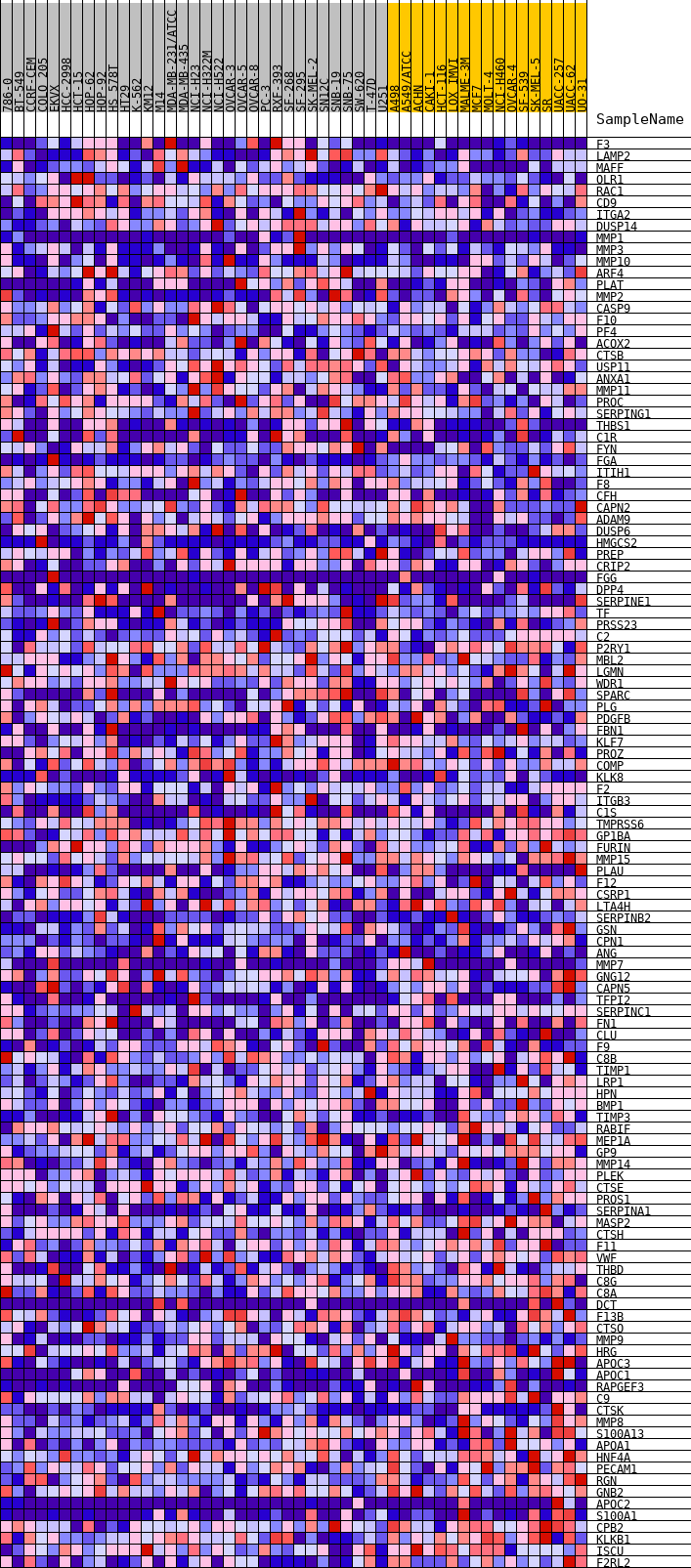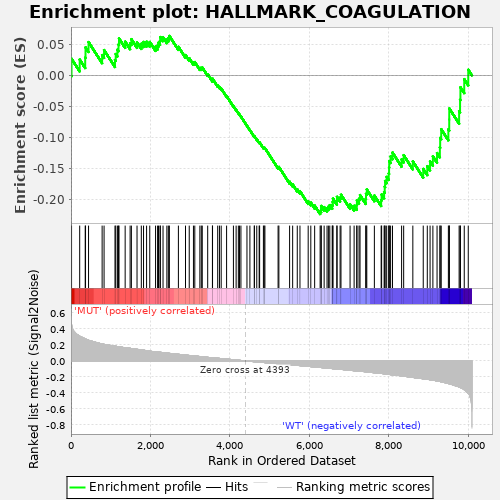
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_COAGULATION |
| Enrichment Score (ES) | -0.22384477 |
| Normalized Enrichment Score (NES) | -0.83412325 |
| Nominal p-value | 0.675 |
| FDR q-value | 0.9656248 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | F3 | na | 14 | 0.454 | 0.0270 | No | ||
| 2 | LAMP2 | na | 219 | 0.308 | 0.0258 | No | ||
| 3 | MAFF | na | 358 | 0.274 | 0.0291 | No | ||
| 4 | OLR1 | na | 366 | 0.272 | 0.0454 | No | ||
| 5 | RAC1 | na | 442 | 0.258 | 0.0540 | No | ||
| 6 | CD9 | na | 784 | 0.211 | 0.0330 | No | ||
| 7 | ITGA2 | na | 833 | 0.206 | 0.0411 | No | ||
| 8 | DUSP14 | na | 1108 | 0.184 | 0.0252 | No | ||
| 9 | MMP1 | na | 1127 | 0.183 | 0.0348 | No | ||
| 10 | MMP3 | na | 1169 | 0.178 | 0.0418 | No | ||
| 11 | MMP10 | na | 1197 | 0.176 | 0.0501 | No | ||
| 12 | ARF4 | na | 1209 | 0.175 | 0.0600 | No | ||
| 13 | PLAT | na | 1364 | 0.165 | 0.0548 | No | ||
| 14 | MMP2 | na | 1489 | 0.156 | 0.0522 | No | ||
| 15 | CASP9 | na | 1521 | 0.154 | 0.0587 | No | ||
| 16 | F10 | na | 1663 | 0.145 | 0.0536 | No | ||
| 17 | PF4 | na | 1771 | 0.137 | 0.0514 | No | ||
| 18 | ACOX2 | na | 1827 | 0.133 | 0.0542 | No | ||
| 19 | CTSB | na | 1904 | 0.128 | 0.0546 | No | ||
| 20 | USP11 | na | 1984 | 0.121 | 0.0542 | No | ||
| 21 | ANXA1 | na | 2130 | 0.112 | 0.0467 | No | ||
| 22 | MMP11 | na | 2181 | 0.110 | 0.0486 | No | ||
| 23 | PROC | na | 2204 | 0.109 | 0.0532 | No | ||
| 24 | SERPING1 | na | 2243 | 0.107 | 0.0561 | No | ||
| 25 | THBS1 | na | 2250 | 0.106 | 0.0621 | No | ||
| 26 | C1R | na | 2318 | 0.103 | 0.0618 | No | ||
| 27 | FYN | na | 2415 | 0.097 | 0.0583 | No | ||
