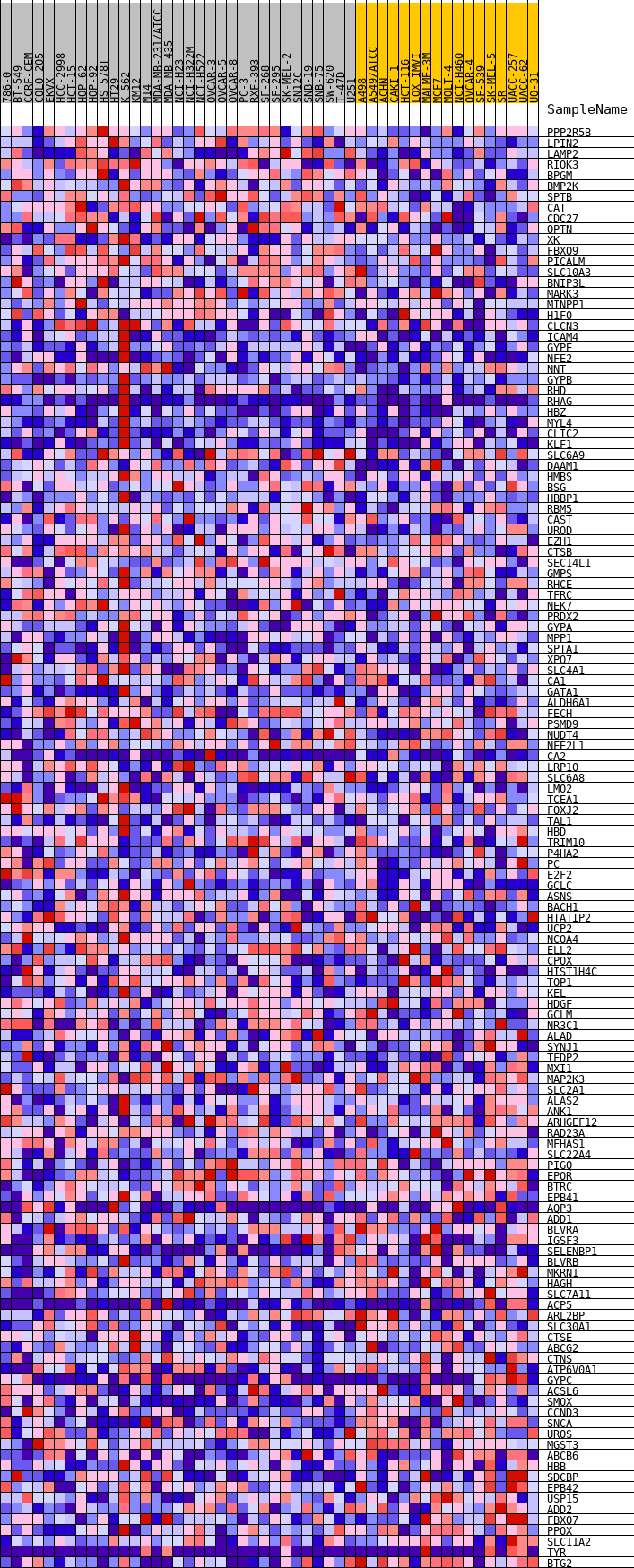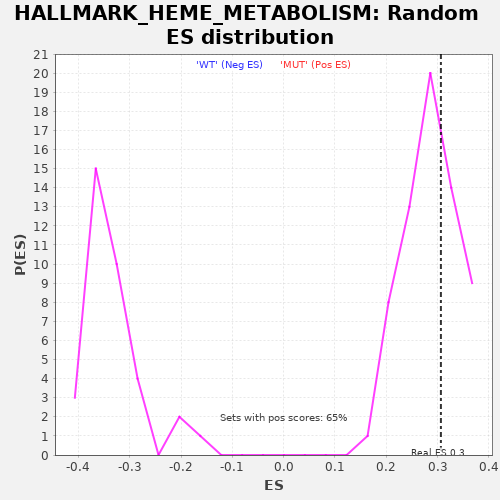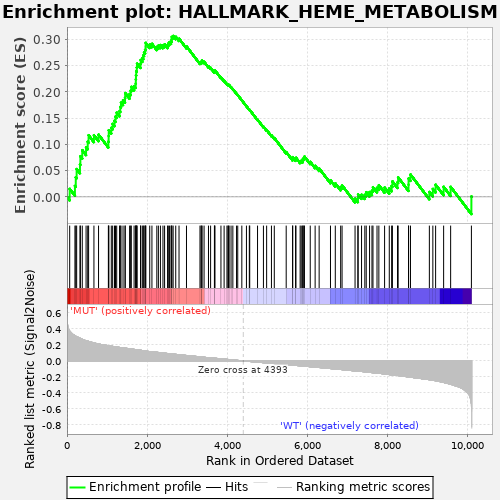
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_HEME_METABOLISM |
| Enrichment Score (ES) | 0.30647236 |
| Normalized Enrichment Score (NES) | 1.0738864 |
| Nominal p-value | 0.35384616 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PPP2R5B | na | 67 | 0.372 | 0.0153 | Yes | ||
| 2 | LPIN2 | na | 198 | 0.316 | 0.0210 | Yes | ||
| 3 | LAMP2 | na | 219 | 0.308 | 0.0373 | Yes | ||
| 4 | RIOK3 | na | 241 | 0.302 | 0.0531 | Yes | ||
| 5 | BPGM | na | 324 | 0.284 | 0.0617 | Yes | ||
| 6 | BMP2K | na | 334 | 0.281 | 0.0775 | Yes | ||
| 7 | SPTB | na | 381 | 0.270 | 0.0889 | Yes | ||
| 8 | CAT | na | 476 | 0.252 | 0.0945 | Yes | ||
| 9 | CDC27 | na | 517 | 0.247 | 0.1051 | Yes | ||
| 10 | OPTN | na | 539 | 0.243 | 0.1174 | Yes | ||
| 11 | XK | na | 672 | 0.225 | 0.1175 | Yes | ||
| 12 | FBXO9 | na | 787 | 0.210 | 0.1185 | Yes | ||
| 13 | PICALM | na | 1032 | 0.190 | 0.1053 | Yes | ||
| 14 | SLC10A3 | na | 1039 | 0.189 | 0.1159 | Yes | ||
| 15 | BNIP3L | na | 1042 | 0.189 | 0.1269 | Yes | ||
| 16 | MARK3 | na | 1105 | 0.184 | 0.1316 | Yes | ||
| 17 | MINPP1 | na | 1139 | 0.182 | 0.1391 | Yes | ||
| 18 | H1F0 | na | 1185 | 0.177 | 0.1451 | Yes | ||
| 19 | CLCN3 | na | 1210 | 0.175 | 0.1530 | Yes | ||
| 20 | ICAM4 | na | 1240 | 0.173 | 0.1604 | Yes | ||
| 21 | GYPE | na | 1311 | 0.168 | 0.1633 | Yes | ||
| 22 | NFE2 | na | 1330 | 0.166 | 0.1714 | Yes | ||
| 23 | NNT | na | 1347 | 0.165 | 0.1796 | Yes | ||
| 24 | GYPB | na | 1398 | 0.162 | 0.1842 | Yes | ||
| 25 | RHD | na | 1448 | 0.159 | 0.1887 | Yes | ||
| 26 | RHAG | na | 1453 | 0.159 | 0.1978 | Yes | ||
| 27 | HBZ | na | 1562 | 0.151 | 0.1959 | Yes | ||
| 28 | MYL4 | na | 1588 | 0.149 | 0.2022 | Yes | ||
| 29 | CLIC2 | na | 1604 | 0.148 | 0.2095 | Yes | ||
