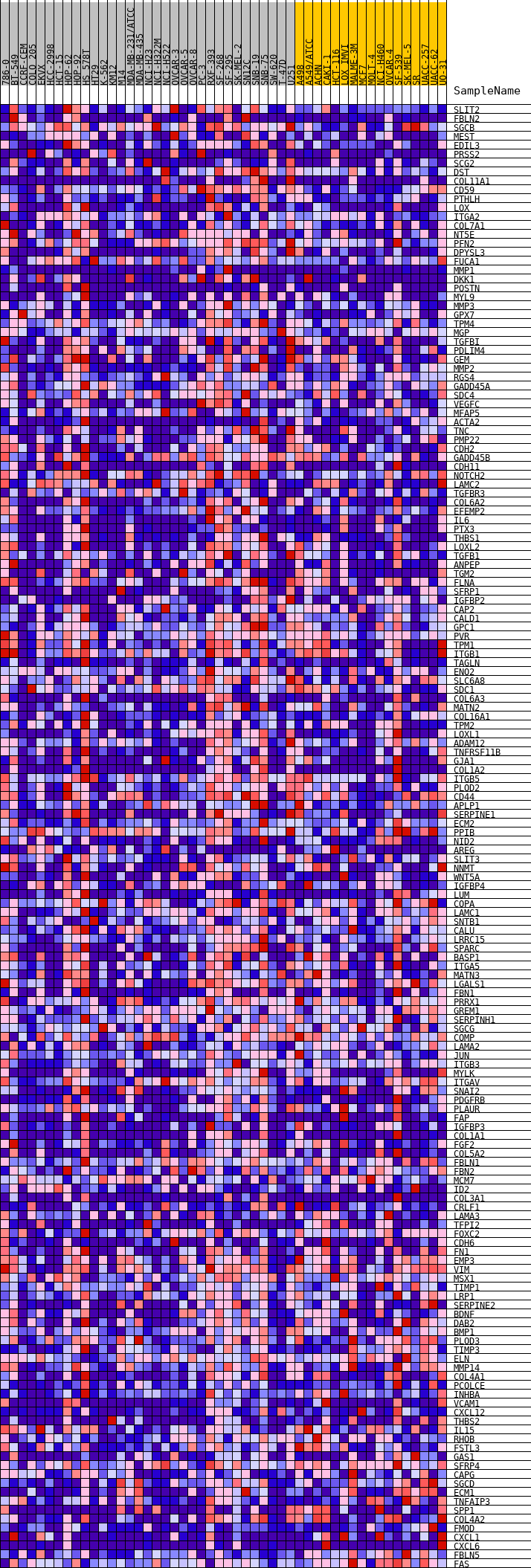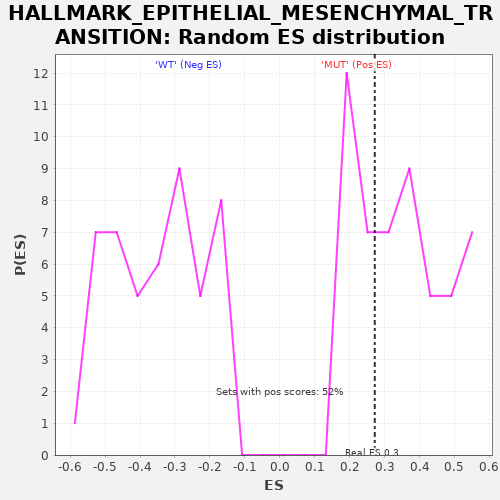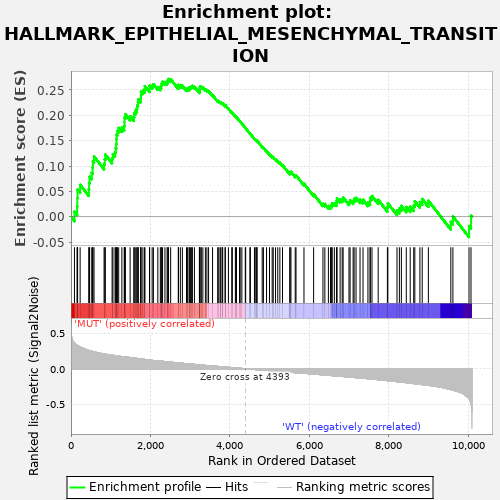
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION |
| Enrichment Score (ES) | 0.2717964 |
| Normalized Enrichment Score (NES) | 0.77809626 |
| Nominal p-value | 0.63461536 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLIT2 | na | 87 | 0.360 | 0.0100 | Yes | ||
| 2 | FBLN2 | na | 155 | 0.328 | 0.0204 | Yes | ||
| 3 | SGCB | na | 159 | 0.326 | 0.0371 | Yes | ||
| 4 | MEST | na | 166 | 0.325 | 0.0535 | Yes | ||
| 5 | EDIL3 | na | 230 | 0.305 | 0.0630 | Yes | ||
| 6 | PRSS2 | na | 449 | 0.256 | 0.0545 | Yes | ||
| 7 | SCG2 | na | 457 | 0.255 | 0.0671 | Yes | ||
| 8 | DST | na | 469 | 0.253 | 0.0792 | Yes | ||
| 9 | COL11A1 | na | 523 | 0.246 | 0.0867 | Yes | ||
| 10 | CD59 | na | 544 | 0.243 | 0.0973 | Yes | ||
| 11 | PTHLH | na | 551 | 0.241 | 0.1093 | Yes | ||
| 12 | LOX | na | 580 | 0.237 | 0.1189 | Yes | ||
| 13 | ITGA2 | na | 833 | 0.206 | 0.1043 | Yes | ||
| 14 | COL7A1 | na | 851 | 0.205 | 0.1133 | Yes | ||
| 15 | NT5E | na | 864 | 0.203 | 0.1227 | Yes | ||
| 16 | PFN2 | na | 1035 | 0.189 | 0.1155 | Yes | ||
| 17 | DPYSL3 | na | 1056 | 0.188 | 0.1232 | Yes | ||
| 18 | FUCA1 | na | 1109 | 0.184 | 0.1276 | Yes | ||
| 19 | MMP1 | na | 1127 | 0.183 | 0.1354 | Yes | ||
| 20 | DKK1 | na | 1140 | 0.181 | 0.1437 | Yes | ||
| 21 | POSTN | na | 1144 | 0.181 | 0.1528 | Yes | ||
| 22 | MYL9 | na | 1148 | 0.180 | 0.1619 | Yes | ||
| 23 | MMP3 | na | 1169 | 0.178 | 0.1692 | Yes | ||
| 24 | GPX7 | na | 1201 | 0.176 | 0.1752 | Yes | ||
| 25 | TPM4 | na | 1281 | 0.171 | 0.1762 | Yes | ||
| 26 | MGP | na | 1340 | 0.166 | 0.1790 | Yes | ||
| 27 | TGFBI | na | 1348 | 0.165 | 0.1869 | Yes | ||
| 28 | PDLIM4 | na | 1349 | 0.165 | 0.1955 | Yes | ||
