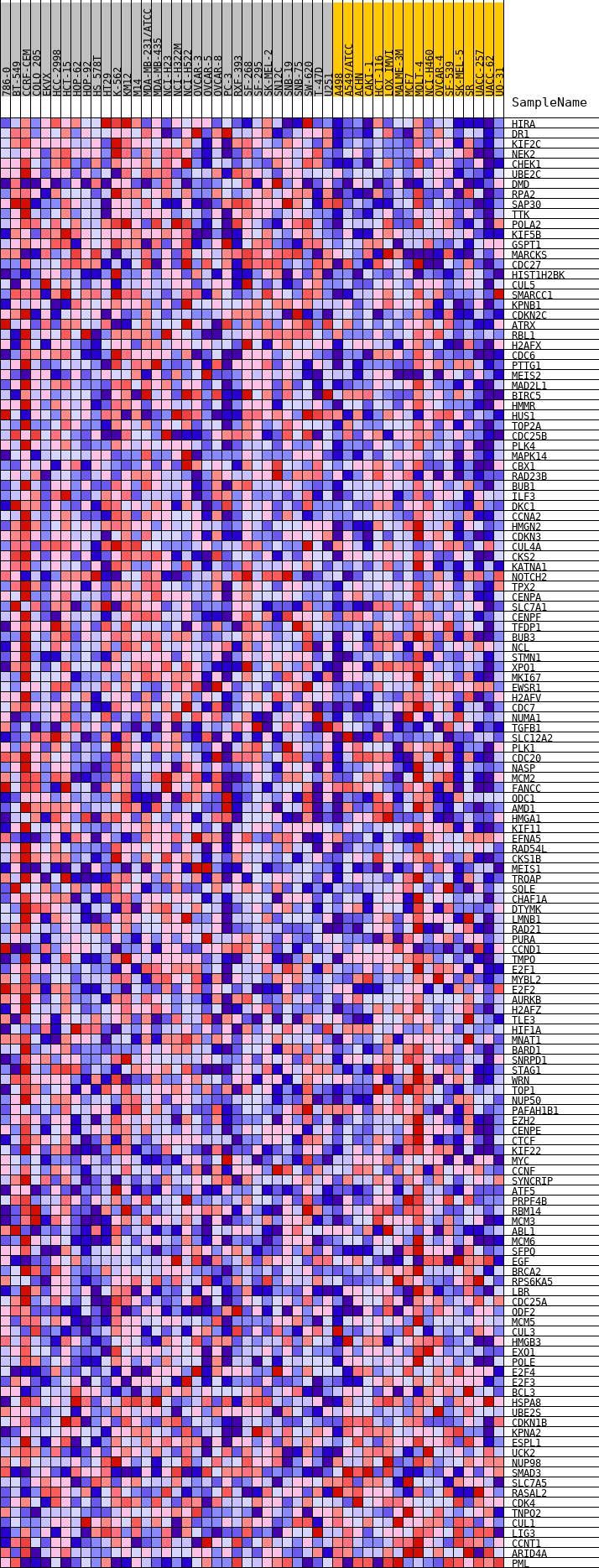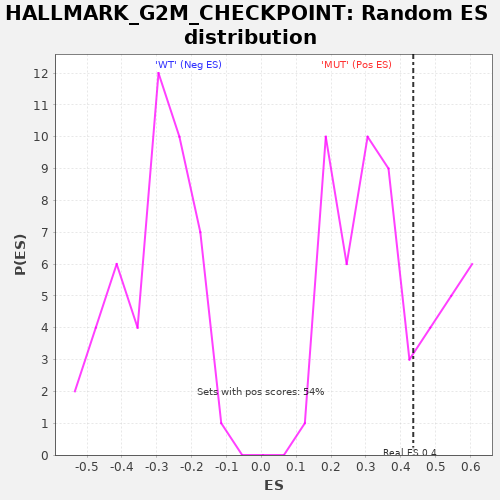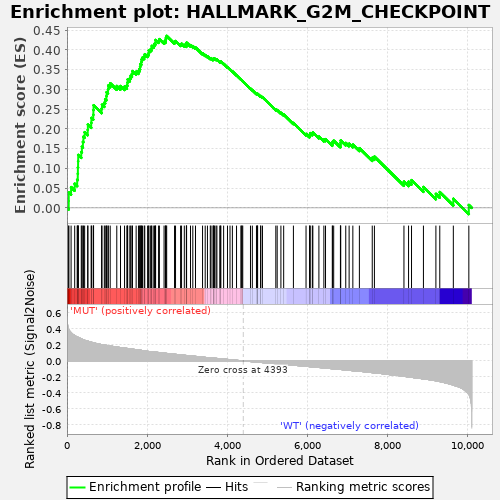
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_G2M_CHECKPOINT |
| Enrichment Score (ES) | 0.43566397 |
| Normalized Enrichment Score (NES) | 1.2074549 |
| Nominal p-value | 0.2962963 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.98 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HIRA | na | 43 | 0.395 | 0.0174 | Yes | ||
| 2 | DR1 | na | 44 | 0.393 | 0.0390 | Yes | ||
| 3 | KIF2C | na | 102 | 0.351 | 0.0526 | Yes | ||
| 4 | NEK2 | na | 192 | 0.318 | 0.0611 | Yes | ||
| 5 | CHEK1 | na | 254 | 0.299 | 0.0714 | Yes | ||
| 6 | UBE2C | na | 269 | 0.297 | 0.0863 | Yes | ||
| 7 | DMD | na | 272 | 0.296 | 0.1024 | Yes | ||
| 8 | RPA2 | na | 277 | 0.295 | 0.1182 | Yes | ||
| 9 | SAP30 | na | 280 | 0.294 | 0.1341 | Yes | ||
| 10 | TTK | na | 356 | 0.274 | 0.1417 | Yes | ||
| 11 | POLA2 | na | 372 | 0.271 | 0.1551 | Yes | ||
| 12 | KIF5B | na | 397 | 0.265 | 0.1673 | Yes | ||
| 13 | GSPT1 | na | 410 | 0.262 | 0.1805 | Yes | ||
| 14 | MARCKS | na | 440 | 0.258 | 0.1918 | Yes | ||
| 15 | CDC27 | na | 517 | 0.247 | 0.1977 | Yes | ||
| 16 | HIST1H2BK | na | 521 | 0.246 | 0.2109 | Yes | ||
| 17 | CUL5 | na | 604 | 0.233 | 0.2155 | Yes | ||
| 18 | SMARCC1 | na | 609 | 0.233 | 0.2279 | Yes | ||
| 19 | KPNB1 | na | 655 | 0.226 | 0.2358 | Yes | ||
| 20 | CDKN2C | na | 659 | 0.226 | 0.2479 | Yes | ||
| 21 | ATRX | na | 663 | 0.225 | 0.2600 | Yes | ||
| 22 | RBL1 | na | 863 | 0.204 | 0.2512 | Yes | ||
| 23 | H2AFX | na | 873 | 0.203 | 0.2614 | Yes | ||
| 24 | CDC6 | na | 931 | 0.198 | 0.2666 | Yes | ||
| 25 | PTTG1 | na | 953 | 0.196 | 0.2753 | Yes | ||
| 26 | MEIS2 | na | 983 | 0.193 | 0.2830 | Yes | ||
| 27 | MAD2L1 | na | 988 | 0.193 | 0.2932 | Yes | ||
| 28 | BIRC5 | na | 1023 | 0.191 | 0.3003 | Yes | ||
| 29 | HMMR | na | 1030 | 0.190 | 0.3101 | Yes | ||
| 30 | HUS1 | na | 1078 | 0.187 | 0.3157 | Yes | ||
| 31 | TOP2A | na | 1242 | 0.173 | 0.3088 | Yes | ||
