0106 | No | ||
| 62 | NR1H4 | na | 4532 | -0.006 | 0.0097 | No | ||
| 63 | GALNT3 | na | 4573 | -0.009 | 0.0061 | No | ||
| 64 | MMD | na | 4614 | -0.010 | 0.0027 | No | ||
| 65 | CMKLR1 | na | 4649 | -0.012 | -0.0001 | No | ||
| 66 | PRKG2 | na | 4662 | -0.012 | -0.0006 | No | ||
| 67 | TPH1 | na | 4725 | -0.015 | -0.0060 | No | ||
| 68 | ITGB2 | na | 4895 | -0.022 | -0.0218 | No | ||
| 69 | IL2RG | na | 4952 | -0.025 | -0.0260 | No | ||
| 70 | CROT | na | 4958 | -0.025 | -0.0252 | No | ||
| 71 | GADD45G | na | 5049 | -0.028 | -0.0327 | No | ||
| 72 | ADAMDEC1 | na | 5116 | -0.031 | -0.0376 | No | ||
| 73 | PLAUR | na | 5152 | -0.033 | -0.0393 | No | ||
| 74 | CBL | na | 5153 | -0.033 | -0.0375 | No | ||
| 75 | DCBLD2 | na | 5185 | -0.035 | -0.0388 | No | ||
| 76 | STRN | na | 5241 | -0.037 | -0.0423 | No | ||
| 77 | IGFBP3 | na | 5261 | -0.038 | -0.0421 | No | ||
| 78 | PSMB8 | na | 5333 | -0.041 | -0.0470 | No | ||
| 79 | SNAP91 | na | 5465 | -0.047 | -0.0576 | No | ||
| 80 | ALDH1A2 | na | 5485 | -0.048 | -0.0569 | No | ||
| 81 | BMP2 | na | 5501 | -0.049 | -0.0557 | No | ||
| 82 | CBR4 | na | 5557 | -0.051 | -0.0585 | No | ||
| 83 | PLAU | na | 5583 | -0.052 | -0.0581 | No | ||
| 84 | IL1RL2 | na | 5753 | -0.059 | -0.0719 | No | ||
| 85 | PCP4 | na | 5764 | -0.059 | -0.0696 | No | ||
| 86 | ID2 | na | 5868 | -0.064 | -0.0765 | No | ||
| 87 | IL10RA | na | 5988 | -0.069 | -0.0847 | No | ||
| 88 | HSD11B1 | na | 6030 | -0.071 | -0.0849 | No | ||
| 89 | CSF2RA | na | 6060 | -0.073 | -0.0839 | No | ||
| 90 | GFPT2 | na | 6106 | -0.074 | -0.0843 | No | ||
| 91 | EREG | na | 6155 | -0.076 | -0.0850 | No | ||
| 92 | TRIB2 | na | 6187 | -0.078 | -0.0838 | No | ||
| 93 | LAPTM5 | na | 6489 | -0.092 | -0.1091 | No | ||
| 94 | LCP1 | na | 6636 | -0.100 | -0.1183 | No | ||
| 95 | ABCB1 | na | 6746 | -0.105 | -0.1235 | No | ||
| 96 | G0S2 | na | 6755 | -0.105 | -0.1186 | No | ||
| 97 | CXCR4 | na | 6781 | -0.106 | -0.1153 | No | ||
| 98 | ENG | na | 6849 | -0.109 | -0.1161 | No | ||
| 99 | SCN1B | na | 6850 | -0.109 | -0.1102 | No | ||
| 100 | ACE | na | 6962 | -0.115 | -0.1151 | No | ||
| 101 | LIF | na | 7032 | -0.118 | -0.1156 | No | ||
| 102 | ETV4 | na | 7036 | -0.118 | -0.1094 | No | ||
| 103 | AMMECR1 | na | 7060 | -0.120 | -0.1052 | No | ||
| 104 | SNAP25 | na | 7062 | -0.120 | -0.0988 | No | ||
| 105 | RELN | na | 7339 | -0.132 | -0.1193 | No | ||
| 106 | MYCN | na | 7342 | -0.133 | -0.1123 | No | ||
| 107 | EMP1 | na | 7394 | -0.135 | -0.1100 | No | ||
| 108 | MAP4K1 | na | 7463 | -0.139 | -0.1093 | No | ||
| 109 | TFPI | na | 7470 | -0.139 | -0.1023 | No | ||
| 110 | TRIB1 | na | 7516 | -0.142 | -0.0991 | No | ||
| 111 | F13A1 | na | 7526 | -0.142 | -0.0923 | No | ||
| 112 | INHBA | na | 7533 | -0.143 | -0.0851 | No | ||
| 113 | MAP7 | na | 7557 | -0.145 | -0.0795 | No | ||
| 114 | GYPC | na | 7634 | -0.148 | -0.0791 | No | ||
| 115 | C3AR1 | na | 7705 | -0.152 | -0.0779 | No | ||
| 116 | SERPINA3 | na | 7712 | -0.152 | -0.0702 | No | ||
| 117 | ETS1 | na | 7730 | -0.153 | -0.0636 | No | ||
| 118 | AKAP12 | na | 7871 | -0.161 | -0.0688 | No | ||
| 119 | MMP9 | na | 8098 | -0.175 | -0.0820 | No | ||
| 120 | CCND2 | na | 8250 | -0.184 | -0.0872 | No | ||
| 121 | APOD | na | 8465 | -0.198 | -0.0979 | No | ||
| 122 | CCL20 | na | 8488 | -0.199 | -0.0893 | No | ||
| 123 | TNFAIP3 | na | 8660 | -0.211 | -0.0950 | No | ||
| 124 | IL1B | na | 8703 | -0.214 | -0.0876 | No | ||
| 125 | PTGS2 | na | 8737 | -0.216 | -0.0791 | No | ||
| 126 | PIGR | na | 8773 | -0.218 | -0.0708 | No | ||
| 127 | SPP1 | na | 8788 | -0.219 | -0.0602 | No | ||
| 128 | GPNMB | na | 8904 | -0.227 | -0.0594 | No | ||
| 129 | PRDM1 | na | 8935 | -0.229 | -0.0500 | No | ||
| 130 | WNT7A | na | 9092 | -0.241 | -0.0526 | No | ||
| 131 | FCER1G | na | 9134 | -0.243 | -0.0434 | No | ||
| 132 | EPB41L3 | na | 9164 | -0.246 | -0.0329 | No | ||
| 133 | IGF2 | na | 9239 | -0.252 | -0.0266 | No | ||
| 134 | DOCK2 | na | 9298 | -0.259 | -0.0184 | No | ||
| 135 | PECAM1 | na | 9325 | -0.262 | -0.0067 | No | ||
| 136 | CTSS | na | 9408 | -0.272 | -0.0001 | No | ||
| 137 | ARG1 | na | 9541 | -0.289 | 0.0024 | No | ||
| 138 | GLRX | na | 9563 | -0.292 | 0.0161 | No | ||
| 139 | GPRC5B | na | 9716 | -0.316 | 0.0181 | No | ||
| 140 | LY96 | na | 9932 | -0.372 | 0.0168 | No |