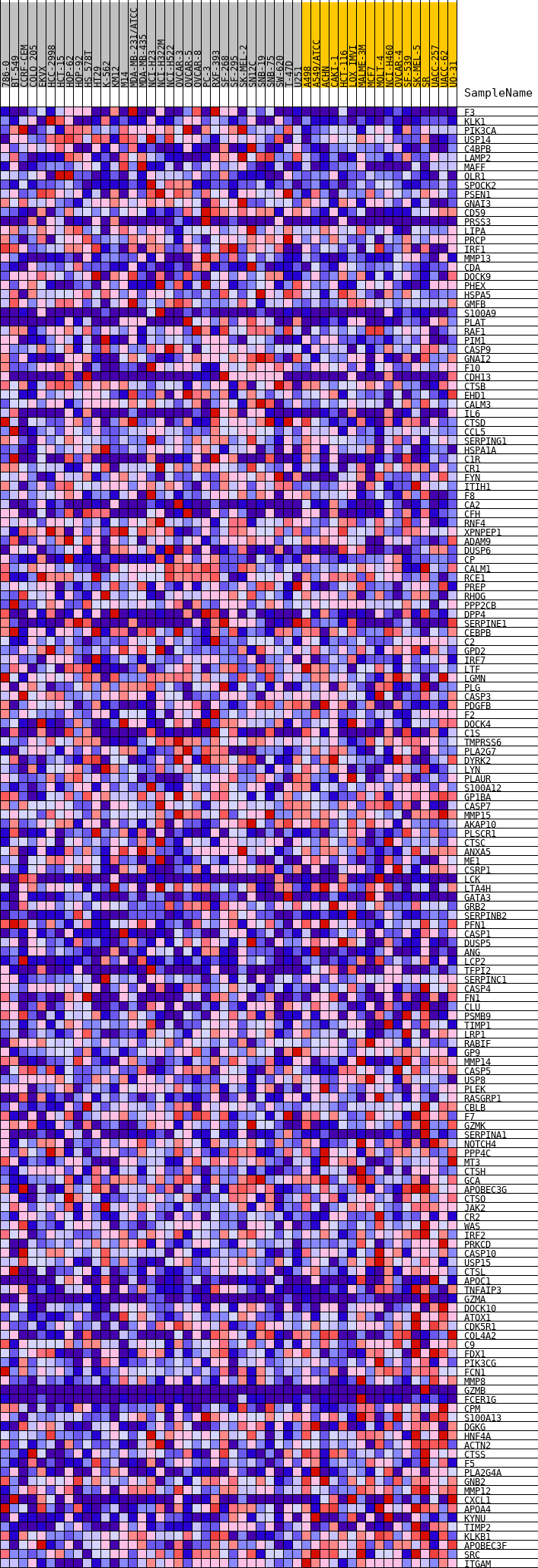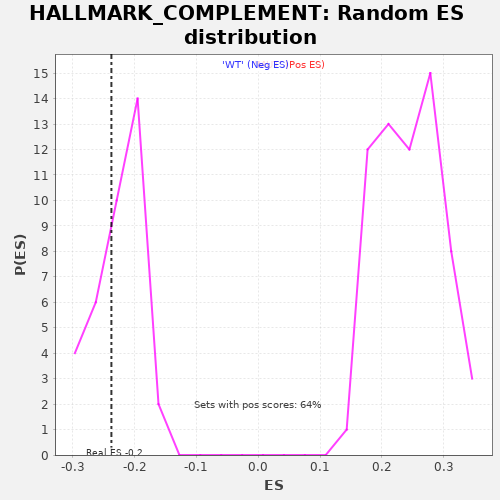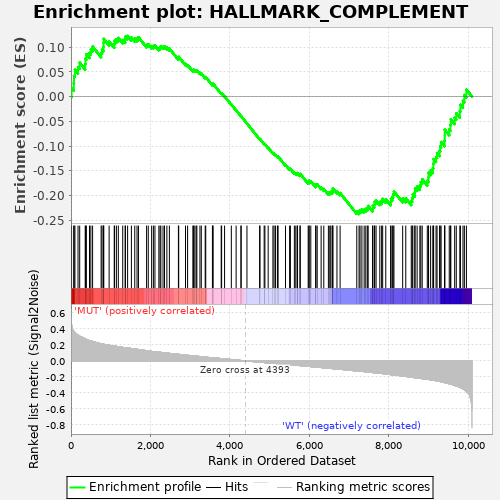
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_COMPLEMENT |
| Enrichment Score (ES) | -0.23740296 |
| Normalized Enrichment Score (NES) | -1.0569826 |
| Nominal p-value | 0.33333334 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.98 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | F3 | na | 14 | 0.454 | 0.0174 | No | ||
| 2 | KLK1 | na | 70 | 0.371 | 0.0272 | No | ||
| 3 | PIK3CA | na | 76 | 0.366 | 0.0419 | No | ||
| 4 | USP14 | na | 99 | 0.355 | 0.0544 | No | ||
| 5 | C4BPB | na | 177 | 0.322 | 0.0599 | No | ||
| 6 | LAMP2 | na | 219 | 0.308 | 0.0685 | No | ||
| 7 | MAFF | na | 358 | 0.274 | 0.0660 | No | ||
| 8 | OLR1 | na | 366 | 0.272 | 0.0766 | No | ||
| 9 | SPOCK2 | na | 387 | 0.268 | 0.0857 | No | ||
| 10 | PSEN1 | na | 466 | 0.254 | 0.0883 | No | ||
| 11 | GNAI3 | na | 500 | 0.249 | 0.0953 | No | ||
| 12 | CD59 | na | 544 | 0.243 | 0.1010 | No | ||
| 13 | PRSS3 | na | 759 | 0.214 | 0.0884 | No | ||
| 14 | LIPA | na | 782 | 0.211 | 0.0949 | No | ||
| 15 | PRCP | na | 817 | 0.208 | 0.1001 | No | ||
| 16 | IRF1 | na | 820 | 0.208 | 0.1085 | No | ||
| 17 | MMP13 | na | 825 | 0.207 | 0.1167 | No | ||
| 18 | CDA | na | 960 | 0.196 | 0.1113 | No | ||
| 19 | DOCK9 | na | 1089 | 0.185 | 0.1061 | No | ||
| 20 | PHEX | na | 1097 | 0.185 | 0.1131 | No | ||
| 21 | HSPA5 | na | 1143 | 0.181 | 0.1160 | No | ||
| 22 | GMFB | na | 1192 | 0.177 | 0.1185 | No | ||
| 23 | S100A9 | na | 1303 | 0.168 | 0.1144 | No | ||
| 24 | PLAT | na | 1364 | 0.165 | 0.1152 | No | ||
| 25 | RAF1 | na | 1371 | 0.164 | 0.1214 | No | ||
| 26 | PIM1 | na | 1426 | 0.161 | 0.1226 | No | ||
| 27 | CASP9 | na | 1521 | 0.154 | 0.1195 | No | ||
