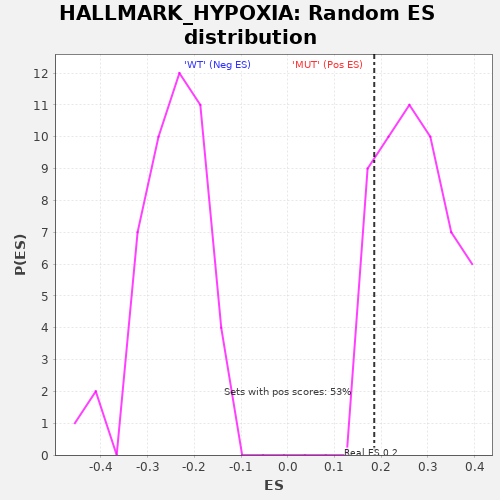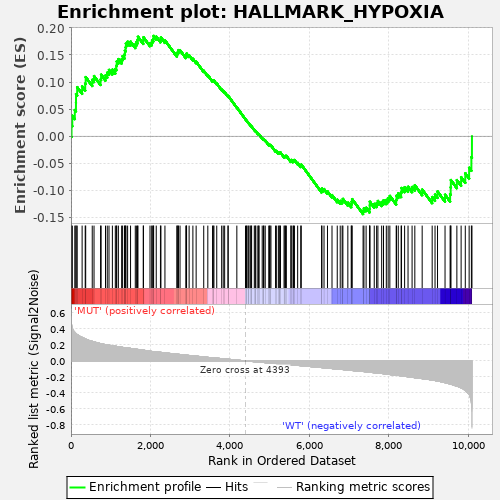
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_HYPOXIA |
| Enrichment Score (ES) | 0.18517184 |
| Normalized Enrichment Score (NES) | 0.6762265 |
| Nominal p-value | 0.8679245 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | F3 | na | 14 | 0.454 | 0.0202 | Yes | ||
| 2 | ADORA2B | na | 33 | 0.409 | 0.0377 | Yes | ||
| 3 | VHL | na | 96 | 0.356 | 0.0484 | Yes | ||
| 4 | TPST2 | na | 125 | 0.341 | 0.0618 | Yes | ||
| 5 | ENO1 | na | 127 | 0.340 | 0.0778 | Yes | ||
| 6 | GPC3 | na | 156 | 0.327 | 0.0905 | Yes | ||
| 7 | SAP30 | na | 280 | 0.294 | 0.0921 | Yes | ||
| 8 | MAFF | na | 358 | 0.274 | 0.0973 | Yes | ||
| 9 | STC2 | na | 370 | 0.272 | 0.1091 | Yes | ||
| 10 | TIPARP | na | 535 | 0.244 | 0.1042 | Yes | ||
| 11 | LOX | na | 580 | 0.237 | 0.1110 | Yes | ||
| 12 | CCNG2 | na | 745 | 0.216 | 0.1048 | Yes | ||
| 13 | ANXA2 | na | 758 | 0.214 | 0.1137 | Yes | ||
| 14 | DPYSL4 | na | 869 | 0.203 | 0.1123 | Yes | ||
| 15 | PDK1 | na | 913 | 0.200 | 0.1174 | Yes | ||
| 16 | PGK1 | na | 957 | 0.196 | 0.1224 | Yes | ||
| 17 | BNIP3L | na | 1042 | 0.189 | 0.1229 | Yes | ||
| 18 | HK2 | na | 1116 | 0.183 | 0.1243 | Yes | ||
| 19 | HSPA5 | na | 1143 | 0.181 | 0.1303 | Yes | ||
| 20 | CAV1 | na | 1152 | 0.180 | 0.1380 | Yes | ||
| 21 | MAP3K1 | na | 1193 | 0.177 | 0.1424 | Yes | ||
| 22 | ATP7A | na | 1276 | 0.171 | 0.1422 | Yes | ||
| 23 | CHST3 | na | 1300 | 0.169 | 0.1479 | Yes | ||
| 24 | TGFBI | na | 1348 | 0.165 | 0.1510 | Yes | ||
| 25 | SLC37A4 | na | 1358 | 0.165 | 0.1579 | Yes | ||
| 26 | P4HA1 | na | 1376 | 0.163 | 0.1640 | Yes | ||
| 27 | FOS | na | 1383 | 0.163 | 0.1711 | Yes | ||
| 28 | PIM1 | na | 1426 | 0.161 | 0.1745 | Yes | ||
| 29 | EGFR | na | 1500 | 0.156 | 0.1745 | Yes | ||
| 30 | SDC4 | na | 1619 | 0.147 | 0.1697 | Yes | ||
| 31 | ZNF292 | na | 1647 | 0.146 | 0.1739 | Yes | ||
| 32 | TPBG | na | 1676 | 0.143 | 0.1778 | Yes | ||
| 33 | TKTL1 | na | 1686 | 0.143 | 0.1837 | Yes | ||
| 34 | MYH9 | na | 1820 | 0.134 | 0.1767 | Yes | ||
| 35 | PPP1R15A | na | 1825 | 0.134 | 0.1826 | Yes | ||
| 36 | STC1 | na | 1990 | 0.120 | 0.1719 | Yes | ||