| 30 | KLF1 | na | 1674 | 0.143 | 0.2111 | Yes | ||
| 31 | SLC6A9 | na | 1714 | 0.140 | 0.2155 | Yes | ||
| 32 | DAAM1 | na | 1717 | 0.140 | 0.2236 | Yes | ||
| 33 | HMBS | na | 1721 | 0.140 | 0.2316 | Yes | ||
| 34 | BSG | na | 1728 | 0.140 | 0.2393 | Yes | ||
| 35 | HBBP1 | na | 1741 | 0.139 | 0.2463 | Yes | ||
| 36 | RBM5 | na | 1748 | 0.138 | 0.2539 | Yes | ||
| 37 | CAST | na | 1834 | 0.133 | 0.2533 | Yes | ||
| 38 | UROD | na | 1839 | 0.133 | 0.2607 | Yes | ||
| 39 | EZH1 | na | 1879 | 0.129 | 0.2645 | Yes | ||
| 40 | CTSB | na | 1904 | 0.128 | 0.2696 | Yes | ||
| 41 | SEC14L1 | na | 1923 | 0.126 | 0.2753 | Yes | ||
| 42 | GMPS | na | 1949 | 0.123 | 0.2801 | Yes | ||
| 43 | RHCE | na | 1963 | 0.122 | 0.2860 | Yes | ||
| 44 | TFRC | na | 1964 | 0.122 | 0.2933 | Yes | ||
| 45 | NEK7 | na | 2064 | 0.116 | 0.2902 | Yes | ||
| 46 | PRDX2 | na | 2119 | 0.113 | 0.2915 | Yes | ||
| 47 | GYPA | na | 2241 | 0.107 | 0.2857 | Yes | ||
| 48 | MPP1 | na | 2280 | 0.105 | 0.2881 | Yes | ||
| 49 | SPTA1 | na | 2331 | 0.102 | 0.2892 | Yes | ||
| 50 | XPO7 | na | 2394 | 0.098 | 0.2888 | Yes | ||
| 51 | SLC4A1 | na | 2435 | 0.096 | 0.2905 | Yes | ||
| 52 | CA1 | na | 2509 | 0.092 | 0.2886 | Yes | ||
| 53 | GATA1 | na | 2527 | 0.091 | 0.2923 | Yes | ||
| 54 | ALDH6A1 | na | 2556 | 0.090 | 0.2948 | Yes | ||
| 55 | FECH | na | 2595 | 0.088 | 0.2962 | Yes | ||
| 56 | PSMD9 | na | 2611 | 0.087 | 0.2998 | Yes | ||
| 57 | NUDT4 | na | 2615 | 0.087 | 0.3047 | Yes | ||
| 58 | NFE2L1 | na | 2648 | 0.085 | 0.3065 | Yes | ||
| 59 | CA2 | na | 2713 | 0.081 | 0.3048 | No | ||
| 60 | LRP10 | na | 2795 | 0.076 | 0.3012 | No | ||
| 61 | SLC6A8 | na | 2981 | 0.067 | 0.2867 | No | ||
| 62 | LMO2 | na | 3318 | 0.050 | 0.2559 | No | ||
| 63 | TCEA1 | na | 3345 | 0.049 | 0.2562 | No | ||
| 64 | FOXJ2 | na | 3364 | 0.048 | 0.2572 | No | ||