| 28 | GNAI2 | na | 1606 | 0.148 | 0.1172 | No | ||
| 29 | F10 | na | 1663 | 0.145 | 0.1175 | No | ||
| 30 | CDH13 | na | 1699 | 0.142 | 0.1199 | No | ||
| 31 | CTSB | na | 1904 | 0.128 | 0.1046 | No | ||
| 32 | EHD1 | na | 1946 | 0.124 | 0.1056 | No | ||
| 33 | CALM3 | na | 2031 | 0.118 | 0.1020 | No | ||
| 34 | IL6 | na | 2075 | 0.115 | 0.1025 | No | ||
| 35 | CTSD | na | 2114 | 0.113 | 0.1034 | No | ||
| 36 | CCL5 | na | 2213 | 0.109 | 0.0980 | No | ||
| 37 | SERPING1 | na | 2243 | 0.107 | 0.0995 | No | ||
| 38 | HSPA1A | na | 2263 | 0.106 | 0.1020 | No | ||
| 39 | C1R | na | 2318 | 0.103 | 0.1008 | No | ||
| 40 | CR1 | na | 2355 | 0.100 | 0.1013 | No | ||
| 41 | FYN | na | 2415 | 0.097 | 0.0994 | No | ||
| 42 | ITIH1 | na | 2478 | 0.093 | 0.0971 | No | ||
| 43 | F8 | na | 2705 | 0.081 | 0.0777 | No | ||
| 44 | CA2 | na | 2713 | 0.081 | 0.0803 | No | ||
| 45 | CFH | na | 2884 | 0.072 | 0.0662 | No | ||
| 46 | RNF4 | na | 2937 | 0.069 | 0.0638 | No | ||
| 47 | XPNPEP1 | na | 3070 | 0.063 | 0.0532 | No | ||
| 48 | ADAM9 | na | 3084 | 0.062 | 0.0544 | No | ||
| 49 | DUSP6 | na | 3118 | 0.060 | 0.0536 | No | ||
| 50 | CP | na | 3152 | 0.059 | 0.0527 | No | ||
| 51 | CALM1 | na | 3170 | 0.058 | 0.0534 | No | ||
| 52 | RCE1 | na | 3248 | 0.054 | 0.0479 | No | ||
| 53 | PREP | na | 3282 | 0.052 | 0.0467 | No | ||
| 54 | RHOG | na | 3380 | 0.047 | 0.0389 | No | ||
| 55 | PPP2CB | na | 3394 | 0.046 | 0.0395 | No | ||
| 56 | DPP4 | na | 3562 | 0.038 | 0.0243 | No | ||
| 57 | SERPINE1 | na | 3565 | 0.038 | 0.0257 | No | ||
| 58 | CEBPB | na | 3580 | 0.037 | 0.0258 | No | ||
| 59 | C2 | na | 3783 | 0.028 | 0.0067 | No | ||
| 60 | GPD2 | na | 3794 | 0.028 | 0.0068 | No | ||
| 61 | IRF7 | na | 3868 | 0.024 | 0.0005 | No | ||
| 62 | LTF | na | 4038 | 0.016 | -0.0159 | No | ||
