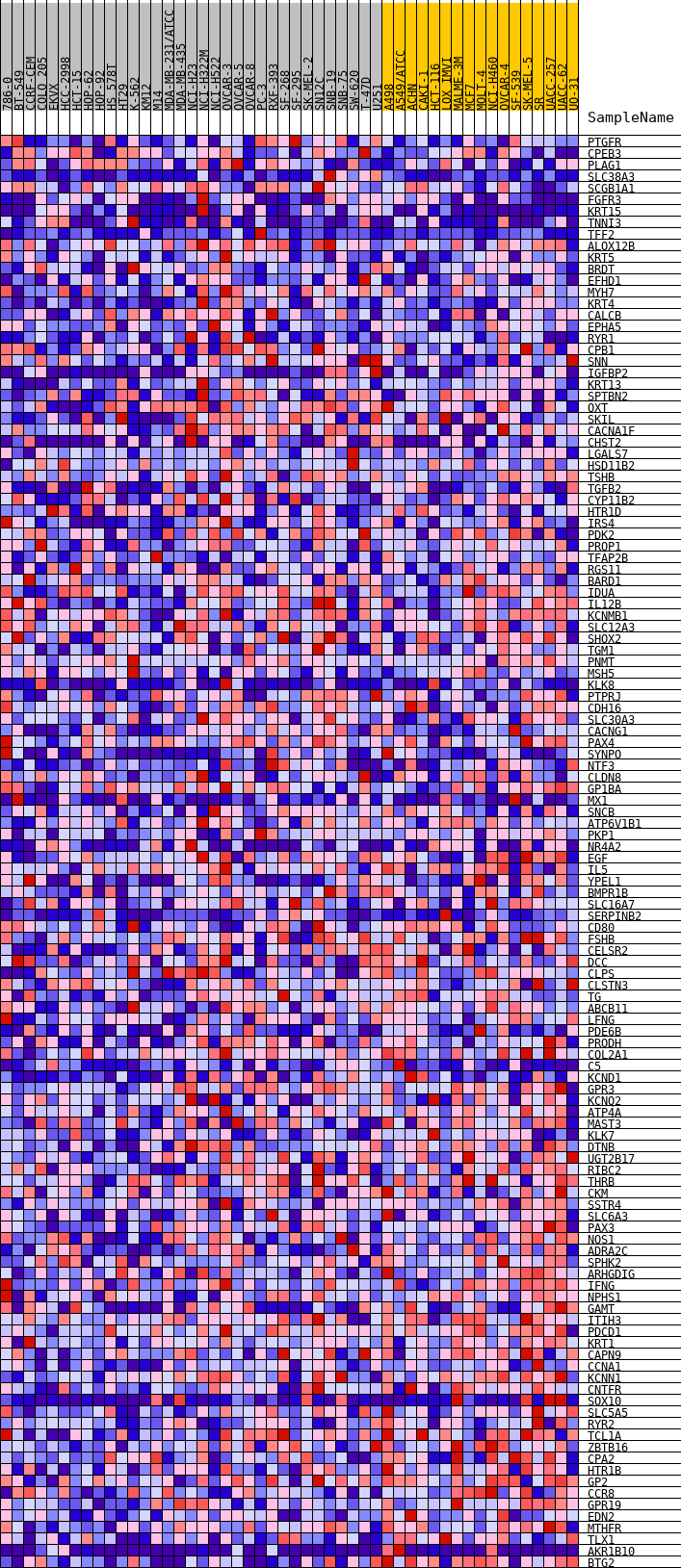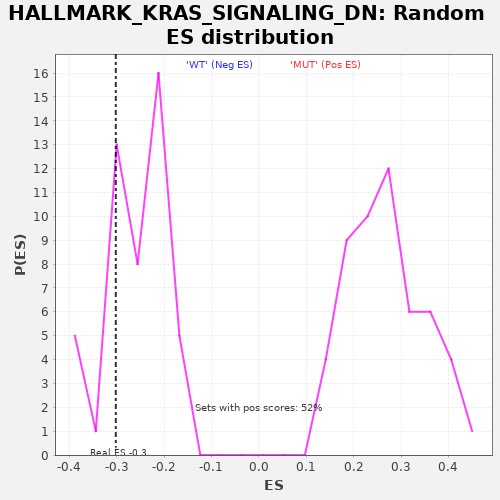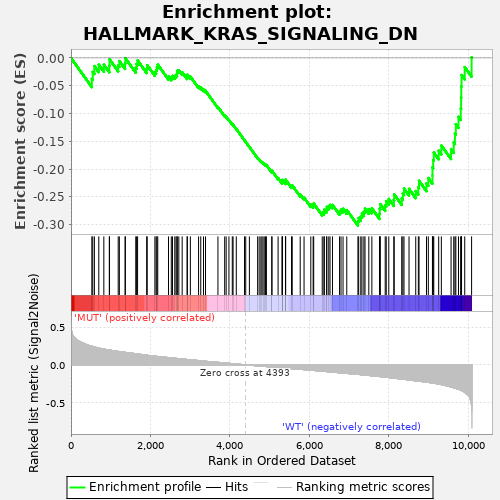
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_KRAS_SIGNALING_DN |
| Enrichment Score (ES) | -0.3017706 |
| Normalized Enrichment Score (NES) | -1.1620005 |
| Nominal p-value | 0.27083334 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.97 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PTGFR | na | 525 | 0.246 | -0.0376 | No | ||
| 2 | CPEB3 | na | 548 | 0.242 | -0.0250 | No | ||
| 3 | PLAG1 | na | 589 | 0.236 | -0.0146 | No | ||
| 4 | SLC38A3 | na | 700 | 0.221 | -0.0122 | No | ||
| 5 | SCGB1A1 | na | 827 | 0.207 | -0.0121 | No | ||
| 6 | FGFR3 | na | 965 | 0.196 | -0.0139 | No | ||
| 7 | KRT15 | na | 970 | 0.195 | -0.0024 | No | ||
| 8 | TNNI3 | na | 1189 | 0.177 | -0.0135 | No | ||
| 9 | TFF2 | na | 1218 | 0.175 | -0.0056 | No | ||
| 10 | ALOX12B | na | 1362 | 0.165 | -0.0099 | No | ||
| 11 | KRT5 | na | 1369 | 0.164 | -0.0005 | No | ||
| 12 | BRDT | na | 1631 | 0.147 | -0.0177 | No | ||
| 13 | EFHD1 | na | 1652 | 0.145 | -0.0108 | No | ||
| 14 | MYH7 | na | 1677 | 0.143 | -0.0045 | No | ||
| 15 | KRT4 | na | 1906 | 0.128 | -0.0195 | No | ||
| 16 | CALCB | na | 1919 | 0.127 | -0.0130 | No | ||
| 17 | EPHA5 | na | 2113 | 0.113 | -0.0254 | No | ||
| 18 | RYR1 | na | 2144 | 0.112 | -0.0216 | No | ||
| 19 | CPB1 | na | 2163 | 0.111 | -0.0167 | No | ||
| 20 | SNN | na | 2185 | 0.110 | -0.0121 | No | ||
| 21 | IGFBP2 | na | 2456 | 0.095 | -0.0333 | No | ||
| 22 | KRT13 | na | 2526 | 0.091 | -0.0347 | No | ||
| 23 | SPTBN2 | na | 2557 | 0.090 | -0.0322 | No | ||
| 24 | OXT | na | 2614 | 0.087 | -0.0325 | No | ||
| 25 | SKIL | na | 2646 | 0.085 | -0.0305 | No | ||
| 26 | CACNA1F | na | 2671 | 0.083 | -0.0278 | No | ||
