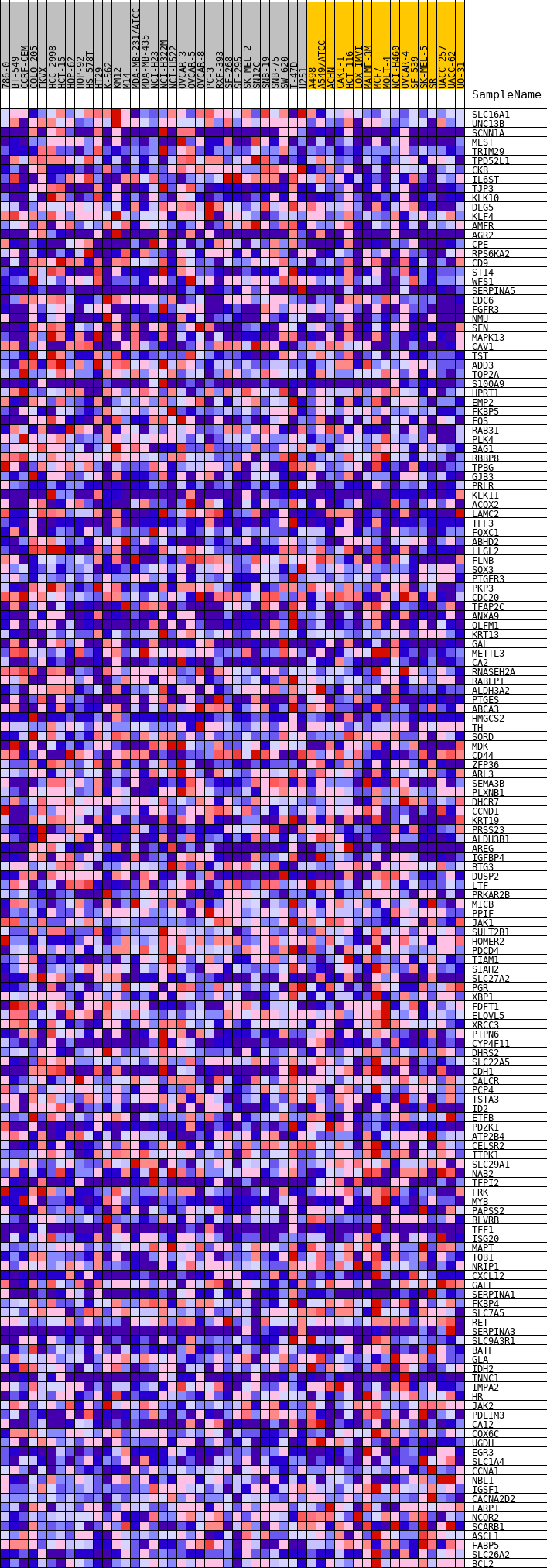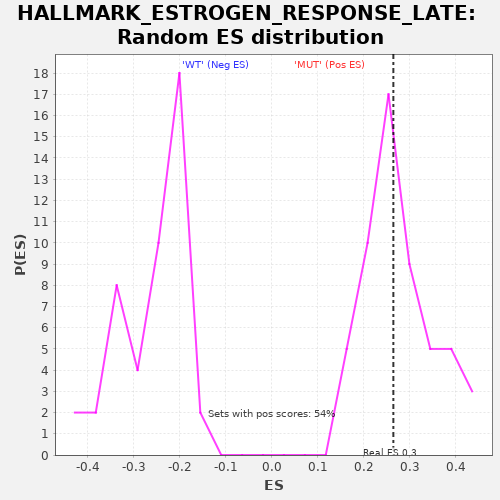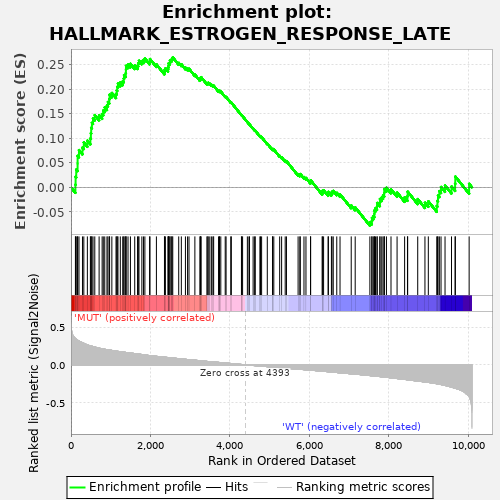
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_ESTROGEN_RESPONSE_LATE |
| Enrichment Score (ES) | 0.26437652 |
| Normalized Enrichment Score (NES) | 0.9584295 |
| Nominal p-value | 0.46296296 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLC16A1 | na | 111 | 0.347 | 0.0055 | Yes | ||
| 2 | UNC13B | na | 116 | 0.346 | 0.0216 | Yes | ||
| 3 | SCNN1A | na | 132 | 0.338 | 0.0363 | Yes | ||
| 4 | MEST | na | 166 | 0.325 | 0.0485 | Yes | ||
| 5 | TRIM29 | na | 167 | 0.325 | 0.0641 | Yes | ||
| 6 | TPD52L1 | na | 201 | 0.314 | 0.0758 | Yes | ||
| 7 | CKB | na | 289 | 0.292 | 0.0810 | Yes | ||
| 8 | IL6ST | na | 323 | 0.284 | 0.0913 | Yes | ||
| 9 | TJP3 | na | 412 | 0.262 | 0.0950 | Yes | ||
| 10 | KLK10 | na | 488 | 0.250 | 0.0995 | Yes | ||
| 11 | DLG5 | na | 501 | 0.249 | 0.1102 | Yes | ||
| 12 | KLF4 | na | 509 | 0.247 | 0.1213 | Yes | ||
| 13 | AMFR | na | 527 | 0.245 | 0.1314 | Yes | ||
| 14 | AGR2 | na | 554 | 0.241 | 0.1403 | Yes | ||
| 15 | CPE | na | 599 | 0.234 | 0.1471 | Yes | ||
| 16 | RPS6KA2 | na | 708 | 0.220 | 0.1468 | Yes | ||
| 17 | CD9 | na | 784 | 0.211 | 0.1493 | Yes | ||
| 18 | ST14 | na | 813 | 0.208 | 0.1565 | Yes | ||
| 19 | WFS1 | na | 850 | 0.205 | 0.1627 | Yes | ||
| 20 | SERPINA5 | na | 906 | 0.200 | 0.1667 | Yes | ||
| 21 | CDC6 | na | 931 | 0.198 | 0.1738 | Yes | ||
| 22 | FGFR3 | na | 965 | 0.196 | 0.1798 | Yes | ||
| 23 | NMU | na | 972 | 0.195 | 0.1886 | Yes | ||
| 24 | SFN | na | 1029 | 0.190 | 0.1920 | Yes | ||
| 25 | MAPK13 | na | 1134 | 0.182 | 0.1903 | Yes | ||
| 26 | CAV1 | na | 1152 | 0.180 | 0.1972 | Yes | ||
| 27 | TST | na | 1165 | 0.178 | 0.2045 | Yes | ||
| 28 | ADD3 | na | 1183 | 0.177 | 0.2113 | Yes | ||
| 29 | TOP2A | na | 1242 | 0.173 | 0.2138 | Yes | ||
| 30 | S100A9 | na | 1303 | 0.168 | 0.2158 | Yes | ||
| 31 | HPRT1 | na | 1329 | 0.166 | 0.2212 | Yes | ||
| 32 | EMP2 | na | 1337 | 0.166 | 0.2285 | Yes | ||
| 33 | FKBP5 | na | 1379 | 0.163 | 0.2322 | Yes | ||
| 34 | FOS | na | 1383 | 0.163 | 0.2397 | Yes | ||
| 35 | RAB31 | na | 1385 | 0.163 | 0.2474 | Yes | ||