| 37 | EFNA1 | na | 2034 | 0.118 | 0.1731 | Yes | ||
| 38 | EXT1 | na | 2047 | 0.117 | 0.1775 | Yes | ||
| 39 | IL6 | na | 2075 | 0.115 | 0.1802 | Yes | ||
| 40 | PGAM2 | na | 2081 | 0.115 | 0.1852 | Yes | ||
| 41 | KDELR3 | na | 2147 | 0.112 | 0.1839 | No | ||
| 42 | PNRC1 | na | 2253 | 0.106 | 0.1784 | No | ||
| 43 | CSRP2 | na | 2265 | 0.106 | 0.1823 | No | ||
| 44 | TGM2 | na | 2366 | 0.100 | 0.1770 | No | ||
| 45 | CASP6 | na | 2667 | 0.084 | 0.1508 | No | ||
| 46 | CHST2 | na | 2676 | 0.083 | 0.1540 | No | ||
| 47 | SLC25A1 | na | 2696 | 0.082 | 0.1559 | No | ||
| 48 | GPC1 | na | 2706 | 0.081 | 0.1589 | No | ||
| 49 | PHKG1 | na | 2746 | 0.080 | 0.1587 | No | ||
| 50 | ADM | na | 2892 | 0.072 | 0.1476 | No | ||
| 51 | SLC6A6 | na | 2908 | 0.071 | 0.1494 | No | ||
| 52 | TGFB3 | na | 2911 | 0.071 | 0.1526 | No | ||
| 53 | ENO2 | na | 2975 | 0.067 | 0.1494 | No | ||
| 54 | XPNPEP1 | na | 3070 | 0.063 | 0.1429 | No | ||
| 55 | CP | na | 3152 | 0.059 | 0.1376 | No | ||
| 56 | GBE1 | na | 3343 | 0.049 | 0.1208 | No | ||
| 57 | ZFP36 | na | 3446 | 0.043 | 0.1126 | No | ||
| 58 | SERPINE1 | na | 3565 | 0.038 | 0.1025 | No | ||
| 59 | P4HA2 | na | 3583 | 0.037 | 0.1026 | No | ||
| 60 | UGP2 | na | 3600 | 0.037 | 0.1027 | No | ||
| 61 | PPFIA4 | na | 3671 | 0.033 | 0.0973 | No | ||
| 62 | CITED2 | na | 3795 | 0.028 | 0.0862 | No | ||
| 63 | TPI1 | na | 3834 | 0.026 | 0.0836 | No | ||
| 64 | NDRG1 | na | 3870 | 0.024 | 0.0812 | No | ||
| 65 | HDLBP | na | 3956 | 0.020 | 0.0736 | No | ||
| 66 | TES | na | 3957 | 0.020 | 0.0746 | No | ||
| 67 | SDC3 | na | 4177 | 0.009 | 0.0530 | No | ||
| 68 | PCK1 | na | 4395 | -0.000 | 0.0312 | No | ||
| 69 | IGFBP1 | na | 4421 | -0.002 | 0.0288 | No | ||
| 70 | PDGFB | na | 4432 | -0.002 | 0.0278 | No | ||
| 71 | PGF | na | 4464 | -0.003 | 0.0249 | No | ||
| 72 | SULT2B1 | na | 4482 | -0.004 | 0.0234 | No | ||
| 73 | HS3ST1 | na | 4516 | -0.006 | 0.0203 | No | ||
| 74 | LDHA | na | 4537 | -0.007 | 0.0186 | No | ||
| 75 | NR3C1 | na | 4549 | -0.007 | 0.0179 | No | ||
| 76 | KLF7 | na | 4617 | -0.010 | 0.0116 | No | ||
| 77 | SIAH2 | na | 4641 | -0.011 | 0.0098 | No | ||