| 63 | LGMN | na | 4159 | 0.010 | -0.0275 | No | ||
| 64 | PLG | na | 4280 | 0.005 | -0.0394 | No | ||
| 65 | CASP3 | na | 4286 | 0.004 | -0.0397 | No | ||
| 66 | PDGFB | na | 4432 | -0.002 | -0.0542 | No | ||
| 67 | F2 | na | 4751 | -0.017 | -0.0855 | No | ||
| 68 | DOCK4 | na | 4760 | -0.017 | -0.0856 | No | ||
| 69 | C1S | na | 4865 | -0.021 | -0.0952 | No | ||
| 70 | TMPRSS6 | na | 4884 | -0.022 | -0.0961 | No | ||
| 71 | PLA2G7 | na | 4966 | -0.025 | -0.1032 | No | ||
| 72 | DYRK2 | na | 5086 | -0.030 | -0.1140 | No | ||
| 73 | LYN | na | 5115 | -0.031 | -0.1155 | No | ||
| 74 | PLAUR | na | 5152 | -0.033 | -0.1178 | No | ||
| 75 | S100A12 | na | 5200 | -0.035 | -0.1210 | No | ||
| 76 | GP1BA | na | 5216 | -0.036 | -0.1210 | No | ||
| 77 | CASP7 | na | 5403 | -0.044 | -0.1379 | No | ||
| 78 | MMP15 | na | 5506 | -0.049 | -0.1461 | No | ||
| 79 | AKAP10 | na | 5526 | -0.050 | -0.1460 | No | ||
| 80 | PLSCR1 | na | 5626 | -0.054 | -0.1537 | No | ||
| 81 | CTSC | na | 5662 | -0.056 | -0.1549 | No | ||
| 82 | ANXA5 | na | 5697 | -0.057 | -0.1560 | No | ||
| 83 | ME1 | na | 5706 | -0.057 | -0.1544 | No | ||
| 84 | CSRP1 | na | 5767 | -0.059 | -0.1580 | No | ||
| 85 | LCK | na | 5774 | -0.060 | -0.1561 | No | ||
| 86 | LTA4H | na | 5974 | -0.069 | -0.1733 | No | ||
| 87 | GATA3 | na | 5996 | -0.070 | -0.1725 | No | ||
| 88 | GRB2 | na | 5997 | -0.070 | -0.1696 | No | ||
| 89 | SERPINB2 | na | 6040 | -0.072 | -0.1709 | No | ||
| 90 | PFN1 | na | 6162 | -0.077 | -0.1799 | No | ||
| 91 | CASP1 | na | 6169 | -0.077 | -0.1773 | No | ||
| 92 | DUSP5 | na | 6201 | -0.079 | -0.1771 | No | ||
| 93 | ANG | na | 6300 | -0.084 | -0.1835 | No | ||
| 94 | LCP2 | na | 6367 | -0.086 | -0.1866 | No | ||
| 95 | TFPI2 | na | 6483 | -0.092 | -0.1944 | No | ||