| 29 | GEM | na | 1370 | 0.164 | 0.2021 | Yes | ||
| 30 | MMP2 | na | 1489 | 0.156 | 0.1984 | Yes | ||
| 31 | RGS4 | na | 1582 | 0.150 | 0.1969 | Yes | ||
| 32 | GADD45A | na | 1592 | 0.149 | 0.2038 | Yes | ||
| 33 | SDC4 | na | 1619 | 0.147 | 0.2089 | Yes | ||
| 34 | VEGFC | na | 1656 | 0.145 | 0.2128 | Yes | ||
| 35 | MFAP5 | na | 1666 | 0.144 | 0.2195 | Yes | ||
| 36 | ACTA2 | na | 1688 | 0.143 | 0.2248 | Yes | ||
| 37 | TNC | na | 1695 | 0.142 | 0.2316 | Yes | ||
| 38 | PMP22 | na | 1754 | 0.138 | 0.2329 | Yes | ||
| 39 | CDH2 | na | 1761 | 0.137 | 0.2395 | Yes | ||
| 40 | GADD45B | na | 1763 | 0.137 | 0.2466 | Yes | ||
| 41 | CDH11 | na | 1811 | 0.135 | 0.2489 | Yes | ||
| 42 | NOTCH2 | na | 1854 | 0.132 | 0.2515 | Yes | ||
| 43 | LAMC2 | na | 1861 | 0.130 | 0.2577 | Yes | ||
| 44 | TGFBR3 | na | 1978 | 0.122 | 0.2524 | Yes | ||
| 45 | COL6A2 | na | 1981 | 0.121 | 0.2585 | Yes | ||
| 46 | EFEMP2 | na | 2044 | 0.117 | 0.2584 | Yes | ||
| 47 | IL6 | na | 2075 | 0.115 | 0.2614 | Yes | ||
| 48 | PTX3 | na | 2187 | 0.110 | 0.2559 | Yes | ||
| 49 | THBS1 | na | 2250 | 0.106 | 0.2552 | Yes | ||
| 50 | LOXL2 | na | 2271 | 0.105 | 0.2587 | Yes | ||
| 51 | TGFB1 | na | 2285 | 0.105 | 0.2629 | Yes | ||
| 52 | ANPEP | na | 2306 | 0.103 | 0.2662 | Yes | ||
| 53 | TGM2 | na | 2366 | 0.100 | 0.2655 | Yes | ||
| 54 | FLNA | na | 2420 | 0.097 | 0.2653 | Yes | ||
| 55 | SFRP1 | na | 2436 | 0.096 | 0.2688 | Yes | ||
| 56 | IGFBP2 | na | 2456 | 0.095 | 0.2718 | Yes | ||
| 57 | CAP2 | na | 2510 | 0.092 | 0.2712 | No | ||
| 58 | CALD1 | na | 2703 | 0.082 | 0.2562 | No | ||
| 59 | GPC1 | na | 2706 | 0.081 | 0.2602 | No | ||
| 60 | PVR | na | 2754 | 0.079 | 0.2596 | No | ||
| 61 | TPM1 | na | 2805 | 0.076 | 0.2585 | No | ||
| 62 | ITGB1 | na | 2909 | 0.071 | 0.2519 | No | ||
