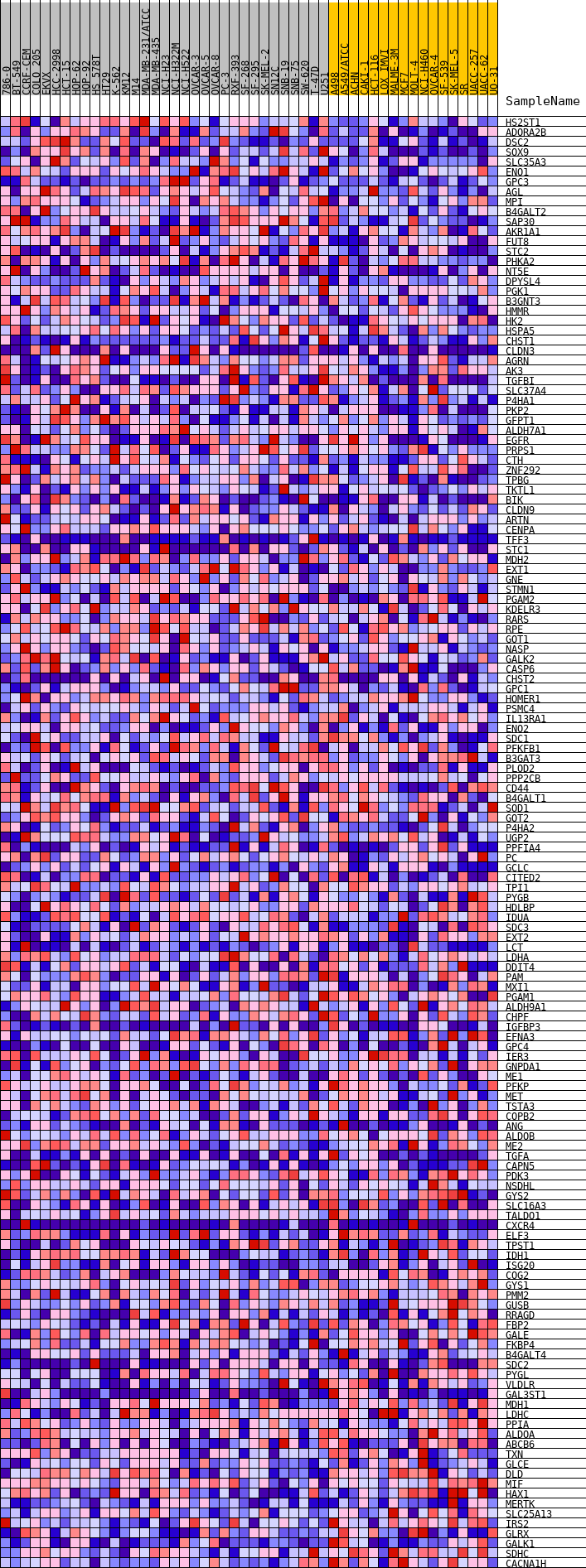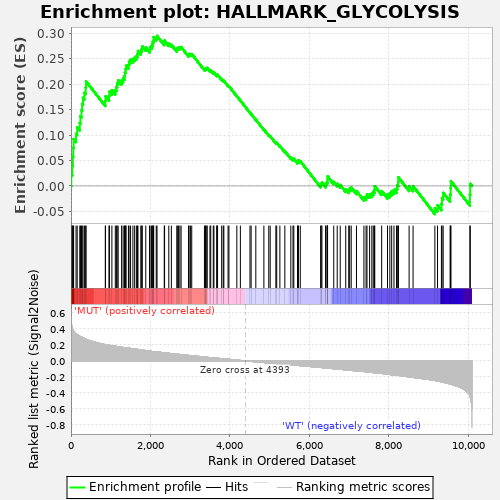
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_GLYCOLYSIS |
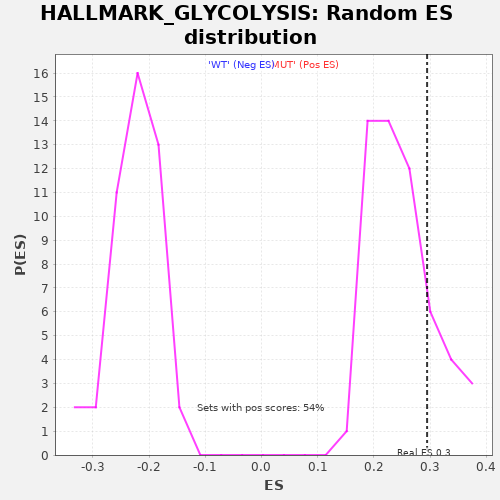
| Enrichment Score (ES) | 0.2947977 |
| Normalized Enrichment Score (NES) | 1.176057 |
| Nominal p-value | 0.18518518 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HS2ST1 | na | 7 | 0.487 | 0.0228 | Yes | ||
| 2 | ADORA2B | na | 33 | 0.409 | 0.0401 | Yes | ||
| 3 | DSC2 | na | 47 | 0.390 | 0.0576 | Yes | ||
| 4 | SOX9 | na | 57 | 0.378 | 0.0750 | Yes | ||
| 5 | SLC35A3 | na | 66 | 0.372 | 0.0921 | Yes | ||
| 6 | ENO1 | na | 127 | 0.340 | 0.1025 | Yes | ||
| 7 | GPC3 | na | 156 | 0.327 | 0.1155 | Yes | ||
| 8 | AGL | na | 221 | 0.308 | 0.1239 | Yes | ||
| 9 | MPI | na | 238 | 0.303 | 0.1369 | Yes | ||
| 10 | B4GALT2 | na | 265 | 0.297 | 0.1487 | Yes | ||
| 11 | SAP30 | na | 280 | 0.294 | 0.1615 | Yes | ||
| 12 | AKR1A1 | na | 302 | 0.289 | 0.1733 | Yes | ||
| 13 | FUT8 | na | 338 | 0.280 | 0.1833 | Yes | ||
| 14 | STC2 | na | 370 | 0.272 | 0.1934 | Yes | ||
| 15 | PHKA2 | na | 379 | 0.271 | 0.2056 | Yes | ||
| 16 | NT5E | na | 864 | 0.203 | 0.1668 | Yes | ||
| 17 | DPYSL4 | na | 869 | 0.203 | 0.1762 | Yes | ||
| 18 | PGK1 | na | 957 | 0.196 | 0.1770 | Yes | ||
| 19 | B3GNT3 | na | 967 | 0.195 | 0.1855 | Yes | ||
| 20 | HMMR | na | 1030 | 0.190 | 0.1885 | Yes | ||
| 21 | HK2 | na | 1116 | 0.183 | 0.1888 | Yes | ||
| 22 | HSPA5 | na | 1143 | 0.181 | 0.1949 | Yes | ||
| 23 | CHST1 | na | 1163 | 0.179 | 0.2016 | Yes | ||
| 24 | CLDN3 | na | 1188 | 0.177 | 0.2077 | Yes | ||
| 25 | AGRN | na | 1275 | 0.171 | 0.2074 | Yes | ||
| 26 | AK3 | na | 1313 | 0.167 | 0.2117 | Yes | ||
| 27 | TGFBI | na | 1348 | 0.165 | 0.2163 | Yes | ||
| 28 | SLC37A4 | na | 1358 | 0.165 | 0.2233 | Yes | ||
| 29 | P4HA1 | na | 1376 | 0.163 | 0.2295 | Yes | ||
| 30 | PKP2 | na | 1387 | 0.163 | 0.2364 | Yes | ||
| 31 | GFPT1 | na | 1447 | 0.159 | 0.2381 | Yes | ||
| 32 | ALDH7A1 | na | 1462 | 0.158 | 0.2444 | Yes | ||