| 96 | SERPINC1 | na | 6518 | -0.093 | -0.1939 | No | ||
| 97 | CASP4 | na | 6543 | -0.095 | -0.1924 | No | ||
| 98 | FN1 | na | 6579 | -0.097 | -0.1919 | No | ||
| 99 | CLU | na | 6591 | -0.097 | -0.1890 | No | ||
| 100 | PSMB9 | na | 6596 | -0.098 | -0.1853 | No | ||
| 101 | TIMP1 | na | 6699 | -0.102 | -0.1914 | No | ||
| 102 | LRP1 | na | 6778 | -0.106 | -0.1948 | No | ||
| 103 | RABIF | na | 7198 | -0.125 | -0.2318 | No | ||
| 104 | GP9 | na | 7255 | -0.128 | -0.2321 | Yes | ||
| 105 | MMP14 | na | 7280 | -0.130 | -0.2291 | Yes | ||
| 106 | CASP5 | na | 7327 | -0.132 | -0.2283 | Yes | ||
| 107 | USP8 | na | 7384 | -0.135 | -0.2284 | Yes | ||
| 108 | PLEK | na | 7425 | -0.137 | -0.2267 | Yes | ||
| 109 | RASGRP1 | na | 7464 | -0.139 | -0.2248 | Yes | ||
| 110 | CBLB | na | 7485 | -0.140 | -0.2210 | Yes | ||
| 111 | F7 | na | 7595 | -0.147 | -0.2259 | Yes | ||
| 112 | GZMK | na | 7603 | -0.147 | -0.2205 | Yes | ||
| 113 | SERPINA1 | na | 7641 | -0.148 | -0.2181 | Yes | ||
| 114 | NOTCH4 | na | 7649 | -0.149 | -0.2127 | Yes | ||
| 115 | PPP4C | na | 7684 | -0.151 | -0.2098 | Yes | ||
| 116 | MT3 | na | 7778 | -0.156 | -0.2127 | Yes | ||
| 117 | CTSH | na | 7824 | -0.158 | -0.2107 | Yes | ||
| 118 | GCA | na | 7850 | -0.159 | -0.2066 | Yes | ||
| 119 | APOBEC3G | na | 7928 | -0.164 | -0.2076 | Yes | ||
| 120 | CTSO | na | 8050 | -0.172 | -0.2126 | Yes | ||
| 121 | JAK2 | na | 8060 | -0.173 | -0.2064 | Yes | ||
| 122 | CR2 | na | 8094 | -0.174 | -0.2025 | Yes | ||
| 123 | WAS | na | 8114 | -0.176 | -0.1971 | Yes | ||
| 124 | IRF2 | na | 8134 | -0.177 | -0.1917 | Yes | ||
| 125 | PRKCD | na | 8354 | -0.190 | -0.2058 | Yes | ||
| 126 | CASP10 | na | 8432 | -0.195 | -0.2055 | Yes | ||
| 127 | USP15 | na | 8570 | -0.205 | -0.2108 | Yes | ||
| 128 | CTSL | na | 8594 | -0.206 | -0.2046 | Yes | ||