| 65 | TAL1 | na | 3367 | 0.047 | 0.2598 | No | ||
| 66 | HBD | na | 3420 | 0.045 | 0.2573 | No | ||
| 67 | TRIM10 | na | 3530 | 0.040 | 0.2487 | No | ||
| 68 | P4HA2 | na | 3583 | 0.037 | 0.2457 | No | ||
| 69 | PC | na | 3678 | 0.033 | 0.2382 | No | ||
| 70 | E2F2 | na | 3682 | 0.033 | 0.2399 | No | ||
| 71 | GCLC | na | 3691 | 0.033 | 0.2410 | No | ||
| 72 | ASNS | na | 3841 | 0.026 | 0.2276 | No | ||
| 73 | BACH1 | na | 3920 | 0.022 | 0.2211 | No | ||
| 74 | HTATIP2 | na | 3991 | 0.019 | 0.2151 | No | ||
| 75 | UCP2 | na | 4016 | 0.017 | 0.2138 | No | ||
| 76 | NCOA4 | na | 4030 | 0.017 | 0.2134 | No | ||
| 77 | ELL2 | na | 4045 | 0.015 | 0.2129 | No | ||
| 78 | CPOX | na | 4093 | 0.013 | 0.2090 | No | ||
| 79 | HIST1H4C | na | 4138 | 0.011 | 0.2052 | No | ||
| 80 | TOP1 | na | 4230 | 0.007 | 0.1965 | No | ||
| 81 | KEL | na | 4260 | 0.005 | 0.1939 | No | ||
| 82 | HDGF | na | 4362 | 0.001 | 0.1839 | No | ||
| 83 | GCLM | na | 4477 | -0.004 | 0.1727 | No | ||
| 84 | NR3C1 | na | 4549 | -0.007 | 0.1660 | No | ||
| 85 | ALAD | na | 4559 | -0.008 | 0.1655 | No | ||
| 86 | SYNJ1 | na | 4754 | -0.017 | 0.1471 | No | ||
| 87 | TFDP2 | na | 4901 | -0.022 | 0.1338 | No | ||
| 88 | MXI1 | na | 4981 | -0.026 | 0.1274 | No | ||
| 89 | MAP2K3 | na | 5101 | -0.031 | 0.1172 | No | ||
| 90 | SLC2A1 | na | 5172 | -0.034 | 0.1122 | No | ||
| 91 | ALAS2 | na | 5469 | -0.047 | 0.0853 | No | ||
| 92 | ANK1 | na | 5629 | -0.054 | 0.0726 | No | ||
| 93 | ARHGEF12 | na | 5631 | -0.054 | 0.0757 | No | ||
| 94 | RAD23A | na | 5703 | -0.057 | 0.0720 | No | ||
| 95 | MFHAS1 | na | 5712 | -0.058 | 0.0746 | No | ||
| 96 | SLC22A4 | na | 5817 | -0.062 | 0.0678 | No | ||
| 97 | PIGQ | na | 5844 | -0.063 | 0.0690 | No | ||
| 98 | EPOR | na | 5880 | -0.065 | 0.0693 | No | ||
| 99 | BTRC | na | 5888 | -0.065 | 0.0724 | No | ||
| 100 | EPB41 | na | 5910 | -0.066 | 0.0742 | No | ||
