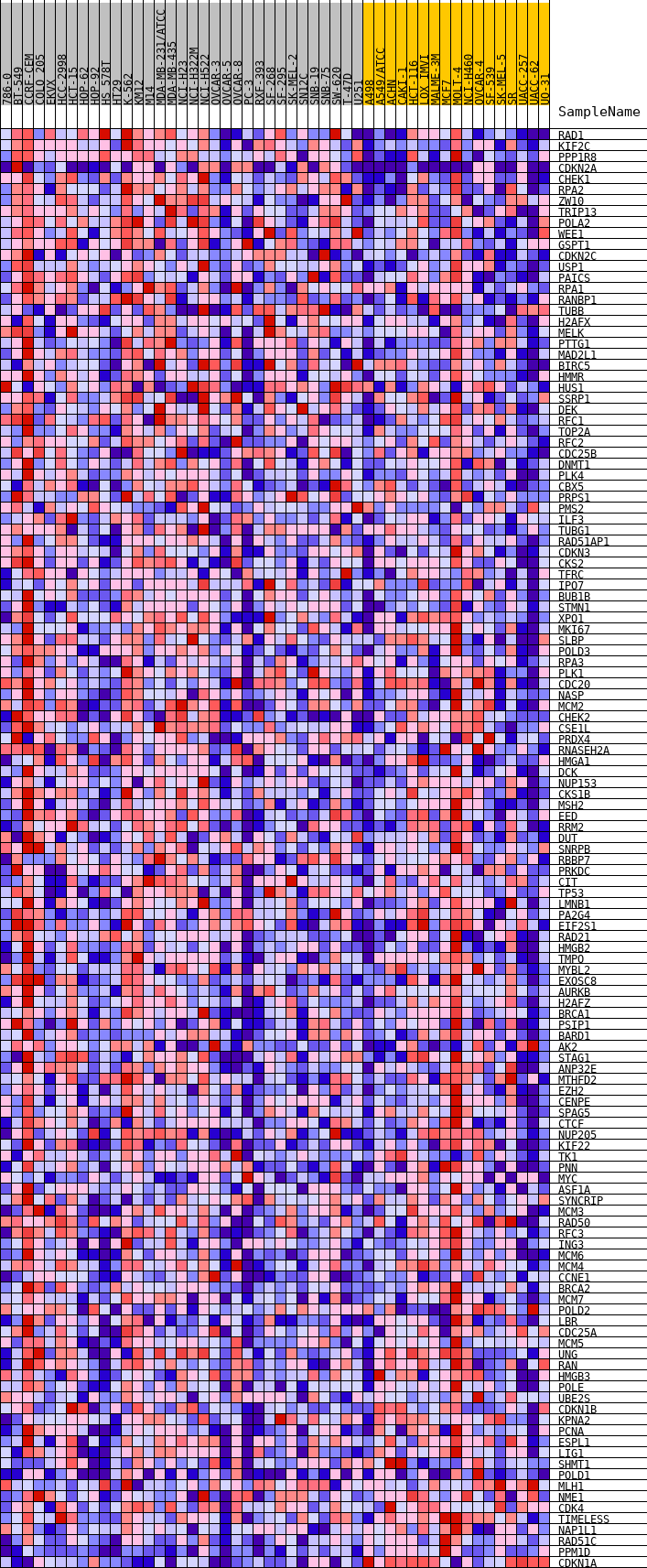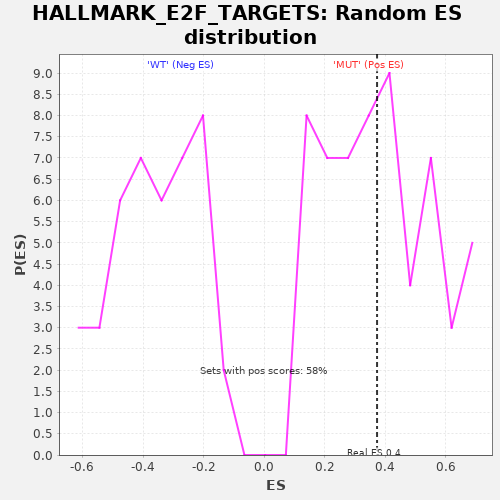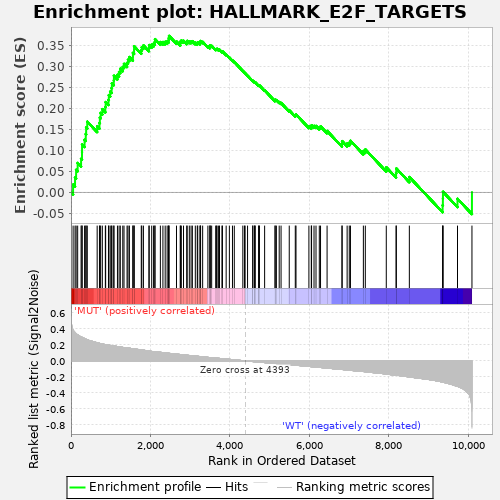
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_E2F_TARGETS |
| Enrichment Score (ES) | 0.3729696 |
| Normalized Enrichment Score (NES) | 0.97688365 |
| Nominal p-value | 0.51724136 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RAD1 | na | 53 | 0.380 | 0.0182 | Yes | ||
| 2 | KIF2C | na | 102 | 0.351 | 0.0352 | Yes | ||
| 3 | PPP1R8 | na | 129 | 0.339 | 0.0536 | Yes | ||
| 4 | CDKN2A | na | 168 | 0.324 | 0.0698 | Yes | ||
| 5 | CHEK1 | na | 254 | 0.299 | 0.0798 | Yes | ||
| 6 | RPA2 | na | 277 | 0.295 | 0.0959 | Yes | ||
| 7 | ZW10 | na | 279 | 0.294 | 0.1140 | Yes | ||
| 8 | TRIP13 | na | 342 | 0.279 | 0.1250 | Yes | ||
| 9 | POLA2 | na | 372 | 0.271 | 0.1389 | Yes | ||
| 10 | WEE1 | na | 382 | 0.270 | 0.1548 | Yes | ||
| 11 | GSPT1 | na | 410 | 0.262 | 0.1683 | Yes | ||
| 12 | CDKN2C | na | 659 | 0.226 | 0.1574 | Yes | ||
| 13 | USP1 | na | 717 | 0.219 | 0.1652 | Yes | ||
| 14 | PAICS | na | 727 | 0.218 | 0.1778 | Yes | ||
| 15 | RPA1 | na | 747 | 0.216 | 0.1893 | Yes | ||
| 16 | RANBP1 | na | 789 | 0.210 | 0.1982 | Yes | ||
| 17 | TUBB | na | 865 | 0.203 | 0.2032 | Yes | ||
| 18 | H2AFX | na | 873 | 0.203 | 0.2151 | Yes | ||
| 19 | MELK | na | 944 | 0.197 | 0.2203 | Yes | ||
| 20 | PTTG1 | na | 953 | 0.196 | 0.2316 | Yes | ||
| 21 | MAD2L1 | na | 988 | 0.193 | 0.2402 | Yes | ||
| 22 | BIRC5 | na | 1023 | 0.191 | 0.2486 | Yes | ||
| 23 | HMMR | na | 1030 | 0.190 | 0.2598 | Yes | ||
| 24 | HUS1 | na | 1078 | 0.187 | 0.2666 | Yes | ||
| 25 | SSRP1 | na | 1081 | 0.186 | 0.2779 | Yes | ||
| 26 | DEK | na | 1170 | 0.178 | 0.2801 | Yes | ||
| 27 | RFC1 | na | 1211 | 0.175 | 0.2870 | Yes | ||
| 28 | TOP2A | na | 1242 | 0.173 | 0.2947 | Yes | ||
