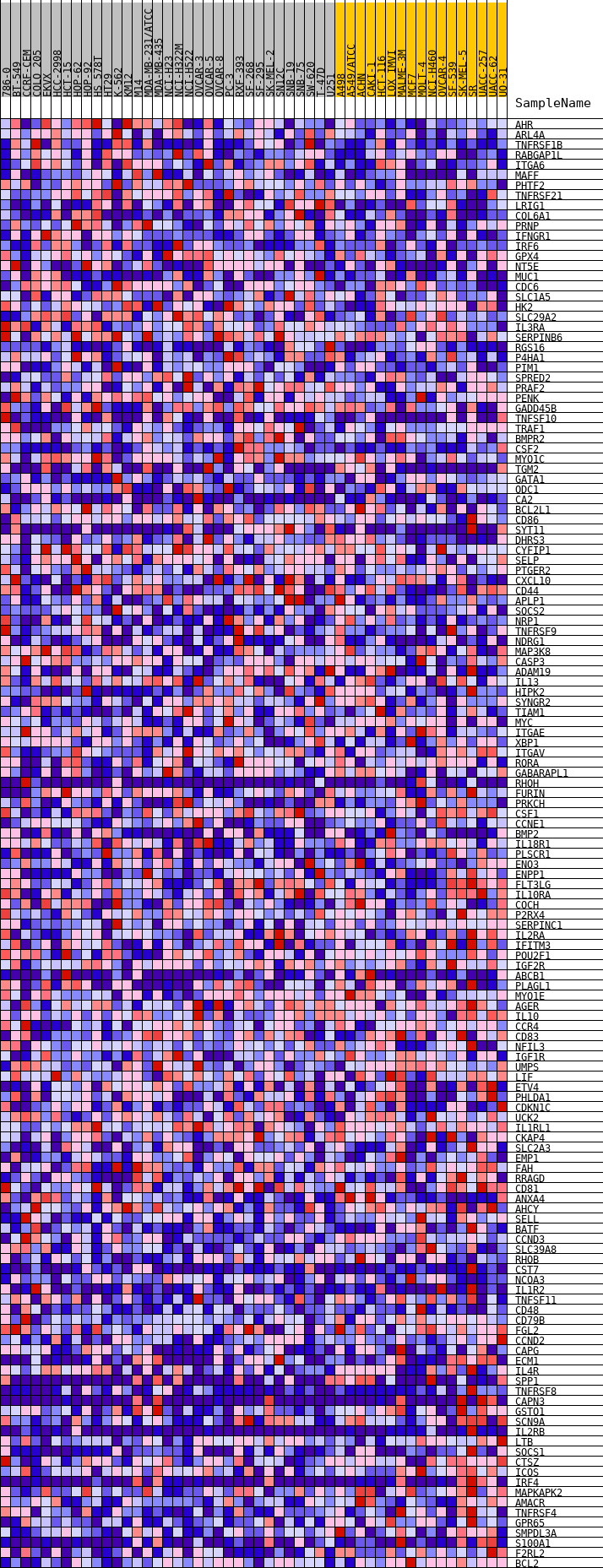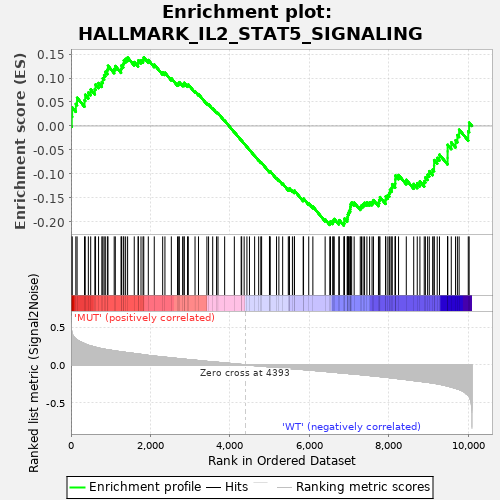
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_IL2_STAT5_SIGNALING |
| Enrichment Score (ES) | -0.20764275 |
| Normalized Enrichment Score (NES) | -0.9271519 |
| Nominal p-value | 0.5483871 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AHR | na | 19 | 0.448 | 0.0197 | No | ||
| 2 | ARL4A | na | 29 | 0.414 | 0.0388 | No | ||
| 3 | TNFRSF1B | na | 122 | 0.342 | 0.0461 | No | ||
| 4 | RABGAP1L | na | 153 | 0.328 | 0.0589 | No | ||
| 5 | ITGA6 | na | 340 | 0.279 | 0.0537 | No | ||
| 6 | MAFF | na | 358 | 0.274 | 0.0652 | No | ||
| 7 | PHTF2 | na | 437 | 0.258 | 0.0698 | No | ||
| 8 | TNFRSF21 | na | 496 | 0.250 | 0.0760 | No | ||
| 9 | LRIG1 | na | 601 | 0.234 | 0.0769 | No | ||
| 10 | COL6A1 | na | 618 | 0.232 | 0.0864 | No | ||
| 11 | PRNP | na | 694 | 0.221 | 0.0896 | No | ||
| 12 | IFNGR1 | na | 777 | 0.212 | 0.0916 | No | ||
| 13 | IRF6 | na | 801 | 0.209 | 0.0994 | No | ||
| 14 | GPX4 | na | 834 | 0.206 | 0.1061 | No | ||
| 15 | NT5E | na | 864 | 0.203 | 0.1130 | No | ||
| 16 | MUC1 | na | 916 | 0.199 | 0.1175 | No | ||
| 17 | CDC6 | na | 931 | 0.198 | 0.1256 | No | ||
| 18 | SLC1A5 | na | 1088 | 0.186 | 0.1189 | No | ||
| 19 | HK2 | na | 1116 | 0.183 | 0.1250 | No | ||
| 20 | SLC29A2 | na | 1261 | 0.172 | 0.1189 | No | ||
| 21 | IL3RA | na | 1271 | 0.171 | 0.1262 | No | ||
| 22 | SERPINB6 | na | 1316 | 0.167 | 0.1299 | No | ||
| 23 | RGS16 | na | 1331 | 0.166 | 0.1365 | No | ||
| 24 | P4HA1 | na | 1376 | 0.163 | 0.1399 | No | ||
| 25 | PIM1 | na | 1426 | 0.161 | 0.1428 | No | ||
| 26 | SPRED2 | na | 1593 | 0.149 | 0.1333 | No | ||
| 27 | PRAF2 | na | 1690 | 0.143 | 0.1305 | No | ||
| 28 | PENK | na | 1697 | 0.142 | 0.1368 | No | ||
| 29 | GADD45B | na | 1763 | 0.137 | 0.1369 | No | ||
| 30 | TNFSF10 | na | 1814 | 0.134 | 0.1383 | No | ||
| 31 | TRAF1 | na | 1833 | 0.133 | 0.1429 | No | ||
| 32 | BMPR2 | na | 1947 | 0.123 | 0.1375 | No | ||
| 33 | CSF2 | na | 2097 | 0.114 | 0.1281 | No | ||