| 36 | PLK4 | na | 1436 | 0.160 | 0.2500 | Yes | ||
| 37 | BAG1 | na | 1497 | 0.156 | 0.2514 | Yes | ||
| 38 | RBBP8 | na | 1605 | 0.148 | 0.2478 | Yes | ||
| 39 | TPBG | na | 1676 | 0.143 | 0.2476 | Yes | ||
| 40 | GJB3 | na | 1694 | 0.142 | 0.2527 | Yes | ||
| 41 | PRLR | na | 1712 | 0.141 | 0.2577 | Yes | ||
| 42 | KLK11 | na | 1785 | 0.136 | 0.2570 | Yes | ||
| 43 | ACOX2 | na | 1827 | 0.133 | 0.2592 | Yes | ||
| 44 | LAMC2 | na | 1861 | 0.130 | 0.2622 | Yes | ||
| 45 | TFF3 | na | 1980 | 0.121 | 0.2561 | Yes | ||
| 46 | FOXC1 | na | 1989 | 0.121 | 0.2611 | Yes | ||
| 47 | ABHD2 | na | 2151 | 0.111 | 0.2502 | Yes | ||
| 48 | LLGL2 | na | 2354 | 0.101 | 0.2347 | Yes | ||
| 49 | FLNB | na | 2359 | 0.100 | 0.2391 | Yes | ||
| 50 | SOX3 | na | 2377 | 0.099 | 0.2422 | Yes | ||
| 51 | PTGER3 | na | 2444 | 0.096 | 0.2401 | Yes | ||
| 52 | PKP3 | na | 2449 | 0.095 | 0.2443 | Yes | ||
| 53 | CDC20 | na | 2457 | 0.095 | 0.2481 | Yes | ||
| 54 | TFAP2C | na | 2460 | 0.095 | 0.2524 | Yes | ||
| 55 | ANXA9 | na | 2490 | 0.093 | 0.2540 | Yes | ||
| 56 | OLFM1 | na | 2492 | 0.093 | 0.2583 | Yes | ||
| 57 | KRT13 | na | 2526 | 0.091 | 0.2593 | Yes | ||
| 58 | GAL | na | 2544 | 0.090 | 0.2619 | Yes | ||
| 59 | METTL3 | na | 2563 | 0.089 | 0.2644 | Yes | ||
| 60 | CA2 | na | 2713 | 0.081 | 0.2533 | No | ||
| 61 | RNASEH2A | na | 2779 | 0.077 | 0.2504 | No | ||
| 62 | RABEP1 | na | 2882 | 0.072 | 0.2436 | No | ||
| 63 | ALDH3A2 | na | 2936 | 0.069 | 0.2416 | No | ||
| 64 | PTGES | na | 2972 | 0.067 | 0.2413 | No | ||
| 65 | ABCA3 | na | 3120 | 0.060 | 0.2294 | No | ||
| 66 | HMGCS2 | na | 3246 | 0.054 | 0.2194 | No | ||
| 67 | TH | na | 3261 | 0.053 | 0.2205 | No | ||
| 68 | SORD | na | 3263 | 0.053 | 0.2230 | No | ||
| 69 | MDK | na | 3278 | 0.052 | 0.2241 | No | ||
| 70 | CD44 | na | 3423 | 0.045 | 0.2118 | No | ||
| 71 | ZFP36 | na | 3446 | 0.043 | 0.2116 | No | ||
| 72 | ARL3 | na | 3455 | 0.043 | 0.2129 | No | ||
| 73 | SEMA3B | na | 3496 | 0.041 | 0.2108 | No | ||
| 74 | PLXNB1 | na | 3536 | 0.039 | 0.2088 | No | ||
| 75 | DHCR7 | na | 3574 | 0.038 | 0.2069 | No | ||
| 76 | CCND1 | na | 3593 | 0.037 | 0.2068 | No | ||
| 77 | KRT19 | na | 3716 | 0.031 | 0.1961 | No | ||