| 63 | TAGLN | na | 2932 | 0.069 | 0.2533 | No | ||
| 64 | ENO2 | na | 2975 | 0.067 | 0.2525 | No | ||
| 65 | SLC6A8 | na | 2981 | 0.067 | 0.2555 | No | ||
| 66 | SDC1 | na | 3012 | 0.066 | 0.2559 | No | ||
| 67 | COL6A3 | na | 3038 | 0.064 | 0.2568 | No | ||
| 68 | MATN2 | na | 3058 | 0.064 | 0.2582 | No | ||
| 69 | COL16A1 | na | 3109 | 0.061 | 0.2563 | No | ||
| 70 | TPM2 | na | 3234 | 0.054 | 0.2467 | No | ||
| 71 | LOXL1 | na | 3236 | 0.054 | 0.2494 | No | ||
| 72 | ADAM12 | na | 3243 | 0.054 | 0.2516 | No | ||
| 73 | TNFRSF11B | na | 3245 | 0.054 | 0.2544 | No | ||
| 74 | GJA1 | na | 3249 | 0.054 | 0.2569 | No | ||
| 75 | COL1A2 | na | 3283 | 0.052 | 0.2563 | No | ||
| 76 | ITGB5 | na | 3321 | 0.050 | 0.2551 | No | ||
| 77 | PLOD2 | na | 3385 | 0.046 | 0.2512 | No | ||
| 78 | CD44 | na | 3423 | 0.045 | 0.2498 | No | ||
| 79 | APLP1 | na | 3465 | 0.042 | 0.2479 | No | ||
| 80 | SERPINE1 | na | 3565 | 0.038 | 0.2399 | No | ||
| 81 | ECM2 | na | 3694 | 0.032 | 0.2287 | No | ||
| 82 | PPIB | na | 3721 | 0.031 | 0.2278 | No | ||
| 83 | NID2 | na | 3756 | 0.029 | 0.2259 | No | ||
| 84 | AREG | na | 3773 | 0.029 | 0.2257 | No | ||
| 85 | SLIT3 | na | 3807 | 0.027 | 0.2238 | No | ||
| 86 | NNMT | na | 3821 | 0.026 | 0.2239 | No | ||
| 87 | WNT5A | na | 3883 | 0.024 | 0.2190 | No | ||
| 88 | IGFBP4 | na | 3884 | 0.024 | 0.2202 | No | ||
| 89 | LUM | na | 3965 | 0.020 | 0.2132 | No | ||
| 90 | COPA | na | 4050 | 0.015 | 0.2055 | No | ||
| 91 | LAMC1 | na | 4063 | 0.014 | 0.2051 | No | ||
| 92 | SNTB1 | na | 4140 | 0.011 | 0.1980 | No | ||
| 93 | CALU | na | 4156 | 0.010 | 0.1970 | No | ||
| 94 | LRRC15 | na | 4168 | 0.010 | 0.1964 | No | ||
| 95 | SPARC | na | 4245 | 0.006 | 0.1891 | No | ||
| 96 | BASP1 | na | 4261 | 0.005 | 0.1879 | No | ||
| 97 | ITGA5 | na | 4303 | 0.004 | 0.1840 | No | ||