| 78 | DDIT4 | na | 4655 | -0.012 | 0.0091 | No | ||
| 79 | IDS | na | 4706 | -0.015 | 0.0048 | No | ||
| 80 | GRHPR | na | 4726 | -0.015 | 0.0036 | No | ||
| 81 | NDST2 | na | 4737 | -0.016 | 0.0033 | No | ||
| 82 | JUN | na | 4816 | -0.019 | -0.0036 | No | ||
| 83 | PGM1 | na | 4849 | -0.021 | -0.0058 | No | ||
| 84 | PAM | na | 4858 | -0.021 | -0.0056 | No | ||
| 85 | GAA | na | 4892 | -0.022 | -0.0079 | No | ||
| 86 | MXI1 | na | 4981 | -0.026 | -0.0155 | No | ||
| 87 | RORA | na | 5001 | -0.026 | -0.0162 | No | ||
| 88 | PRKCA | na | 5013 | -0.027 | -0.0160 | No | ||
| 89 | DDIT3 | na | 5036 | -0.028 | -0.0169 | No | ||
| 90 | PLAUR | na | 5152 | -0.033 | -0.0269 | No | ||
| 91 | SLC2A1 | na | 5172 | -0.034 | -0.0272 | No | ||
| 92 | GCK | na | 5222 | -0.036 | -0.0304 | No | ||
| 93 | S100A4 | na | 5243 | -0.037 | -0.0307 | No | ||
| 94 | IGFBP3 | na | 5261 | -0.038 | -0.0306 | No | ||
| 95 | PFKL | na | 5276 | -0.039 | -0.0302 | No | ||
| 96 | FOSL2 | na | 5366 | -0.043 | -0.0371 | No | ||
| 97 | EFNA3 | na | 5386 | -0.044 | -0.0369 | No | ||
| 98 | PKP1 | na | 5402 | -0.044 | -0.0363 | No | ||
| 99 | GCNT2 | na | 5424 | -0.045 | -0.0363 | No | ||
| 100 | GPC4 | na | 5537 | -0.050 | -0.0452 | No | ||
| 101 | NDST1 | na | 5548 | -0.051 | -0.0438 | No | ||
| 102 | IER3 | na | 5592 | -0.053 | -0.0456 | No | ||
| 103 | HEXA | na | 5605 | -0.053 | -0.0443 | No | ||
| 104 | ENO3 | na | 5628 | -0.054 | -0.0439 | No | ||
| 105 | PFKP | na | 5709 | -0.057 | -0.0492 | No | ||
| 106 | DTNA | na | 5787 | -0.060 | -0.0541 | No | ||
| 107 | AMPD3 | na | 5799 | -0.061 | -0.0523 | No | ||
| 108 | BTG1 | na | 6312 | -0.084 | -0.0998 | No | ||
| 109 | ALDOB | na | 6320 | -0.085 | -0.0965 | No | ||
| 110 | PKLR | na | 6373 | -0.087 | -0.0976 | No | ||
| 111 | PDK3 | na | 6460 | -0.091 | -0.1020 | No | ||
| 112 | SELENBP1 | na | 6574 | -0.096 | -0.1088 | No | ||
| 113 | PPP1R3C | na | 6706 | -0.103 | -0.1171 | No | ||
| 114 | CXCR4 | na | 6781 | -0.106 | -0.1195 | No | ||
| 115 | CDKN1B | na | 6829 | -0.108 | -0.1191 | No | ||
| 116 | WSB1 | na | 6848 | -0.109 | -0.1157 | No | ||
| 117 | NFIL3 | na | 6973 | -0.115 | -0.1227 | No | ||