| 32 | CDC25B | na | 1335 | 0.166 | 0.3087 | Yes | ||
| 33 | PLK4 | na | 1436 | 0.160 | 0.3074 | Yes | ||
| 34 | MAPK14 | na | 1494 | 0.156 | 0.3103 | Yes | ||
| 35 | CBX1 | na | 1509 | 0.155 | 0.3174 | Yes | ||
| 36 | RAD23B | na | 1514 | 0.154 | 0.3255 | Yes | ||
| 37 | BUB1 | na | 1568 | 0.151 | 0.3284 | Yes | ||
| 38 | ILF3 | na | 1589 | 0.149 | 0.3346 | Yes | ||
| 39 | DKC1 | na | 1621 | 0.147 | 0.3396 | Yes | ||
| 40 | CCNA2 | na | 1634 | 0.146 | 0.3464 | Yes | ||
| 41 | HMGN2 | na | 1725 | 0.140 | 0.3451 | Yes | ||
| 42 | CDKN3 | na | 1783 | 0.136 | 0.3468 | Yes | ||
| 43 | CUL4A | na | 1806 | 0.135 | 0.3520 | Yes | ||
| 44 | CKS2 | na | 1824 | 0.134 | 0.3577 | Yes | ||
| 45 | KATNA1 | na | 1830 | 0.133 | 0.3645 | Yes | ||
| 46 | NOTCH2 | na | 1854 | 0.132 | 0.3694 | Yes | ||
| 47 | TPX2 | na | 1860 | 0.130 | 0.3761 | Yes | ||
| 48 | CENPA | na | 1883 | 0.129 | 0.3810 | Yes | ||
| 49 | SLC7A1 | na | 1926 | 0.126 | 0.3837 | Yes | ||
| 50 | CENPF | na | 1940 | 0.125 | 0.3892 | Yes | ||
| 51 | TFDP1 | na | 2011 | 0.119 | 0.3887 | Yes | ||
| 52 | BUB3 | na | 2038 | 0.118 | 0.3926 | Yes | ||
| 53 | NCL | na | 2040 | 0.117 | 0.3989 | Yes | ||
| 54 | STMN1 | na | 2080 | 0.115 | 0.4013 | Yes | ||
| 55 | XPO1 | na | 2111 | 0.113 | 0.4046 | Yes | ||
| 56 | MKI67 | na | 2116 | 0.113 | 0.4104 | Yes | ||
| 57 | EWSR1 | na | 2166 | 0.111 | 0.4115 | Yes | ||
| 58 | H2AFV | na | 2184 | 0.110 | 0.4159 | Yes | ||
| 59 | CDC7 | na | 2209 | 0.109 | 0.4194 | Yes | ||
| 60 | NUMA1 | na | 2210 | 0.109 | 0.4254 | Yes | ||
| 61 | TGFB1 | na | 2285 | 0.105 | 0.4237 | Yes | ||
| 62 | SLC12A2 | na | 2305 | 0.103 | 0.4275 | Yes | ||
| 63 | PLK1 | na | 2422 | 0.097 | 0.4212 | Yes | ||
| 64 | CDC20 | na | 2457 | 0.095 | 0.4230 | Yes | ||
| 65 | NASP | na | 2463 | 0.094 | 0.4276 | Yes | ||
| 66 | MCM2 | na | 2470 | 0.094 | 0.4322 | Yes | ||
| 67 | FANCC | na | 2487 | 0.093 | 0.4357 | Yes | ||
| 68 | ODC1 | na | 2683 | 0.083 | 0.4206 | No | ||