| 28 | FGA | na | 2452 | 0.095 | 0.0606 | No | ||
| 29 | ITIH1 | na | 2478 | 0.093 | 0.0640 | No | ||
| 30 | F8 | na | 2705 | 0.081 | 0.0464 | No | ||
| 31 | CFH | na | 2884 | 0.072 | 0.0330 | No | ||
| 32 | CAPN2 | na | 2978 | 0.067 | 0.0279 | No | ||
| 33 | ADAM9 | na | 3084 | 0.062 | 0.0213 | No | ||
| 34 | DUSP6 | na | 3118 | 0.060 | 0.0217 | No | ||
| 35 | HMGCS2 | na | 3246 | 0.054 | 0.0124 | No | ||
| 36 | PREP | na | 3282 | 0.052 | 0.0122 | No | ||
| 37 | CRIP2 | na | 3306 | 0.051 | 0.0130 | No | ||
| 38 | FGG | na | 3443 | 0.043 | 0.0021 | No | ||
| 39 | DPP4 | na | 3562 | 0.038 | -0.0073 | No | ||
| 40 | SERPINE1 | na | 3565 | 0.038 | -0.0051 | No | ||
| 41 | TF | na | 3695 | 0.032 | -0.0161 | No | ||
| 42 | PRSS23 | na | 3739 | 0.030 | -0.0185 | No | ||
| 43 | C2 | na | 3783 | 0.028 | -0.0210 | No | ||
| 44 | P2RY1 | na | 3919 | 0.022 | -0.0332 | No | ||
| 45 | MBL2 | na | 4091 | 0.013 | -0.0495 | No | ||
| 46 | LGMN | na | 4159 | 0.010 | -0.0556 | No | ||
| 47 | WDR1 | na | 4215 | 0.008 | -0.0606 | No | ||
| 48 | SPARC | na | 4245 | 0.006 | -0.0631 | No | ||
| 49 | PLG | na | 4280 | 0.005 | -0.0662 | No | ||
| 50 | PDGFB | na | 4432 | -0.002 | -0.0812 | No | ||
| 51 | FBN1 | na | 4506 | -0.005 | -0.0882 | No | ||
| 52 | KLF7 | na | 4617 | -0.010 | -0.0986 | No | ||
| 53 | PROZ | na | 4619 | -0.010 | -0.0981 | No | ||
| 54 | COMP | na | 4679 | -0.013 | -0.1031 | No | ||
| 55 | KLK8 | na | 4739 | -0.016 | -0.1080 | No | ||
| 56 | F2 | na | 4751 | -0.017 | -0.1081 | No | ||
| 57 | ITGB3 | na | 4846 | -0.021 | -0.1162 | No | ||
| 58 | C1S | na | 4865 | -0.021 | -0.1167 | No | ||
| 59 | TMPRSS6 | na | 4884 | -0.022 | -0.1172 | No | ||
| 60 | GP1BA | na | 5216 | -0.036 | -0.1481 | No | ||