| 34 | MYO1C | na | 2309 | 0.103 | 0.1119 | No | ||
| 35 | TGM2 | na | 2366 | 0.100 | 0.1111 | No | ||
| 36 | GATA1 | na | 2527 | 0.091 | 0.0994 | No | ||
| 37 | ODC1 | na | 2683 | 0.083 | 0.0878 | No | ||
| 38 | CA2 | na | 2713 | 0.081 | 0.0888 | No | ||
| 39 | BCL2L1 | na | 2728 | 0.080 | 0.0913 | No | ||
| 40 | CD86 | na | 2811 | 0.076 | 0.0867 | No | ||
| 41 | SYT11 | na | 2848 | 0.074 | 0.0866 | No | ||
| 42 | DHRS3 | na | 2850 | 0.074 | 0.0900 | No | ||
| 43 | CYFIP1 | na | 2935 | 0.069 | 0.0849 | No | ||
| 44 | SELP | na | 2954 | 0.068 | 0.0864 | No | ||
| 45 | PTGER2 | na | 3124 | 0.060 | 0.0723 | No | ||
| 46 | CXCL10 | na | 3211 | 0.056 | 0.0664 | No | ||
| 47 | CD44 | na | 3423 | 0.045 | 0.0474 | No | ||
| 48 | APLP1 | na | 3465 | 0.042 | 0.0453 | No | ||
| 49 | SOCS2 | na | 3569 | 0.038 | 0.0368 | No | ||
| 50 | NRP1 | na | 3666 | 0.034 | 0.0288 | No | ||
| 51 | TNFRSF9 | na | 3702 | 0.032 | 0.0268 | No | ||
| 52 | NDRG1 | na | 3870 | 0.024 | 0.0112 | No | ||
| 53 | MAP3K8 | na | 4114 | 0.012 | -0.0126 | No | ||
| 54 | CASP3 | na | 4286 | 0.004 | -0.0296 | No | ||
| 55 | ADAM19 | na | 4299 | 0.004 | -0.0306 | No | ||
| 56 | IL13 | na | 4361 | 0.001 | -0.0367 | No | ||
| 57 | HIPK2 | na | 4431 | -0.002 | -0.0435 | No | ||
| 58 | SYNGR2 | na | 4495 | -0.005 | -0.0496 | No | ||
| 59 | TIAM1 | na | 4625 | -0.011 | -0.0621 | No | ||
| 60 | MYC | na | 4727 | -0.015 | -0.0715 | No | ||
| 61 | ITGAE | na | 4781 | -0.018 | -0.0759 | No | ||
| 62 | XBP1 | na | 4801 | -0.019 | -0.0769 | No | ||
| 63 | ITGAV | na | 4999 | -0.026 | -0.0954 | No | ||
| 64 | RORA | na | 5001 | -0.026 | -0.0942 | No | ||
| 65 | GABARAPL1 | na | 5018 | -0.027 | -0.0945 | No | ||
| 66 | RHOH | na | 5177 | -0.034 | -0.1088 | No | ||
| 67 | FURIN | na | 5236 | -0.037 | -0.1128 | No | ||
| 68 | PRKCH | na | 5331 | -0.041 | -0.1203 | No | ||
| 69 | CSF1 | na | 5473 | -0.047 | -0.1321 | No | ||
| 70 | CCNE1 | na | 5497 | -0.049 | -0.1321 | No | ||
| 71 | BMP2 | na | 5501 | -0.049 | -0.1300 | No | ||