| 29 | RFC2 | na | 1304 | 0.168 | 0.2990 | Yes | ||
| 30 | CDC25B | na | 1335 | 0.166 | 0.3062 | Yes | ||
| 31 | DNMT1 | na | 1412 | 0.162 | 0.3086 | Yes | ||
| 32 | PLK4 | na | 1436 | 0.160 | 0.3162 | Yes | ||
| 33 | CBX5 | na | 1471 | 0.158 | 0.3226 | Yes | ||
| 34 | PRPS1 | na | 1559 | 0.151 | 0.3232 | Yes | ||
| 35 | PMS2 | na | 1560 | 0.151 | 0.3325 | Yes | ||
| 36 | ILF3 | na | 1589 | 0.149 | 0.3390 | Yes | ||
| 37 | TUBG1 | na | 1590 | 0.149 | 0.3482 | Yes | ||
| 38 | RAD51AP1 | na | 1770 | 0.137 | 0.3388 | Yes | ||
| 39 | CDKN3 | na | 1783 | 0.136 | 0.3460 | Yes | ||
| 40 | CKS2 | na | 1824 | 0.134 | 0.3503 | Yes | ||
| 41 | TFRC | na | 1964 | 0.122 | 0.3439 | Yes | ||
| 42 | IPO7 | na | 1968 | 0.122 | 0.3511 | Yes | ||
| 43 | BUB1B | na | 2033 | 0.118 | 0.3520 | Yes | ||
| 44 | STMN1 | na | 2080 | 0.115 | 0.3545 | Yes | ||
| 45 | XPO1 | na | 2111 | 0.113 | 0.3585 | Yes | ||
| 46 | MKI67 | na | 2116 | 0.113 | 0.3652 | Yes | ||
| 47 | SLBP | na | 2251 | 0.106 | 0.3583 | Yes | ||
| 48 | POLD3 | na | 2315 | 0.103 | 0.3584 | Yes | ||
| 49 | RPA3 | na | 2372 | 0.099 | 0.3589 | Yes | ||
| 50 | PLK1 | na | 2422 | 0.097 | 0.3600 | Yes | ||
| 51 | CDC20 | na | 2457 | 0.095 | 0.3625 | Yes | ||
| 52 | NASP | na | 2463 | 0.094 | 0.3678 | Yes | ||
| 53 | MCM2 | na | 2470 | 0.094 | 0.3730 | Yes | ||
| 54 | CHEK2 | na | 2657 | 0.084 | 0.3595 | No | ||
| 55 | CSE1L | na | 2753 | 0.079 | 0.3549 | No | ||
| 56 | PRDX4 | na | 2755 | 0.079 | 0.3597 | No | ||
| 57 | RNASEH2A | na | 2779 | 0.077 | 0.3621 | No | ||
| 58 | HMGA1 | na | 2834 | 0.074 | 0.3613 | No | ||
| 59 | DCK | na | 2916 | 0.070 | 0.3576 | No | ||
| 60 | NUP153 | na | 2925 | 0.070 | 0.3611 | No | ||
| 61 | CKS1B | na | 2980 | 0.067 | 0.3598 | No | ||
| 62 | MSH2 | na | 3024 | 0.065 | 0.3595 | No | ||
| 63 | EED | na | 3057 | 0.064 | 0.3603 | No | ||