| 129 | APOC1 | na | 8610 | -0.207 | -0.1975 | Yes | ||
| 130 | TNFAIP3 | na | 8660 | -0.211 | -0.1937 | Yes | ||
| 131 | GZMA | na | 8665 | -0.211 | -0.1854 | Yes | ||
| 132 | DOCK10 | na | 8718 | -0.215 | -0.1817 | Yes | ||
| 133 | ATOX1 | na | 8787 | -0.219 | -0.1795 | Yes | ||
| 134 | CDK5R1 | na | 8811 | -0.221 | -0.1726 | Yes | ||
| 135 | COL4A2 | na | 8846 | -0.223 | -0.1668 | Yes | ||
| 136 | C9 | na | 8974 | -0.232 | -0.1700 | Yes | ||
| 137 | FDX1 | na | 9003 | -0.233 | -0.1632 | Yes | ||
| 138 | PIK3CG | na | 9006 | -0.233 | -0.1537 | Yes | ||
| 139 | FCN1 | na | 9058 | -0.237 | -0.1490 | Yes | ||
| 140 | MMP8 | na | 9115 | -0.243 | -0.1446 | Yes | ||
| 141 | GZMB | na | 9126 | -0.243 | -0.1356 | Yes | ||
| 142 | FCER1G | na | 9134 | -0.243 | -0.1262 | Yes | ||
| 143 | CPM | na | 9195 | -0.249 | -0.1219 | Yes | ||
| 144 | S100A13 | na | 9220 | -0.251 | -0.1140 | Yes | ||
| 145 | DGKG | na | 9283 | -0.257 | -0.1096 | Yes | ||
| 146 | HNF4A | na | 9299 | -0.259 | -0.1003 | Yes | ||
| 147 | ACTN2 | na | 9322 | -0.262 | -0.0917 | Yes | ||
| 148 | CTSS | na | 9408 | -0.272 | -0.0890 | Yes | ||
| 149 | F5 | na | 9413 | -0.273 | -0.0781 | Yes | ||
| 150 | PLA2G4A | na | 9414 | -0.273 | -0.0668 | Yes | ||
| 151 | GNB2 | na | 9527 | -0.287 | -0.0662 | Yes | ||
| 152 | MMP12 | na | 9557 | -0.291 | -0.0571 | Yes | ||
| 153 | CXCL1 | na | 9567 | -0.292 | -0.0459 | Yes | ||
| 154 | APOA4 | na | 9664 | -0.309 | -0.0428 | Yes | ||
| 155 | KYNU | na | 9704 | -0.315 | -0.0337 | Yes | ||
| 156 | TIMP2 | na | 9794 | -0.330 | -0.0290 | Yes | ||
| 157 | KLKB1 | na | 9809 | -0.334 | -0.0165 | Yes | ||
| 158 | APOBEC3F | na | 9873 | -0.349 | -0.0084 | Yes | ||
| 159 | SRC | na | 9908 | -0.362 | 0.0031 | Yes | ||
| 160 | ITGAM | na | 9959 | -0.386 | 0.0141 | Yes |