| 72 | IL18R1 | na | 5577 | -0.052 | -0.1351 | No | ||
| 73 | PLSCR1 | na | 5626 | -0.054 | -0.1373 | No | ||
| 74 | ENO3 | na | 5628 | -0.054 | -0.1347 | No | ||
| 75 | ENPP1 | na | 5848 | -0.063 | -0.1537 | No | ||
| 76 | FLT3LG | na | 5857 | -0.063 | -0.1514 | No | ||
| 77 | IL10RA | na | 5988 | -0.069 | -0.1611 | No | ||
| 78 | COCH | na | 6092 | -0.074 | -0.1679 | No | ||
| 79 | P2RX4 | na | 6401 | -0.088 | -0.1946 | No | ||
| 80 | SERPINC1 | na | 6518 | -0.093 | -0.2017 | No | ||
| 81 | IL2RA | na | 6536 | -0.095 | -0.1989 | No | ||
| 82 | IFITM3 | na | 6592 | -0.097 | -0.1997 | No | ||
| 83 | POU2F1 | na | 6608 | -0.098 | -0.1965 | No | ||
| 84 | IGF2R | na | 6630 | -0.099 | -0.1938 | No | ||
| 85 | ABCB1 | na | 6746 | -0.105 | -0.2003 | No | ||
| 86 | PLAGL1 | na | 6758 | -0.106 | -0.1963 | No | ||
| 87 | MYO1E | na | 6872 | -0.110 | -0.2023 | Yes | ||
| 88 | AGER | na | 6887 | -0.111 | -0.1984 | Yes | ||
| 89 | IL10 | na | 6888 | -0.111 | -0.1930 | Yes | ||
| 90 | CCR4 | na | 6956 | -0.114 | -0.1943 | Yes | ||
| 91 | CD83 | na | 6967 | -0.115 | -0.1897 | Yes | ||
| 92 | NFIL3 | na | 6973 | -0.115 | -0.1847 | Yes | ||
| 93 | IGF1R | na | 6989 | -0.116 | -0.1806 | Yes | ||
| 94 | UMPS | na | 7007 | -0.117 | -0.1766 | Yes | ||
| 95 | LIF | na | 7032 | -0.118 | -0.1733 | Yes | ||
| 96 | ETV4 | na | 7036 | -0.118 | -0.1679 | Yes | ||
| 97 | PHLDA1 | na | 7045 | -0.119 | -0.1630 | Yes | ||
| 98 | CDKN1C | na | 7069 | -0.120 | -0.1595 | Yes | ||
| 99 | UCK2 | na | 7132 | -0.123 | -0.1598 | Yes | ||
| 100 | IL1RL1 | na | 7286 | -0.130 | -0.1689 | Yes | ||
| 101 | CKAP4 | na | 7317 | -0.132 | -0.1655 | Yes | ||
| 102 | SLC2A3 | na | 7357 | -0.133 | -0.1630 | Yes | ||
| 103 | EMP1 | na | 7394 | -0.135 | -0.1601 | Yes | ||
| 104 | FAH | na | 7453 | -0.138 | -0.1593 | Yes | ||
| 105 | RRAGD | na | 7521 | -0.142 | -0.1591 | Yes | ||
| 106 | CD81 | na | 7583 | -0.146 | -0.1582 | Yes | ||
| 107 | ANXA4 | na | 7618 | -0.147 | -0.1545 | Yes | ||