| 98 | MATN3 | na | 4390 | 0.000 | 0.1753 | No | ||
| 99 | LGALS1 | na | 4400 | -0.001 | 0.1744 | No | ||
| 100 | FBN1 | na | 4506 | -0.005 | 0.1641 | No | ||
| 101 | PRRX1 | na | 4518 | -0.006 | 0.1633 | No | ||
| 102 | GREM1 | na | 4621 | -0.010 | 0.1536 | No | ||
| 103 | SERPINH1 | na | 4637 | -0.011 | 0.1527 | No | ||
| 104 | SGCG | na | 4665 | -0.012 | 0.1506 | No | ||
| 105 | COMP | na | 4679 | -0.013 | 0.1500 | No | ||
| 106 | LAMA2 | na | 4687 | -0.013 | 0.1500 | No | ||
| 107 | JUN | na | 4816 | -0.019 | 0.1381 | No | ||
| 108 | ITGB3 | na | 4846 | -0.021 | 0.1363 | No | ||
| 109 | MYLK | na | 4925 | -0.023 | 0.1296 | No | ||
| 110 | ITGAV | na | 4999 | -0.026 | 0.1237 | No | ||
| 111 | SNAI2 | na | 5067 | -0.029 | 0.1184 | No | ||
| 112 | PDGFRB | na | 5093 | -0.030 | 0.1175 | No | ||
| 113 | PLAUR | na | 5152 | -0.033 | 0.1134 | No | ||
| 114 | FAP | na | 5208 | -0.036 | 0.1097 | No | ||
| 115 | IGFBP3 | na | 5261 | -0.038 | 0.1065 | No | ||
| 116 | COL1A1 | na | 5327 | -0.041 | 0.1020 | No | ||
| 117 | FGF2 | na | 5511 | -0.049 | 0.0862 | No | ||
| 118 | COL5A2 | na | 5516 | -0.049 | 0.0884 | No | ||
| 119 | FBLN1 | na | 5540 | -0.050 | 0.0887 | No | ||
| 120 | FBN2 | na | 5647 | -0.055 | 0.0809 | No | ||
| 121 | MCM7 | na | 5667 | -0.056 | 0.0819 | No | ||
| 122 | ID2 | na | 5868 | -0.064 | 0.0651 | No | ||
| 123 | COL3A1 | na | 6111 | -0.075 | 0.0446 | No | ||
| 124 | CRLF1 | na | 6346 | -0.085 | 0.0255 | No | ||
| 125 | LAMA3 | na | 6394 | -0.088 | 0.0254 | No | ||
| 126 | TFPI2 | na | 6483 | -0.092 | 0.0213 | No | ||
| 127 | FOXC2 | na | 6537 | -0.095 | 0.0209 | No | ||
| 128 | CDH6 | na | 6564 | -0.096 | 0.0233 | No | ||
| 129 | FN1 | na | 6579 | -0.097 | 0.0269 | No | ||
| 130 | EMP3 | na | 6633 | -0.099 | 0.0268 | No | ||
| 131 | VIM | na | 6687 | -0.101 | 0.0268 | No | ||
