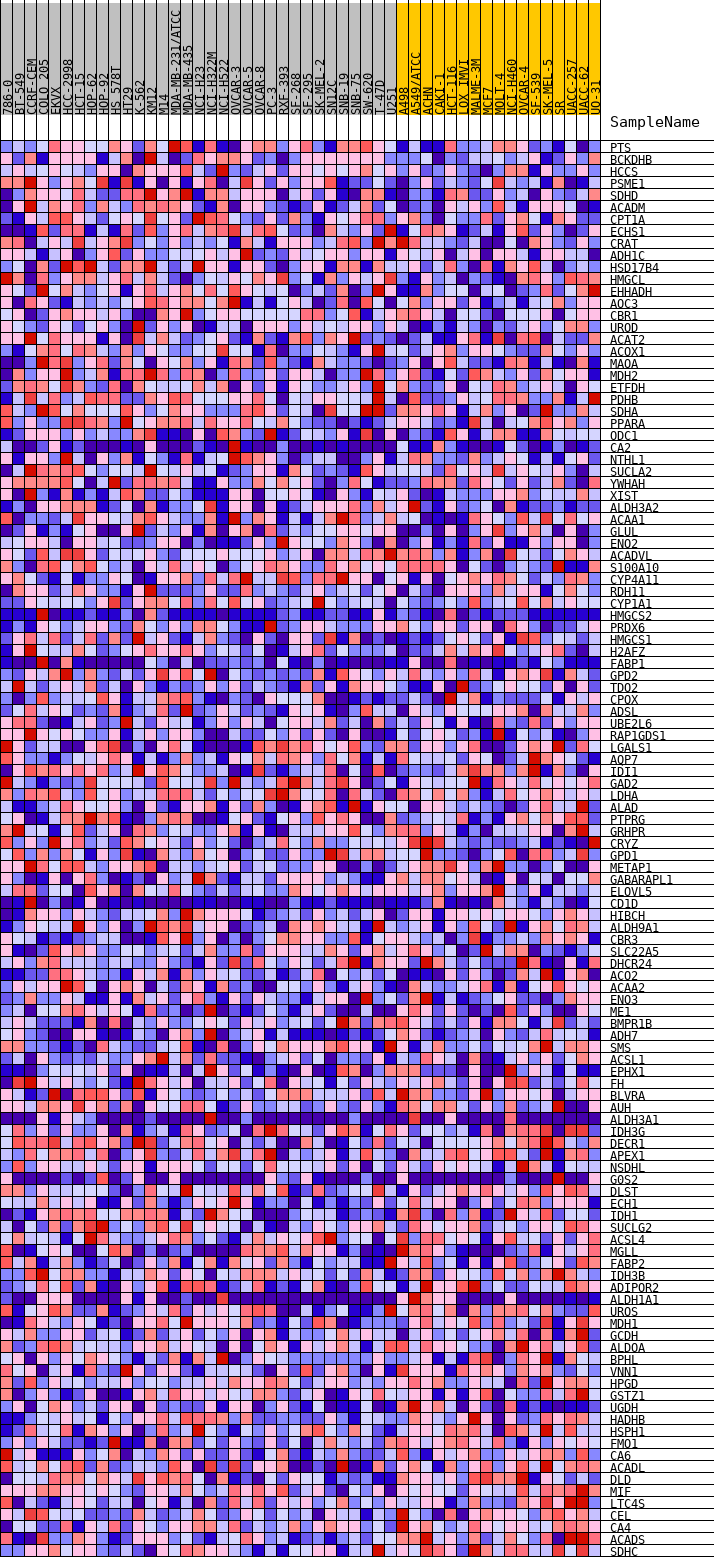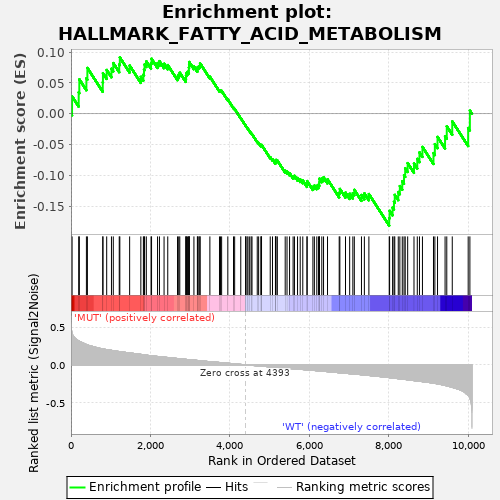
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_FATTY_ACID_METABOLISM |
| Enrichment Score (ES) | -0.18113494 |
| Normalized Enrichment Score (NES) | -0.72004426 |
| Nominal p-value | 0.8974359 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PTS | na | 27 | 0.424 | 0.0282 | No | ||
| 2 | BCKDHB | na | 194 | 0.317 | 0.0347 | No | ||
| 3 | HCCS | na | 210 | 0.310 | 0.0558 | No | ||
| 4 | PSME1 | na | 388 | 0.268 | 0.0576 | No | ||
| 5 | SDHD | na | 413 | 0.262 | 0.0743 | No | ||
| 6 | ACADM | na | 800 | 0.209 | 0.0509 | No | ||
| 7 | CPT1A | na | 805 | 0.209 | 0.0657 | No | ||
| 8 | ECHS1 | na | 899 | 0.200 | 0.0710 | No | ||
| 9 | CRAT | na | 1019 | 0.191 | 0.0730 | No | ||
| 10 | ADH1C | na | 1065 | 0.187 | 0.0822 | No | ||
| 11 | HSD17B4 | na | 1215 | 0.175 | 0.0800 | No | ||
| 12 | HMGCL | na | 1230 | 0.174 | 0.0913 | No | ||
| 13 | EHHADH | na | 1477 | 0.157 | 0.0781 | No | ||
| 14 | AOC3 | na | 1759 | 0.138 | 0.0600 | No | ||
| 15 | CBR1 | na | 1823 | 0.134 | 0.0635 | No | ||
| 16 | UROD | na | 1839 | 0.133 | 0.0716 | No | ||
| 17 | ACAT2 | na | 1853 | 0.132 | 0.0799 | No | ||
| 18 | ACOX1 | na | 1897 | 0.128 | 0.0849 | No | ||
| 19 | MAOA | na | 2016 | 0.119 | 0.0818 | No | ||
| 20 | MDH2 | na | 2029 | 0.118 | 0.0892 | No | ||
| 21 | ETFDH | na | 2180 | 0.110 | 0.0821 | No | ||
| 22 | PDHB | na | 2228 | 0.108 | 0.0853 | No | ||
| 23 | SDHA | na | 2344 | 0.101 | 0.0811 | No | ||
| 24 | PPARA | na | 2437 | 0.096 | 0.0789 | No | ||
| 25 | ODC1 | na | 2683 | 0.083 | 0.0604 | No | ||
| 26 | CA2 | na | 2713 | 0.081 | 0.0634 | No | ||