| 33 | EGFR | na | 1500 | 0.156 | 0.2482 | Yes | ||
| 34 | PRPS1 | na | 1559 | 0.151 | 0.2497 | Yes | ||
| 35 | CTH | na | 1603 | 0.148 | 0.2525 | Yes | ||
| 36 | ZNF292 | na | 1647 | 0.146 | 0.2552 | Yes | ||
| 37 | TPBG | na | 1676 | 0.143 | 0.2593 | Yes | ||
| 38 | TKTL1 | na | 1686 | 0.143 | 0.2653 | Yes | ||
| 39 | BIK | na | 1758 | 0.138 | 0.2648 | Yes | ||
| 40 | CLDN9 | na | 1781 | 0.136 | 0.2692 | Yes | ||
| 41 | ARTN | na | 1795 | 0.136 | 0.2744 | Yes | ||
| 42 | CENPA | na | 1883 | 0.129 | 0.2719 | Yes | ||
| 43 | TFF3 | na | 1980 | 0.121 | 0.2681 | Yes | ||
| 44 | STC1 | na | 1990 | 0.120 | 0.2731 | Yes | ||
| 45 | MDH2 | na | 2029 | 0.118 | 0.2749 | Yes | ||
| 46 | EXT1 | na | 2047 | 0.117 | 0.2789 | Yes | ||
| 47 | GNE | na | 2060 | 0.116 | 0.2833 | Yes | ||
| 48 | STMN1 | na | 2080 | 0.115 | 0.2869 | Yes | ||
| 49 | PGAM2 | na | 2081 | 0.115 | 0.2925 | Yes | ||
| 50 | KDELR3 | na | 2147 | 0.112 | 0.2914 | Yes | ||
| 51 | RARS | na | 2167 | 0.111 | 0.2948 | Yes | ||
| 52 | RPE | na | 2349 | 0.101 | 0.2815 | No | ||
| 53 | GOT1 | na | 2357 | 0.100 | 0.2856 | No | ||
| 54 | NASP | na | 2463 | 0.094 | 0.2796 | No | ||
| 55 | GALK2 | na | 2528 | 0.091 | 0.2776 | No | ||
| 56 | CASP6 | na | 2667 | 0.084 | 0.2678 | No | ||
| 57 | CHST2 | na | 2676 | 0.083 | 0.2710 | No | ||
| 58 | GPC1 | na | 2706 | 0.081 | 0.2720 | No | ||
| 59 | HOMER1 | na | 2742 | 0.080 | 0.2723 | No | ||
| 60 | PSMC4 | na | 2773 | 0.078 | 0.2730 | No | ||
| 61 | IL13RA1 | na | 2960 | 0.068 | 0.2577 | No | ||
| 62 | ENO2 | na | 2975 | 0.067 | 0.2595 | No | ||
| 63 | SDC1 | na | 3012 | 0.066 | 0.2590 | No | ||
| 64 | PFKFB1 | na | 3041 | 0.064 | 0.2593 | No | ||
| 65 | B3GAT3 | na | 3361 | 0.048 | 0.2296 | No | ||
| 66 | PLOD2 | na | 3385 | 0.046 | 0.2295 | No | ||
| 67 | PPP2CB | na | 3394 | 0.046 | 0.2309 | No | ||
| 68 | CD44 | na | 3423 | 0.045 | 0.2303 | No | ||
| 69 | B4GALT1 | na | 3427 | 0.044 | 0.2321 | No | ||
| 70 | SOD1 | na | 3500 | 0.041 | 0.2269 | No | ||
| 71 | GOT2 | na | 3519 | 0.040 | 0.2270 | No | ||