| 27 | CHST2 | na | 2676 | 0.083 | -0.0231 | No | ||
| 28 | LGALS7 | na | 2716 | 0.081 | -0.0221 | No | ||
| 29 | HSD11B2 | na | 2800 | 0.076 | -0.0258 | No | ||
| 30 | TSHB | na | 2924 | 0.070 | -0.0338 | No | ||
| 31 | TGFB2 | na | 2931 | 0.069 | -0.0302 | No | ||
| 32 | CYP11B2 | na | 3006 | 0.066 | -0.0336 | No | ||
| 33 | HTR1D | na | 3213 | 0.056 | -0.0509 | No | ||
| 34 | IRS4 | na | 3268 | 0.053 | -0.0530 | No | ||
| 35 | PDK2 | na | 3337 | 0.049 | -0.0569 | No | ||
| 36 | PROP1 | na | 3388 | 0.046 | -0.0590 | No | ||
| 37 | TFAP2B | na | 3701 | 0.032 | -0.0884 | No | ||
| 38 | RGS11 | na | 3871 | 0.024 | -0.1038 | No | ||
| 39 | BARD1 | na | 3908 | 0.022 | -0.1061 | No | ||
| 40 | IDUA | na | 3978 | 0.019 | -0.1118 | No | ||
| 41 | IL12B | na | 4062 | 0.015 | -0.1192 | No | ||
| 42 | KCNMB1 | na | 4082 | 0.013 | -0.1203 | No | ||
| 43 | SLC12A3 | na | 4163 | 0.010 | -0.1277 | No | ||
| 44 | SHOX2 | na | 4370 | 0.001 | -0.1483 | No | ||
| 45 | TGM1 | na | 4401 | -0.001 | -0.1513 | No | ||
| 46 | PNMT | na | 4493 | -0.005 | -0.1601 | No | ||
| 47 | MSH5 | na | 4701 | -0.014 | -0.1800 | No | ||
| 48 | KLK8 | na | 4739 | -0.016 | -0.1827 | No | ||
| 49 | PTPRJ | na | 4782 | -0.018 | -0.1858 | No | ||
| 50 | CDH16 | na | 4810 | -0.019 | -0.1874 | No | ||
| 51 | SLC30A3 | na | 4839 | -0.020 | -0.1889 | No | ||
| 52 | CACNG1 | na | 4881 | -0.022 | -0.1917 | No | ||
| 53 | PAX4 | na | 4898 | -0.022 | -0.1919 | No | ||
| 54 | SYNPO | na | 4922 | -0.023 | -0.1928 | No | ||
| 55 | NTF3 | na | 5059 | -0.029 | -0.2047 | No | ||
| 56 | CLDN8 | na | 5060 | -0.029 | -0.2029 | No | ||
| 57 | GP1BA | na | 5216 | -0.036 | -0.2163 | No | ||
| 58 | MX1 | na | 5317 | -0.040 | -0.2238 | No | ||
| 59 | SNCB | na | 5322 | -0.041 | -0.2218 | No | ||