| 27 | NTHL1 | na | 2740 | 0.080 | 0.0666 | No | ||
| 28 | SUCLA2 | na | 2891 | 0.072 | 0.0568 | No | ||
| 29 | YWHAH | na | 2897 | 0.071 | 0.0615 | No | ||
| 30 | XIST | na | 2907 | 0.071 | 0.0658 | No | ||
| 31 | ALDH3A2 | na | 2936 | 0.069 | 0.0680 | No | ||
| 32 | ACAA1 | na | 2968 | 0.068 | 0.0699 | No | ||
| 33 | GLUL | na | 2969 | 0.068 | 0.0748 | No | ||
| 34 | ENO2 | na | 2975 | 0.067 | 0.0792 | No | ||
| 35 | ACADVL | na | 2977 | 0.067 | 0.0840 | No | ||
| 36 | S100A10 | na | 3096 | 0.061 | 0.0766 | No | ||
| 37 | CYP4A11 | na | 3183 | 0.057 | 0.0722 | No | ||
| 38 | RDH11 | na | 3191 | 0.057 | 0.0756 | No | ||
| 39 | CYP1A1 | na | 3220 | 0.055 | 0.0768 | No | ||
| 40 | HMGCS2 | na | 3246 | 0.054 | 0.0782 | No | ||
| 41 | PRDX6 | na | 3251 | 0.054 | 0.0818 | No | ||
| 42 | HMGCS1 | na | 3498 | 0.041 | 0.0601 | No | ||
| 43 | H2AFZ | na | 3740 | 0.030 | 0.0382 | No | ||
| 44 | FABP1 | na | 3766 | 0.029 | 0.0378 | No | ||
| 45 | GPD2 | na | 3794 | 0.028 | 0.0371 | No | ||
| 46 | TDO2 | na | 3950 | 0.021 | 0.0231 | No | ||
| 47 | CPOX | na | 4093 | 0.013 | 0.0098 | No | ||
| 48 | ADSL | na | 4122 | 0.012 | 0.0078 | No | ||
| 49 | UBE2L6 | na | 4276 | 0.005 | -0.0072 | No | ||
| 50 | RAP1GDS1 | na | 4399 | -0.000 | -0.0194 | No | ||
| 51 | LGALS1 | na | 4400 | -0.001 | -0.0193 | No | ||
| 52 | AQP7 | na | 4434 | -0.002 | -0.0225 | No | ||
| 53 | IDI1 | na | 4458 | -0.003 | -0.0246 | No | ||
| 54 | GAD2 | na | 4500 | -0.005 | -0.0283 | No | ||
| 55 | LDHA | na | 4537 | -0.007 | -0.0314 | No | ||
| 56 | ALAD | na | 4559 | -0.008 | -0.0330 | No | ||
| 57 | PTPRG | na | 4691 | -0.014 | -0.0451 | No | ||