| 132 | MSX1 | na | 6692 | -0.102 | 0.0317 | No | ||
| 133 | TIMP1 | na | 6699 | -0.102 | 0.0364 | No | ||
| 134 | LRP1 | na | 6778 | -0.106 | 0.0341 | No | ||
| 135 | SERPINE2 | na | 6830 | -0.108 | 0.0346 | No | ||
| 136 | BDNF | na | 6854 | -0.109 | 0.0380 | No | ||
| 137 | DAB2 | na | 7002 | -0.117 | 0.0293 | No | ||
| 138 | BMP1 | na | 7029 | -0.118 | 0.0328 | No | ||
| 139 | PLOD3 | na | 7106 | -0.122 | 0.0315 | No | ||
| 140 | TIMP3 | na | 7131 | -0.123 | 0.0355 | No | ||
| 141 | ELN | na | 7179 | -0.125 | 0.0373 | No | ||
| 142 | MMP14 | na | 7280 | -0.130 | 0.0340 | No | ||
| 143 | COL4A1 | na | 7355 | -0.133 | 0.0335 | No | ||
| 144 | PCOLCE | na | 7479 | -0.140 | 0.0284 | No | ||
| 145 | INHBA | na | 7533 | -0.143 | 0.0306 | No | ||
| 146 | VCAM1 | na | 7537 | -0.143 | 0.0377 | No | ||
| 147 | CXCL12 | na | 7581 | -0.146 | 0.0410 | No | ||
| 148 | THBS2 | na | 7737 | -0.154 | 0.0334 | No | ||
| 149 | IL15 | na | 7969 | -0.167 | 0.0189 | No | ||
| 150 | RHOB | na | 7981 | -0.168 | 0.0266 | No | ||
| 151 | FSTL3 | na | 8211 | -0.182 | 0.0130 | No | ||
| 152 | GAS1 | na | 8271 | -0.185 | 0.0167 | No | ||
| 153 | SFRP4 | na | 8321 | -0.188 | 0.0216 | No | ||
| 154 | CAPG | na | 8445 | -0.196 | 0.0194 | No | ||
| 155 | SGCD | na | 8544 | -0.203 | 0.0201 | No | ||
| 156 | ECM1 | na | 8632 | -0.209 | 0.0223 | No | ||
| 157 | TNFAIP3 | na | 8660 | -0.211 | 0.0306 | No | ||
| 158 | SPP1 | na | 8788 | -0.219 | 0.0292 | No | ||
| 159 | COL4A2 | na | 8846 | -0.223 | 0.0351 | No | ||
| 160 | FMOD | na | 9001 | -0.233 | 0.0318 | No | ||
| 161 | CXCL1 | na | 9567 | -0.292 | -0.0098 | No | ||
| 162 | CXCL6 | na | 9619 | -0.301 | 0.0007 | No | ||
| 163 | FBLN5 | na | 10025 | -0.419 | -0.0182 | No | ||
| 164 | FAS | na | 10075 | -0.489 | 0.0024 | No |