| 61 | FURIN | na | 5236 | -0.037 | -0.1477 | No | ||
| 62 | MMP15 | na | 5506 | -0.049 | -0.1716 | No | ||
| 63 | PLAU | na | 5583 | -0.052 | -0.1759 | No | ||
| 64 | F12 | na | 5699 | -0.057 | -0.1838 | No | ||
| 65 | CSRP1 | na | 5767 | -0.059 | -0.1868 | No | ||
| 66 | LTA4H | na | 5974 | -0.069 | -0.2032 | No | ||
| 67 | SERPINB2 | na | 6040 | -0.072 | -0.2052 | No | ||
| 68 | GSN | na | 6137 | -0.076 | -0.2101 | No | ||
| 69 | CPN1 | na | 6275 | -0.083 | -0.2187 | Yes | ||
| 70 | ANG | na | 6300 | -0.084 | -0.2159 | Yes | ||
| 71 | MMP7 | na | 6301 | -0.084 | -0.2106 | Yes | ||
| 72 | GNG12 | na | 6376 | -0.087 | -0.2126 | Yes | ||
| 73 | CAPN5 | na | 6449 | -0.090 | -0.2142 | Yes | ||
| 74 | TFPI2 | na | 6483 | -0.092 | -0.2117 | Yes | ||
| 75 | SERPINC1 | na | 6518 | -0.093 | -0.2093 | Yes | ||
| 76 | FN1 | na | 6579 | -0.097 | -0.2093 | Yes | ||
| 77 | CLU | na | 6591 | -0.097 | -0.2043 | Yes | ||
| 78 | F9 | na | 6598 | -0.098 | -0.1988 | Yes | ||
| 79 | C8B | na | 6698 | -0.102 | -0.2023 | Yes | ||
| 80 | TIMP1 | na | 6699 | -0.102 | -0.1959 | Yes | ||
| 81 | LRP1 | na | 6778 | -0.106 | -0.1970 | Yes | ||
| 82 | HPN | na | 6800 | -0.107 | -0.1925 | Yes | ||
| 83 | BMP1 | na | 7029 | -0.118 | -0.2080 | Yes | ||
| 84 | TIMP3 | na | 7131 | -0.123 | -0.2104 | Yes | ||
| 85 | RABIF | na | 7198 | -0.125 | -0.2092 | Yes | ||
| 86 | MEP1A | na | 7202 | -0.126 | -0.2016 | Yes | ||
| 87 | GP9 | na | 7255 | -0.128 | -0.1988 | Yes | ||
| 88 | MMP14 | na | 7280 | -0.130 | -0.1931 | Yes | ||
| 89 | PLEK | na | 7425 | -0.137 | -0.1990 | Yes | ||
| 90 | CTSE | na | 7431 | -0.137 | -0.1909 | Yes | ||
| 91 | PROS1 | na | 7450 | -0.138 | -0.1841 | Yes | ||