| 69 | AMD1 | na | 2708 | 0.081 | 0.4227 | No | ||
| 70 | HMGA1 | na | 2834 | 0.074 | 0.4142 | No | ||
| 71 | KIF11 | na | 2858 | 0.073 | 0.4159 | No | ||
| 72 | EFNA5 | na | 2927 | 0.070 | 0.4129 | No | ||
| 73 | RAD54L | na | 2979 | 0.067 | 0.4115 | No | ||
| 74 | CKS1B | na | 2980 | 0.067 | 0.4152 | No | ||
| 75 | MEIS1 | na | 2984 | 0.067 | 0.4185 | No | ||
| 76 | TROAP | na | 3081 | 0.062 | 0.4123 | No | ||
| 77 | SQLE | na | 3139 | 0.059 | 0.4099 | No | ||
| 78 | CHAF1A | na | 3206 | 0.056 | 0.4063 | No | ||
| 79 | DTYMK | na | 3382 | 0.047 | 0.3913 | No | ||
| 80 | LMNB1 | na | 3448 | 0.043 | 0.3871 | No | ||
| 81 | RAD21 | na | 3504 | 0.041 | 0.3839 | No | ||
| 82 | PURA | na | 3575 | 0.038 | 0.3789 | No | ||
| 83 | CCND1 | na | 3593 | 0.037 | 0.3792 | No | ||
| 84 | TMPO | na | 3645 | 0.035 | 0.3760 | No | ||
| 85 | E2F1 | na | 3651 | 0.034 | 0.3774 | No | ||
| 86 | MYBL2 | na | 3662 | 0.034 | 0.3782 | No | ||
| 87 | E2F2 | na | 3682 | 0.033 | 0.3781 | No | ||
| 88 | AURKB | na | 3718 | 0.031 | 0.3764 | No | ||
| 89 | H2AFZ | na | 3740 | 0.030 | 0.3759 | No | ||
| 90 | TLE3 | na | 3812 | 0.027 | 0.3702 | No | ||
| 91 | HIF1A | na | 3833 | 0.026 | 0.3696 | No | ||
| 92 | MNAT1 | na | 3837 | 0.026 | 0.3708 | No | ||
| 93 | BARD1 | na | 3908 | 0.022 | 0.3650 | No | ||
| 94 | SNRPD1 | na | 4004 | 0.018 | 0.3564 | No | ||
| 95 | STAG1 | na | 4067 | 0.014 | 0.3510 | No | ||
| 96 | WRN | na | 4126 | 0.011 | 0.3458 | No | ||
| 97 | TOP1 | na | 4230 | 0.007 | 0.3358 | No | ||
| 98 | NUP50 | na | 4340 | 0.002 | 0.3250 | No | ||
| 99 | PAFAH1B1 | na | 4350 | 0.002 | 0.3242 | No | ||
| 100 | EZH2 | na | 4369 | 0.001 | 0.3224 | No | ||
| 101 | CENPE | na | 4385 | 0.000 | 0.3209 | No | ||
| 102 | CTCF | na | 4578 | -0.009 | 0.3021 | No | ||
| 103 | KIF22 | na | 4624 | -0.010 | 0.2982 | No | ||
| 104 | MYC | na | 4727 | -0.015 | 0.2888 | No | ||
| 105 | CCNF | na | 4745 | -0.016 | 0.2880 | No | ||
| 106 | SYNCRIP | na | 4746 | -0.017 | 0.2889 | No | ||
| 107 | ATF5 | na | 4756 | -0.017 | 0.2889 | No | ||