| 101 | AQP3 | na | 5924 | -0.066 | 0.0769 | No | ||
| 102 | ADD1 | na | 6069 | -0.073 | 0.0667 | No | ||
| 103 | BLVRA | na | 6191 | -0.078 | 0.0592 | No | ||
| 104 | IGSF3 | na | 6291 | -0.083 | 0.0542 | No | ||
| 105 | SELENBP1 | na | 6574 | -0.096 | 0.0317 | No | ||
| 106 | BLVRB | na | 6694 | -0.102 | 0.0258 | No | ||
| 107 | MKRN1 | na | 6828 | -0.108 | 0.0188 | No | ||
| 108 | HAGH | na | 6863 | -0.110 | 0.0219 | No | ||
| 109 | SLC7A11 | na | 7187 | -0.125 | -0.0031 | No | ||
| 110 | ACP5 | na | 7258 | -0.129 | -0.0025 | No | ||
| 111 | ARL2BP | na | 7263 | -0.129 | 0.0048 | No | ||
| 112 | SLC30A1 | na | 7349 | -0.133 | 0.0041 | No | ||
| 113 | CTSE | na | 7431 | -0.137 | 0.0041 | No | ||
| 114 | ABCG2 | na | 7466 | -0.139 | 0.0090 | No | ||
| 115 | CTNS | na | 7549 | -0.144 | 0.0093 | No | ||
| 116 | ATP6V0A1 | na | 7609 | -0.147 | 0.0121 | No | ||
| 117 | GYPC | na | 7634 | -0.148 | 0.0185 | No | ||
| 118 | ACSL6 | na | 7735 | -0.154 | 0.0175 | No | ||
| 119 | SMOX | na | 7781 | -0.156 | 0.0223 | No | ||
| 120 | CCND3 | na | 7924 | -0.164 | 0.0178 | No | ||
| 121 | SNCA | na | 8041 | -0.172 | 0.0163 | No | ||
| 122 | UROS | na | 8100 | -0.175 | 0.0209 | No | ||
| 123 | MGST3 | na | 8119 | -0.176 | 0.0295 | No | ||
| 124 | ABCB6 | na | 8246 | -0.184 | 0.0278 | No | ||
| 125 | HBB | na | 8262 | -0.184 | 0.0372 | No | ||
| 126 | SDCBP | na | 8521 | -0.201 | 0.0232 | No | ||
| 127 | EPB42 | na | 8523 | -0.201 | 0.0351 | No | ||
| 128 | USP15 | na | 8570 | -0.205 | 0.0426 | No | ||
| 129 | ADD2 | na | 9041 | -0.236 | 0.0094 | No | ||
| 130 | FBXO7 | na | 9128 | -0.243 | 0.0152 | No | ||
| 131 | PPOX | na | 9197 | -0.249 | 0.0232 | No | ||
| 132 | SLC11A2 | na | 9397 | -0.270 | 0.0192 | No | ||
| 133 | TYR | na | 9572 | -0.293 | 0.0192 | No | ||
| 134 | BTG2 | na | 10090 | -0.567 | 0.0009 | No |