| 72 | P4HA2 | na | 3583 | 0.037 | 0.2225 | No | ||
| 73 | UGP2 | na | 3600 | 0.037 | 0.2226 | No | ||
| 74 | PPFIA4 | na | 3671 | 0.033 | 0.2172 | No | ||
| 75 | PC | na | 3678 | 0.033 | 0.2182 | No | ||
| 76 | GCLC | na | 3691 | 0.033 | 0.2186 | No | ||
| 77 | CITED2 | na | 3795 | 0.028 | 0.2096 | No | ||
| 78 | TPI1 | na | 3834 | 0.026 | 0.2070 | No | ||
| 79 | PYGB | na | 3850 | 0.025 | 0.2067 | No | ||
| 80 | HDLBP | na | 3956 | 0.020 | 0.1971 | No | ||
| 81 | IDUA | na | 3978 | 0.019 | 0.1959 | No | ||
| 82 | SDC3 | na | 4177 | 0.009 | 0.1765 | No | ||
| 83 | EXT2 | na | 4266 | 0.005 | 0.1679 | No | ||
| 84 | LCT | na | 4508 | -0.005 | 0.1440 | No | ||
| 85 | LDHA | na | 4537 | -0.007 | 0.1415 | No | ||
| 86 | DDIT4 | na | 4655 | -0.012 | 0.1303 | No | ||
| 87 | PAM | na | 4858 | -0.021 | 0.1110 | No | ||
| 88 | MXI1 | na | 4981 | -0.026 | 0.1000 | No | ||
| 89 | PGAM1 | na | 5017 | -0.027 | 0.0978 | No | ||
| 90 | ALDH9A1 | na | 5160 | -0.033 | 0.0851 | No | ||
| 91 | CHPF | na | 5181 | -0.034 | 0.0848 | No | ||
| 92 | IGFBP3 | na | 5261 | -0.038 | 0.0787 | No | ||
| 93 | EFNA3 | na | 5386 | -0.044 | 0.0683 | No | ||
| 94 | GPC4 | na | 5537 | -0.050 | 0.0557 | No | ||
| 95 | IER3 | na | 5592 | -0.053 | 0.0528 | No | ||
| 96 | GNPDA1 | na | 5612 | -0.054 | 0.0535 | No | ||
| 97 | ME1 | na | 5706 | -0.057 | 0.0469 | No | ||
| 98 | PFKP | na | 5709 | -0.057 | 0.0495 | No | ||
| 99 | MET | na | 5729 | -0.058 | 0.0504 | No | ||
| 100 | TSTA3 | na | 5775 | -0.060 | 0.0488 | No | ||
| 101 | COPB2 | na | 6288 | -0.083 | 0.0014 | No | ||
| 102 | ANG | na | 6300 | -0.084 | 0.0043 | No | ||
| 103 | ALDOB | na | 6320 | -0.085 | 0.0065 | No | ||
| 104 | ME2 | na | 6414 | -0.089 | 0.0014 | No | ||
| 105 | TGFA | na | 6417 | -0.089 | 0.0055 | No | ||
| 106 | CAPN5 | na | 6449 | -0.090 | 0.0068 | No | ||
| 107 | PDK3 | na | 6460 | -0.091 | 0.0101 | No | ||
| 108 | NSDHL | na | 6461 | -0.091 | 0.0145 | No | ||
| 109 | GYS2 | na | 6462 | -0.091 | 0.0189 | No | ||