| 58 | GRHPR | na | 4726 | -0.015 | -0.0474 | No | ||
| 59 | CRYZ | na | 4785 | -0.018 | -0.0518 | No | ||
| 60 | GPD1 | na | 4789 | -0.018 | -0.0508 | No | ||
| 61 | METAP1 | na | 4799 | -0.019 | -0.0503 | No | ||
| 62 | GABARAPL1 | na | 5018 | -0.027 | -0.0702 | No | ||
| 63 | ELOVL5 | na | 5075 | -0.029 | -0.0737 | No | ||
| 64 | CD1D | na | 5149 | -0.033 | -0.0786 | No | ||
| 65 | HIBCH | na | 5158 | -0.033 | -0.0770 | No | ||
| 66 | ALDH9A1 | na | 5160 | -0.033 | -0.0746 | No | ||
| 67 | CBR3 | na | 5197 | -0.035 | -0.0757 | No | ||
| 68 | SLC22A5 | na | 5395 | -0.044 | -0.0922 | No | ||
| 69 | DHCR24 | na | 5441 | -0.046 | -0.0934 | No | ||
| 70 | ACO2 | na | 5505 | -0.049 | -0.0961 | No | ||
| 71 | ACAA2 | na | 5594 | -0.053 | -0.1011 | No | ||
| 72 | ENO3 | na | 5628 | -0.054 | -0.1004 | No | ||
| 73 | ME1 | na | 5706 | -0.057 | -0.1040 | No | ||
| 74 | BMPR1B | na | 5773 | -0.060 | -0.1062 | No | ||
| 75 | ADH7 | na | 5838 | -0.063 | -0.1080 | No | ||
| 76 | SMS | na | 5944 | -0.067 | -0.1137 | No | ||
| 77 | ACSL1 | na | 5947 | -0.067 | -0.1090 | No | ||
| 78 | EPHX1 | na | 6093 | -0.074 | -0.1181 | No | ||
| 79 | FH | na | 6130 | -0.075 | -0.1162 | No | ||
| 80 | BLVRA | na | 6191 | -0.078 | -0.1165 | No | ||
| 81 | AUH | na | 6233 | -0.080 | -0.1148 | No | ||
| 82 | ALDH3A1 | na | 6255 | -0.082 | -0.1109 | No | ||
| 83 | IDH3G | na | 6257 | -0.082 | -0.1051 | No | ||
| 84 | DECR1 | na | 6317 | -0.084 | -0.1048 | No | ||
| 85 | APEX1 | na | 6361 | -0.086 | -0.1029 | No | ||
| 86 | NSDHL | na | 6461 | -0.091 | -0.1062 | No | ||
| 87 | G0S2 | na | 6755 | -0.105 | -0.1278 | No | ||
| 88 | DLST | na | 6772 | -0.106 | -0.1217 | No | ||