| 60 | ATP6V1B1 | na | 5325 | -0.041 | -0.2195 | No | ||
| 61 | PKP1 | na | 5402 | -0.044 | -0.2244 | No | ||
| 62 | NR4A2 | na | 5404 | -0.044 | -0.2218 | No | ||
| 63 | EGF | na | 5406 | -0.044 | -0.2192 | No | ||
| 64 | IL5 | na | 5554 | -0.051 | -0.2308 | No | ||
| 65 | YPEL1 | na | 5572 | -0.052 | -0.2293 | No | ||
| 66 | BMPR1B | na | 5773 | -0.060 | -0.2457 | No | ||
| 67 | SLC16A7 | na | 5870 | -0.064 | -0.2515 | No | ||
| 68 | SERPINB2 | na | 6040 | -0.072 | -0.2640 | No | ||
| 69 | CD80 | na | 6099 | -0.074 | -0.2653 | No | ||
| 70 | FSHB | na | 6113 | -0.075 | -0.2621 | No | ||
| 71 | CELSR2 | na | 6328 | -0.085 | -0.2783 | No | ||
| 72 | DCC | na | 6370 | -0.086 | -0.2772 | No | ||
| 73 | CLPS | na | 6381 | -0.087 | -0.2728 | No | ||
| 74 | CLSTN3 | na | 6438 | -0.090 | -0.2730 | No | ||
| 75 | TG | na | 6444 | -0.090 | -0.2680 | No | ||
| 76 | ABCB11 | na | 6493 | -0.092 | -0.2671 | No | ||
| 77 | LFNG | na | 6524 | -0.094 | -0.2644 | No | ||
| 78 | PDE6B | na | 6586 | -0.097 | -0.2646 | No | ||
| 79 | PRODH | na | 6764 | -0.106 | -0.2759 | No | ||
| 80 | COL2A1 | na | 6801 | -0.107 | -0.2730 | No | ||
| 81 | C5 | na | 6852 | -0.109 | -0.2713 | No | ||
| 82 | KCND1 | na | 6946 | -0.114 | -0.2737 | No | ||
| 83 | GPR3 | na | 7227 | -0.127 | -0.2940 | Yes | ||
| 84 | KCNQ2 | na | 7246 | -0.128 | -0.2880 | Yes | ||
| 85 | ATP4A | na | 7299 | -0.131 | -0.2853 | Yes | ||
| 86 | MAST3 | na | 7328 | -0.132 | -0.2800 | Yes | ||
| 87 | KLK7 | na | 7377 | -0.134 | -0.2766 | Yes | ||
| 88 | DTNB | na | 7405 | -0.136 | -0.2710 | Yes | ||
| 89 | UGT2B17 | na | 7501 | -0.141 | -0.2720 | Yes | ||
| 90 | RIBC2 | na | 7578 | -0.146 | -0.2707 | Yes | ||
| 91 | THRB | na | 7770 | -0.155 | -0.2803 | Yes | ||
| 92 | CKM | na | 7776 | -0.156 | -0.2713 | Yes | ||