| 108 | PRPF4B | na | 4831 | -0.020 | 0.2826 | No | ||
| 109 | RBM14 | na | 4835 | -0.020 | 0.2834 | No | ||
| 110 | MCM3 | na | 4877 | -0.022 | 0.2805 | No | ||
| 111 | ABL1 | na | 5210 | -0.036 | 0.2491 | No | ||
| 112 | MCM6 | na | 5244 | -0.037 | 0.2478 | No | ||
| 113 | SFPQ | na | 5337 | -0.042 | 0.2409 | No | ||
| 114 | EGF | na | 5406 | -0.044 | 0.2365 | No | ||
| 115 | BRCA2 | na | 5649 | -0.055 | 0.2152 | No | ||
| 116 | RPS6KA5 | na | 5964 | -0.068 | 0.1874 | No | ||
| 117 | LBR | na | 6052 | -0.072 | 0.1827 | No | ||
| 118 | CDC25A | na | 6059 | -0.073 | 0.1860 | No | ||
| 119 | ODF2 | na | 6067 | -0.073 | 0.1893 | No | ||
| 120 | MCM5 | na | 6121 | -0.075 | 0.1881 | No | ||
| 121 | CUL3 | na | 6134 | -0.076 | 0.1911 | No | ||
| 122 | HMGB3 | na | 6285 | -0.083 | 0.1806 | No | ||
| 123 | EXO1 | na | 6409 | -0.089 | 0.1731 | No | ||
| 124 | POLE | na | 6450 | -0.090 | 0.1741 | No | ||
| 125 | E2F4 | na | 6619 | -0.099 | 0.1626 | No | ||
| 126 | E2F3 | na | 6627 | -0.099 | 0.1674 | No | ||
| 127 | BCL3 | na | 6654 | -0.100 | 0.1703 | No | ||
| 128 | HSPA8 | na | 6822 | -0.108 | 0.1594 | No | ||
| 129 | UBE2S | na | 6825 | -0.108 | 0.1651 | No | ||
| 130 | CDKN1B | na | 6829 | -0.108 | 0.1708 | No | ||
| 131 | KPNA2 | na | 6955 | -0.114 | 0.1645 | No | ||
| 132 | ESPL1 | na | 7039 | -0.119 | 0.1627 | No | ||
| 133 | UCK2 | na | 7132 | -0.123 | 0.1602 | No | ||
| 134 | NUP98 | na | 7295 | -0.131 | 0.1511 | No | ||
| 135 | SMAD3 | na | 7616 | -0.147 | 0.1270 | No | ||
| 136 | SLC7A5 | na | 7670 | -0.150 | 0.1300 | No | ||
| 137 | RASAL2 | na | 8407 | -0.193 | 0.0667 | No | ||
| 138 | CDK4 | na | 8522 | -0.201 | 0.0663 | No | ||
| 139 | TNPO2 | na | 8596 | -0.206 | 0.0703 | No | ||
| 140 | CUL1 | na | 8893 | -0.226 | 0.0529 | No | ||
| 141 | LIG3 | na | 9204 | -0.250 | 0.0355 | No | ||
| 142 | CCNT1 | na | 9301 | -0.259 | 0.0401 | No | ||
| 143 | ARID4A | na | 9639 | -0.305 | 0.0230 | No | ||
| 144 | PML | na | 10026 | -0.420 | 0.0073 | No |