| 78 | PRSS23 | na | 3739 | 0.030 | 0.1953 | No | ||
| 79 | ALDH3B1 | na | 3741 | 0.030 | 0.1966 | No | ||
| 80 | AREG | na | 3773 | 0.029 | 0.1949 | No | ||
| 81 | IGFBP4 | na | 3884 | 0.024 | 0.1849 | No | ||
| 82 | BTG3 | na | 3906 | 0.023 | 0.1839 | No | ||
| 83 | DUSP2 | na | 4024 | 0.017 | 0.1730 | No | ||
| 84 | LTF | na | 4038 | 0.016 | 0.1724 | No | ||
| 85 | PRKAR2B | na | 4293 | 0.004 | 0.1471 | No | ||
| 86 | MICB | na | 4301 | 0.004 | 0.1465 | No | ||
| 87 | PPIF | na | 4322 | 0.003 | 0.1447 | No | ||
| 88 | JAK1 | na | 4442 | -0.002 | 0.1328 | No | ||
| 89 | SULT2B1 | na | 4482 | -0.004 | 0.1291 | No | ||
| 90 | HOMER2 | na | 4496 | -0.005 | 0.1280 | No | ||
| 91 | PDCD4 | na | 4587 | -0.009 | 0.1194 | No | ||
| 92 | TIAM1 | na | 4625 | -0.011 | 0.1162 | No | ||
| 93 | SIAH2 | na | 4641 | -0.011 | 0.1152 | No | ||
| 94 | SLC27A2 | na | 4758 | -0.017 | 0.1044 | No | ||
| 95 | PGR | na | 4773 | -0.018 | 0.1038 | No | ||
| 96 | XBP1 | na | 4801 | -0.019 | 0.1020 | No | ||
| 97 | FDFT1 | na | 4944 | -0.024 | 0.0889 | No | ||
| 98 | ELOVL5 | na | 5075 | -0.029 | 0.0772 | No | ||
| 99 | XRCC3 | na | 5079 | -0.030 | 0.0783 | No | ||
| 100 | PTPN6 | na | 5108 | -0.031 | 0.0770 | No | ||
| 101 | CYP4F11 | na | 5256 | -0.038 | 0.0640 | No | ||
| 102 | DHRS2 | na | 5305 | -0.040 | 0.0611 | No | ||
| 103 | SLC22A5 | na | 5395 | -0.044 | 0.0542 | No | ||
| 104 | CDH1 | na | 5427 | -0.045 | 0.0533 | No | ||
| 105 | CALCR | na | 5720 | -0.058 | 0.0267 | No | ||
| 106 | PCP4 | na | 5764 | -0.059 | 0.0252 | No | ||
| 107 | TSTA3 | na | 5775 | -0.060 | 0.0271 | No | ||
| 108 | ID2 | na | 5868 | -0.064 | 0.0209 | No | ||
| 109 | ETFB | na | 5918 | -0.066 | 0.0191 | No | ||
| 110 | PDZK1 | na | 6033 | -0.072 | 0.0111 | No | ||
| 111 | ATP2B4 | na | 6036 | -0.072 | 0.0143 | No | ||
| 112 | CELSR2 | na | 6328 | -0.085 | -0.0109 | No | ||
| 113 | ITPK1 | na | 6331 | -0.085 | -0.0070 | No | ||
| 114 | SLC29A1 | na | 6362 | -0.086 | -0.0059 | No | ||
| 115 | NAB2 | na | 6476 | -0.091 | -0.0129 | No | ||
| 116 | TFPI2 | na | 6483 | -0.092 | -0.0091 | No | ||
| 117 | FRK | na | 6559 | -0.096 | -0.0121 | No | ||