| 93 | SSTR4 | na | 7791 | -0.157 | -0.2631 | Yes | ||
| 94 | SLC6A3 | na | 7914 | -0.163 | -0.2654 | Yes | ||
| 95 | PAX3 | na | 7940 | -0.165 | -0.2578 | Yes | ||
| 96 | NOS1 | na | 8004 | -0.169 | -0.2538 | Yes | ||
| 97 | ADRA2C | na | 8131 | -0.177 | -0.2556 | Yes | ||
| 98 | SPHK2 | na | 8140 | -0.177 | -0.2456 | Yes | ||
| 99 | ARHGDIG | na | 8330 | -0.188 | -0.2531 | Yes | ||
| 100 | IFNG | na | 8358 | -0.191 | -0.2441 | Yes | ||
| 101 | NPHS1 | na | 8387 | -0.192 | -0.2352 | Yes | ||
| 102 | GAMT | na | 8514 | -0.200 | -0.2356 | Yes | ||
| 103 | ITIH3 | na | 8683 | -0.212 | -0.2395 | Yes | ||
| 104 | PDCD1 | na | 8749 | -0.216 | -0.2328 | Yes | ||
| 105 | KRT1 | na | 8766 | -0.218 | -0.2211 | Yes | ||
| 106 | CAPN9 | na | 8954 | -0.230 | -0.2258 | Yes | ||
| 107 | CCNA1 | na | 9000 | -0.233 | -0.2161 | Yes | ||
| 108 | KCNN1 | na | 9104 | -0.241 | -0.2116 | Yes | ||
| 109 | CNTFR | na | 9105 | -0.242 | -0.1969 | Yes | ||
| 110 | SOX10 | na | 9123 | -0.243 | -0.1838 | Yes | ||
| 111 | SLC5A5 | na | 9137 | -0.243 | -0.1702 | Yes | ||
| 112 | RYR2 | na | 9261 | -0.255 | -0.1670 | Yes | ||
| 113 | TCL1A | na | 9328 | -0.262 | -0.1576 | Yes | ||
| 114 | ZBTB16 | na | 9573 | -0.293 | -0.1641 | Yes | ||
| 115 | CPA2 | na | 9640 | -0.305 | -0.1521 | Yes | ||
| 116 | HTR1B | na | 9671 | -0.310 | -0.1362 | Yes | ||
| 117 | GP2 | na | 9693 | -0.313 | -0.1192 | Yes | ||
| 118 | CCR8 | na | 9760 | -0.324 | -0.1061 | Yes | ||
| 119 | GPR19 | na | 9817 | -0.335 | -0.0912 | Yes | ||
| 120 | EDN2 | na | 9826 | -0.338 | -0.0714 | Yes | ||
| 121 | MTHFR | na | 9830 | -0.339 | -0.0510 | Yes | ||
| 122 | TLX1 | na | 9837 | -0.340 | -0.0308 | Yes | ||
| 123 | AKR1B10 | na | 9917 | -0.365 | -0.0165 | Yes | ||
| 124 | BTG2 | na | 10090 | -0.567 | 0.0009 | Yes |