| 64 | RRM2 | na | 3131 | 0.060 | 0.3566 | No | ||
| 65 | DUT | na | 3180 | 0.057 | 0.3554 | No | ||
| 66 | SNRPB | na | 3192 | 0.057 | 0.3578 | No | ||
| 67 | RBBP7 | na | 3247 | 0.054 | 0.3557 | No | ||
| 68 | PRKDC | na | 3259 | 0.054 | 0.3579 | No | ||
| 69 | CIT | na | 3264 | 0.053 | 0.3608 | No | ||
| 70 | TP53 | na | 3316 | 0.050 | 0.3588 | No | ||
| 71 | LMNB1 | na | 3448 | 0.043 | 0.3483 | No | ||
| 72 | PA2G4 | na | 3499 | 0.041 | 0.3458 | No | ||
| 73 | EIF2S1 | na | 3503 | 0.041 | 0.3481 | No | ||
| 74 | RAD21 | na | 3504 | 0.041 | 0.3506 | No | ||
| 75 | HMGB2 | na | 3543 | 0.039 | 0.3492 | No | ||
| 76 | TMPO | na | 3645 | 0.035 | 0.3412 | No | ||
| 77 | MYBL2 | na | 3662 | 0.034 | 0.3417 | No | ||
| 78 | EXOSC8 | na | 3679 | 0.033 | 0.3421 | No | ||
| 79 | AURKB | na | 3718 | 0.031 | 0.3403 | No | ||
| 80 | H2AFZ | na | 3740 | 0.030 | 0.3400 | No | ||
| 81 | BRCA1 | na | 3802 | 0.027 | 0.3356 | No | ||
| 82 | PSIP1 | na | 3815 | 0.027 | 0.3360 | No | ||
| 83 | BARD1 | na | 3908 | 0.022 | 0.3282 | No | ||
| 84 | AK2 | na | 3989 | 0.019 | 0.3213 | No | ||
| 85 | STAG1 | na | 4067 | 0.014 | 0.3145 | No | ||
| 86 | ANP32E | na | 4113 | 0.012 | 0.3107 | No | ||
| 87 | MTHFD2 | na | 4328 | 0.003 | 0.2894 | No | ||
| 88 | EZH2 | na | 4369 | 0.001 | 0.2855 | No | ||
| 89 | CENPE | na | 4385 | 0.000 | 0.2840 | No | ||
| 90 | SPAG5 | na | 4447 | -0.002 | 0.2780 | No | ||
| 91 | CTCF | na | 4578 | -0.009 | 0.2655 | No | ||
| 92 | NUP205 | na | 4580 | -0.009 | 0.2660 | No | ||
| 93 | KIF22 | na | 4624 | -0.010 | 0.2623 | No | ||
| 94 | TK1 | na | 4630 | -0.011 | 0.2625 | No | ||
| 95 | PNN | na | 4639 | -0.011 | 0.2624 | No | ||
| 96 | MYC | na | 4727 | -0.015 | 0.2546 | No | ||
| 97 | ASF1A | na | 4731 | -0.016 | 0.2553 | No | ||