| 92 | SERPINA1 | na | 7641 | -0.148 | -0.1938 | Yes | ||
| 93 | MASP2 | na | 7812 | -0.158 | -0.2010 | Yes | ||
| 94 | CTSH | na | 7824 | -0.158 | -0.1922 | Yes | ||
| 95 | F11 | na | 7887 | -0.162 | -0.1883 | Yes | ||
| 96 | VWF | na | 7902 | -0.163 | -0.1796 | Yes | ||
| 97 | THBD | na | 7912 | -0.163 | -0.1702 | Yes | ||
| 98 | C8G | na | 7951 | -0.166 | -0.1637 | Yes | ||
| 99 | C8A | na | 8003 | -0.169 | -0.1582 | Yes | ||
| 100 | DCT | na | 8012 | -0.170 | -0.1484 | Yes | ||
| 101 | F13B | na | 8016 | -0.170 | -0.1381 | Yes | ||
| 102 | CTSO | na | 8050 | -0.172 | -0.1306 | Yes | ||
| 103 | MMP9 | na | 8098 | -0.175 | -0.1244 | Yes | ||
| 104 | HRG | na | 8327 | -0.188 | -0.1355 | Yes | ||
| 105 | APOC3 | na | 8379 | -0.192 | -0.1286 | Yes | ||
| 106 | APOC1 | na | 8610 | -0.207 | -0.1387 | Yes | ||
| 107 | RAPGEF3 | na | 8872 | -0.225 | -0.1508 | Yes | ||
| 108 | C9 | na | 8974 | -0.232 | -0.1465 | Yes | ||
| 109 | CTSK | na | 9044 | -0.236 | -0.1386 | Yes | ||
| 110 | MMP8 | na | 9115 | -0.243 | -0.1305 | Yes | ||
| 111 | S100A13 | na | 9220 | -0.251 | -0.1252 | Yes | ||
| 112 | APOA1 | na | 9291 | -0.258 | -0.1161 | Yes | ||
| 113 | HNF4A | na | 9299 | -0.259 | -0.1006 | Yes | ||
| 114 | PECAM1 | na | 9325 | -0.262 | -0.0867 | Yes | ||
| 115 | RGN | na | 9505 | -0.284 | -0.0869 | Yes | ||
| 116 | GNB2 | na | 9527 | -0.287 | -0.0711 | Yes | ||
| 117 | APOC2 | na | 9528 | -0.287 | -0.0531 | Yes | ||
| 118 | S100A1 | na | 9776 | -0.326 | -0.0575 | Yes | ||
| 119 | CPB2 | na | 9800 | -0.332 | -0.0390 | Yes | ||
| 120 | KLKB1 | na | 9809 | -0.334 | -0.0190 | Yes | ||
| 121 | ISCU | na | 9905 | -0.361 | -0.0059 | Yes | ||
| 122 | F2RL2 | na | 10006 | -0.404 | 0.0093 | Yes |