| 89 | ECH1 | na | 6914 | -0.112 | -0.1276 | No | ||
| 90 | IDH1 | na | 7019 | -0.117 | -0.1295 | No | ||
| 91 | SUCLG2 | na | 7099 | -0.122 | -0.1286 | No | ||
| 92 | ACSL4 | na | 7136 | -0.123 | -0.1232 | No | ||
| 93 | MGLL | na | 7316 | -0.132 | -0.1315 | No | ||
| 94 | FABP2 | na | 7389 | -0.135 | -0.1289 | No | ||
| 95 | IDH3B | na | 7503 | -0.141 | -0.1299 | No | ||
| 96 | ADIPOR2 | na | 8015 | -0.170 | -0.1687 | Yes | ||
| 97 | ALDH1A1 | na | 8025 | -0.171 | -0.1572 | Yes | ||
| 98 | UROS | na | 8100 | -0.175 | -0.1518 | Yes | ||
| 99 | MDH1 | na | 8135 | -0.177 | -0.1424 | Yes | ||
| 100 | GCDH | na | 8155 | -0.178 | -0.1313 | Yes | ||
| 101 | ALDOA | na | 8243 | -0.184 | -0.1266 | Yes | ||
| 102 | BPHL | na | 8283 | -0.185 | -0.1170 | Yes | ||
| 103 | VNN1 | na | 8344 | -0.190 | -0.1092 | Yes | ||
| 104 | HPGD | na | 8388 | -0.192 | -0.0995 | Yes | ||
| 105 | GSTZ1 | na | 8418 | -0.194 | -0.0883 | Yes | ||
| 106 | UGDH | na | 8480 | -0.198 | -0.0799 | Yes | ||
| 107 | HADHB | na | 8639 | -0.209 | -0.0805 | Yes | ||
| 108 | HSPH1 | na | 8722 | -0.215 | -0.0730 | Yes | ||
| 109 | FMO1 | na | 8777 | -0.218 | -0.0625 | Yes | ||
| 110 | CA6 | na | 8852 | -0.223 | -0.0536 | Yes | ||
| 111 | ACADL | na | 9130 | -0.243 | -0.0637 | Yes | ||
| 112 | DLD | na | 9165 | -0.246 | -0.0492 | Yes | ||
| 113 | MIF | na | 9232 | -0.252 | -0.0374 | Yes | ||
| 114 | LTC4S | na | 9422 | -0.274 | -0.0364 | Yes | ||
| 115 | CEL | na | 9465 | -0.279 | -0.0203 | Yes | ||
| 116 | CA4 | na | 9603 | -0.298 | -0.0123 | Yes | ||
| 117 | ACADS | na | 10004 | -0.403 | -0.0230 | Yes | ||
| 118 | SDHC | na | 10047 | -0.444 | 0.0052 | Yes |