| 98 | SYNCRIP | na | 4746 | -0.017 | 0.2549 | No | ||
| 99 | MCM3 | na | 4877 | -0.022 | 0.2432 | No | ||
| 100 | RAD50 | na | 5139 | -0.032 | 0.2190 | No | ||
| 101 | RFC3 | na | 5142 | -0.033 | 0.2208 | No | ||
| 102 | ING3 | na | 5173 | -0.034 | 0.2199 | No | ||
| 103 | MCM6 | na | 5244 | -0.037 | 0.2152 | No | ||
| 104 | MCM4 | na | 5291 | -0.039 | 0.2130 | No | ||
| 105 | CCNE1 | na | 5497 | -0.049 | 0.1955 | No | ||
| 106 | BRCA2 | na | 5649 | -0.055 | 0.1837 | No | ||
| 107 | MCM7 | na | 5667 | -0.056 | 0.1855 | No | ||
| 108 | POLD2 | na | 5992 | -0.069 | 0.1573 | No | ||
| 109 | LBR | na | 6052 | -0.072 | 0.1559 | No | ||
| 110 | CDC25A | na | 6059 | -0.073 | 0.1598 | No | ||
| 111 | MCM5 | na | 6121 | -0.075 | 0.1583 | No | ||
| 112 | UNG | na | 6171 | -0.077 | 0.1582 | No | ||
| 113 | RAN | na | 6254 | -0.082 | 0.1550 | No | ||
| 114 | HMGB3 | na | 6285 | -0.083 | 0.1571 | No | ||
| 115 | POLE | na | 6450 | -0.090 | 0.1463 | No | ||
| 116 | UBE2S | na | 6825 | -0.108 | 0.1154 | No | ||
| 117 | CDKN1B | na | 6829 | -0.108 | 0.1218 | No | ||
| 118 | KPNA2 | na | 6955 | -0.114 | 0.1163 | No | ||
| 119 | PCNA | na | 7009 | -0.117 | 0.1183 | No | ||
| 120 | ESPL1 | na | 7039 | -0.119 | 0.1227 | No | ||
| 121 | LIG1 | na | 7364 | -0.134 | 0.0985 | No | ||
| 122 | SHMT1 | na | 7413 | -0.136 | 0.1021 | No | ||
| 123 | POLD1 | na | 7941 | -0.165 | 0.0595 | No | ||
| 124 | MLH1 | na | 8188 | -0.180 | 0.0460 | No | ||
| 125 | NME1 | na | 8194 | -0.181 | 0.0566 | No | ||
| 126 | CDK4 | na | 8522 | -0.201 | 0.0363 | No | ||
| 127 | TIMELESS | na | 9360 | -0.265 | -0.0313 | No | ||
| 128 | NAP1L1 | na | 9367 | -0.267 | -0.0153 | No | ||
| 129 | RAD51C | na | 9369 | -0.267 | 0.0011 | No | ||
| 130 | PPM1D | na | 9735 | -0.319 | -0.0158 | No | ||
| 131 | CDKN1A | na | 10099 | -0.843 | 0.0000 | No |