| 118 | MYB | na | 6573 | -0.096 | -0.0088 | No | ||
| 119 | PAPSS2 | na | 6605 | -0.098 | -0.0072 | No | ||
| 120 | BLVRB | na | 6694 | -0.102 | -0.0112 | No | ||
| 121 | TFF1 | na | 6776 | -0.106 | -0.0142 | No | ||
| 122 | ISG20 | na | 7058 | -0.120 | -0.0367 | No | ||
| 123 | MAPT | na | 7158 | -0.124 | -0.0407 | No | ||
| 124 | TOB1 | na | 7528 | -0.143 | -0.0710 | No | ||
| 125 | NRIP1 | na | 7576 | -0.146 | -0.0688 | No | ||
| 126 | CXCL12 | na | 7581 | -0.146 | -0.0622 | No | ||
| 127 | GALE | na | 7614 | -0.147 | -0.0584 | No | ||
| 128 | SERPINA1 | na | 7641 | -0.148 | -0.0539 | No | ||
| 129 | FKBP4 | na | 7643 | -0.149 | -0.0469 | No | ||
| 130 | SLC7A5 | na | 7670 | -0.150 | -0.0423 | No | ||
| 131 | RET | na | 7708 | -0.152 | -0.0387 | No | ||
| 132 | SERPINA3 | na | 7712 | -0.152 | -0.0317 | No | ||
| 133 | SLC9A3R1 | na | 7780 | -0.156 | -0.0310 | No | ||
| 134 | BATF | na | 7782 | -0.156 | -0.0236 | No | ||
| 135 | GLA | na | 7827 | -0.158 | -0.0205 | No | ||
| 136 | IDH2 | na | 7860 | -0.160 | -0.0160 | No | ||
| 137 | TNNC1 | na | 7890 | -0.162 | -0.0112 | No | ||
| 138 | IMPA2 | na | 7892 | -0.162 | -0.0035 | No | ||
| 139 | HR | na | 7945 | -0.166 | -0.0008 | No | ||
| 140 | JAK2 | na | 8060 | -0.173 | -0.0040 | No | ||
| 141 | PDLIM3 | na | 8213 | -0.182 | -0.0106 | No | ||
| 142 | CA12 | na | 8403 | -0.193 | -0.0203 | No | ||
| 143 | COX6C | na | 8475 | -0.198 | -0.0180 | No | ||
| 144 | UGDH | na | 8480 | -0.198 | -0.0089 | No | ||
| 145 | EGR3 | na | 8733 | -0.216 | -0.0239 | No | ||
| 146 | SLC1A4 | na | 8915 | -0.227 | -0.0312 | No | ||
| 147 | CCNA1 | na | 9000 | -0.233 | -0.0285 | No | ||
| 148 | NBL1 | na | 9213 | -0.250 | -0.0378 | No | ||
| 149 | IGSF1 | na | 9230 | -0.251 | -0.0274 | No | ||
| 150 | CACNA2D2 | na | 9244 | -0.253 | -0.0166 | No | ||
| 151 | FARP1 | na | 9278 | -0.257 | -0.0076 | No | ||
| 152 | NCOR2 | na | 9320 | -0.262 | 0.0008 | No | ||
| 153 | SCARB1 | na | 9421 | -0.274 | 0.0038 | No | ||
| 154 | ASCL1 | na | 9584 | -0.295 | 0.0016 | No | ||
| 155 | FABP5 | na | 9675 | -0.310 | 0.0075 | No | ||
| 156 | SLC26A2 | na | 9680 | -0.311 | 0.0219 | No | ||
| 157 | BCL2 | na | 10028 | -0.420 | 0.0071 | No |