| 118 | ISG20 | na | 7058 | -0.120 | -0.1255 | No | ||
| 119 | CDKN1C | na | 7069 | -0.120 | -0.1208 | No | ||
| 120 | KIF5A | na | 7082 | -0.121 | -0.1163 | No | ||
| 121 | SLC2A3 | na | 7357 | -0.133 | -0.1375 | No | ||
| 122 | GYS1 | na | 7378 | -0.134 | -0.1331 | No | ||
| 123 | PFKFB3 | na | 7433 | -0.137 | -0.1320 | No | ||
| 124 | RRAGD | na | 7521 | -0.142 | -0.1341 | No | ||
| 125 | ATF3 | na | 7525 | -0.142 | -0.1276 | No | ||
| 126 | FBP1 | na | 7527 | -0.143 | -0.1209 | No | ||
| 127 | SRPX | na | 7638 | -0.148 | -0.1250 | No | ||
| 128 | GPI | na | 7698 | -0.151 | -0.1237 | No | ||
| 129 | ETS1 | na | 7730 | -0.153 | -0.1196 | No | ||
| 130 | SDC2 | na | 7825 | -0.158 | -0.1215 | No | ||
| 131 | AKAP12 | na | 7871 | -0.161 | -0.1184 | No | ||
| 132 | MT1E | na | 7942 | -0.165 | -0.1176 | No | ||
| 133 | HMOX1 | na | 7988 | -0.168 | -0.1141 | No | ||
| 134 | VLDLR | na | 8031 | -0.171 | -0.1102 | No | ||
| 135 | PYGM | na | 8191 | -0.180 | -0.1177 | No | ||
| 136 | LDHC | na | 8199 | -0.181 | -0.1098 | No | ||
| 137 | ALDOA | na | 8243 | -0.184 | -0.1054 | No | ||
| 138 | TPD52 | na | 8315 | -0.188 | -0.1036 | No | ||
| 139 | ILVBL | na | 8325 | -0.188 | -0.0956 | No | ||
| 140 | CA12 | na | 8403 | -0.193 | -0.0942 | No | ||
| 141 | HAS1 | na | 8489 | -0.199 | -0.0933 | No | ||
| 142 | SLC2A5 | na | 8591 | -0.206 | -0.0937 | No | ||
| 143 | TNFAIP3 | na | 8660 | -0.211 | -0.0905 | No | ||
| 144 | NEDD4L | na | 8845 | -0.223 | -0.0985 | No | ||
| 145 | ALDOC | na | 9097 | -0.241 | -0.1123 | No | ||
| 146 | DUSP1 | na | 9169 | -0.246 | -0.1077 | No | ||
| 147 | MIF | na | 9232 | -0.252 | -0.1020 | No | ||
| 148 | SCARB1 | na | 9421 | -0.274 | -0.1079 | No | ||
| 149 | IRS2 | na | 9549 | -0.290 | -0.1070 | No | ||
| 150 | GLRX | na | 9563 | -0.292 | -0.0944 | No | ||
| 151 | GALK1 | na | 9568 | -0.293 | -0.0809 | No | ||
| 152 | BRS3 | na | 9719 | -0.316 | -0.0810 | No | ||
| 153 | EDN2 | na | 9826 | -0.338 | -0.0756 | No | ||
| 154 | INHA | na | 9931 | -0.372 | -0.0685 | No | ||
| 155 | BCL2 | na | 10028 | -0.420 | -0.0582 | No | ||
| 156 | LALBA | na | 10085 | -0.531 | -0.0387 | No | ||
| 157 | CDKN1A | na | 10099 | -0.843 | 0.0000 | No |