| 108 | AHCY | na | 7744 | -0.154 | -0.1597 | Yes | ||
| 109 | SELL | na | 7763 | -0.155 | -0.1540 | Yes | ||
| 110 | BATF | na | 7782 | -0.156 | -0.1483 | Yes | ||
| 111 | CCND3 | na | 7924 | -0.164 | -0.1545 | Yes | ||
| 112 | SLC39A8 | na | 7927 | -0.164 | -0.1468 | Yes | ||
| 113 | RHOB | na | 7981 | -0.168 | -0.1440 | Yes | ||
| 114 | CST7 | na | 8017 | -0.170 | -0.1393 | Yes | ||
| 115 | NCOA3 | na | 8035 | -0.172 | -0.1327 | Yes | ||
| 116 | IL1R2 | na | 8079 | -0.174 | -0.1287 | Yes | ||
| 117 | TNFSF11 | na | 8090 | -0.174 | -0.1213 | Yes | ||
| 118 | CD48 | na | 8165 | -0.179 | -0.1201 | Yes | ||
| 119 | CD79B | na | 8168 | -0.179 | -0.1116 | Yes | ||
| 120 | FGL2 | na | 8171 | -0.179 | -0.1032 | Yes | ||
| 121 | CCND2 | na | 8250 | -0.184 | -0.1022 | Yes | ||
| 122 | CAPG | na | 8445 | -0.196 | -0.1122 | Yes | ||
| 123 | ECM1 | na | 8632 | -0.209 | -0.1208 | Yes | ||
| 124 | IL4R | na | 8721 | -0.215 | -0.1193 | Yes | ||
| 125 | SPP1 | na | 8788 | -0.219 | -0.1153 | Yes | ||
| 126 | TNFRSF8 | na | 8897 | -0.226 | -0.1153 | Yes | ||
| 127 | CAPN3 | na | 8926 | -0.228 | -0.1071 | Yes | ||
| 128 | GSTO1 | na | 8982 | -0.232 | -0.1014 | Yes | ||
| 129 | SCN9A | na | 9024 | -0.235 | -0.0942 | Yes | ||
| 130 | IL2RB | na | 9109 | -0.242 | -0.0910 | Yes | ||
| 131 | LTB | na | 9145 | -0.244 | -0.0827 | Yes | ||
| 132 | SOCS1 | na | 9147 | -0.244 | -0.0710 | Yes | ||
| 133 | CTSZ | na | 9224 | -0.251 | -0.0665 | Yes | ||
| 134 | ICOS | na | 9281 | -0.257 | -0.0598 | Yes | ||
| 135 | IRF4 | na | 9485 | -0.282 | -0.0665 | Yes | ||
| 136 | MAPKAPK2 | na | 9486 | -0.282 | -0.0529 | Yes | ||
| 137 | AMACR | na | 9487 | -0.283 | -0.0393 | Yes | ||
| 138 | TNFRSF4 | na | 9578 | -0.294 | -0.0341 | Yes | ||
| 139 | GPR65 | na | 9686 | -0.312 | -0.0298 | Yes | ||
| 140 | SMPDL3A | na | 9732 | -0.319 | -0.0189 | Yes | ||
| 141 | S100A1 | na | 9776 | -0.326 | -0.0075 | Yes | ||
| 142 | F2RL2 | na | 10006 | -0.404 | -0.0110 | Yes | ||
| 143 | BCL2 | na | 10028 | -0.420 | 0.0071 | Yes |