| 110 | SLC16A3 | na | 6618 | -0.099 | 0.0081 | No | ||
| 111 | TALDO1 | na | 6705 | -0.103 | 0.0044 | No | ||
| 112 | CXCR4 | na | 6781 | -0.106 | 0.0020 | No | ||
| 113 | ELF3 | na | 6920 | -0.113 | -0.0064 | No | ||
| 114 | TPST1 | na | 6995 | -0.116 | -0.0082 | No | ||
| 115 | IDH1 | na | 7019 | -0.117 | -0.0048 | No | ||
| 116 | ISG20 | na | 7058 | -0.120 | -0.0029 | No | ||
| 117 | COG2 | na | 7190 | -0.125 | -0.0100 | No | ||
| 118 | GYS1 | na | 7378 | -0.134 | -0.0223 | No | ||
| 119 | PMM2 | na | 7432 | -0.137 | -0.0210 | No | ||
| 120 | GUSB | na | 7455 | -0.139 | -0.0165 | No | ||
| 121 | RRAGD | na | 7521 | -0.142 | -0.0161 | No | ||
| 122 | FBP2 | na | 7577 | -0.146 | -0.0146 | No | ||
| 123 | GALE | na | 7614 | -0.147 | -0.0111 | No | ||
| 124 | FKBP4 | na | 7643 | -0.149 | -0.0068 | No | ||
| 125 | B4GALT4 | na | 7651 | -0.149 | -0.0003 | No | ||
| 126 | SDC2 | na | 7825 | -0.158 | -0.0100 | No | ||
| 127 | PYGL | na | 7972 | -0.167 | -0.0166 | No | ||
| 128 | VLDLR | na | 8031 | -0.171 | -0.0142 | No | ||
| 129 | GAL3ST1 | na | 8077 | -0.174 | -0.0103 | No | ||
| 130 | MDH1 | na | 8135 | -0.177 | -0.0075 | No | ||
| 131 | LDHC | na | 8199 | -0.181 | -0.0051 | No | ||
| 132 | PPIA | na | 8218 | -0.182 | 0.0019 | No | ||
| 133 | ALDOA | na | 8243 | -0.184 | 0.0084 | No | ||
| 134 | ABCB6 | na | 8246 | -0.184 | 0.0171 | No | ||
| 135 | TXN | na | 8516 | -0.200 | -0.0003 | No | ||
| 136 | GLCE | na | 8616 | -0.208 | -0.0002 | No | ||
| 137 | DLD | na | 9165 | -0.246 | -0.0434 | No | ||
| 138 | MIF | na | 9232 | -0.252 | -0.0379 | No | ||
| 139 | HAX1 | na | 9330 | -0.263 | -0.0349 | No | ||
| 140 | MERTK | na | 9344 | -0.264 | -0.0235 | No | ||
| 141 | SLC25A13 | na | 9374 | -0.268 | -0.0135 | No | ||
| 142 | IRS2 | na | 9549 | -0.290 | -0.0169 | No | ||
| 143 | GLRX | na | 9563 | -0.292 | -0.0042 | No | ||
| 144 | GALK1 | na | 9568 | -0.293 | 0.0096 | No | ||
| 145 | SDHC | na | 10047 | -0.444 | -0.0170 | No | ||
| 146 | CACNA1H | na | 10056 | -0.458 | 0.0043 | No |