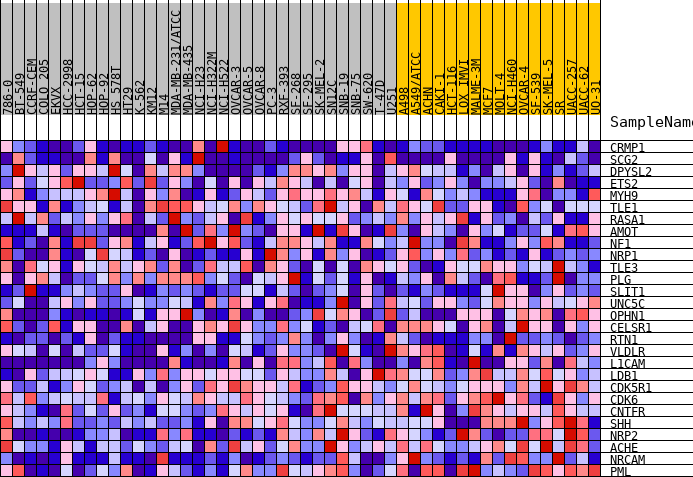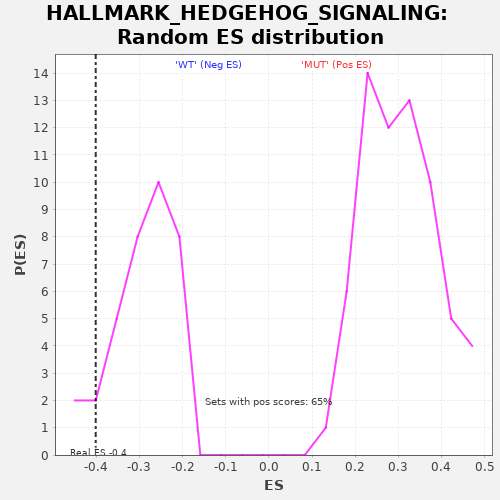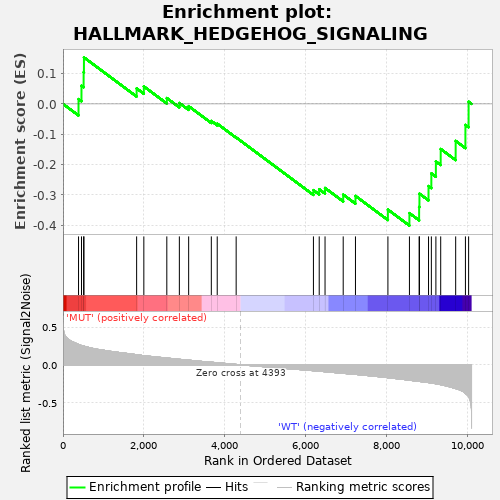
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_HEDGEHOG_SIGNALING |
| Enrichment Score (ES) | -0.40139312 |
| Normalized Enrichment Score (NES) | -1.3817556 |
| Nominal p-value | 0.057142857 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.72 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CRMP1 | na | 384 | 0.269 | 0.0159 | No | ||
| 2 | SCG2 | na | 457 | 0.255 | 0.0599 | No | ||
| 3 | DPYSL2 | na | 510 | 0.247 | 0.1045 | No | ||
| 4 | ETS2 | na | 518 | 0.247 | 0.1534 | No | ||
| 5 | MYH9 | na | 1820 | 0.134 | 0.0511 | No | ||
| 6 | TLE1 | na | 1998 | 0.120 | 0.0577 | No | ||
| 7 | RASA1 | na | 2566 | 0.089 | 0.0192 | No | ||
| 8 | AMOT | na | 2876 | 0.072 | 0.0030 | No | ||
| 9 | NF1 | na | 3106 | 0.061 | -0.0075 | No | ||
| 10 | NRP1 | na | 3666 | 0.034 | -0.0563 | No | ||
| 11 | TLE3 | na | 3812 | 0.027 | -0.0653 | No | ||
| 12 | PLG | na | 4280 | 0.005 | -0.1107 | No | ||
| 13 | SLIT1 | na | 6190 | -0.078 | -0.2845 | No | ||
| 14 | UNC5C | na | 6333 | -0.085 | -0.2816 | No | ||
| 15 | OPHN1 | na | 6477 | -0.092 | -0.2773 | No | ||
| 16 | CELSR1 | na | 6926 | -0.113 | -0.2991 | No | ||
| 17 | RTN1 | na | 7229 | -0.127 | -0.3036 | No | ||
| 18 | VLDLR | na | 8031 | -0.171 | -0.3487 | No | ||
| 19 | L1CAM | na | 8563 | -0.204 | -0.3604 | Yes | ||
| 20 | LDB1 | na | 8803 | -0.220 | -0.3398 | Yes | ||
| 21 | CDK5R1 | na | 8811 | -0.221 | -0.2962 | Yes | ||
| 22 | CDK6 | na | 9035 | -0.236 | -0.2710 | Yes | ||
| 23 | CNTFR | na | 9105 | -0.242 | -0.2293 | Yes | ||
| 24 | SHH | na | 9217 | -0.251 | -0.1899 | Yes | ||
| 25 | NRP2 | na | 9336 | -0.263 | -0.1487 | Yes | ||
| 26 | ACHE | na | 9706 | -0.315 | -0.1221 | Yes | ||
| 27 | NRCAM | na | 9947 | -0.381 | -0.0693 | Yes | ||
| 28 | PML | na | 10026 | -0.420 | 0.0072 | Yes |