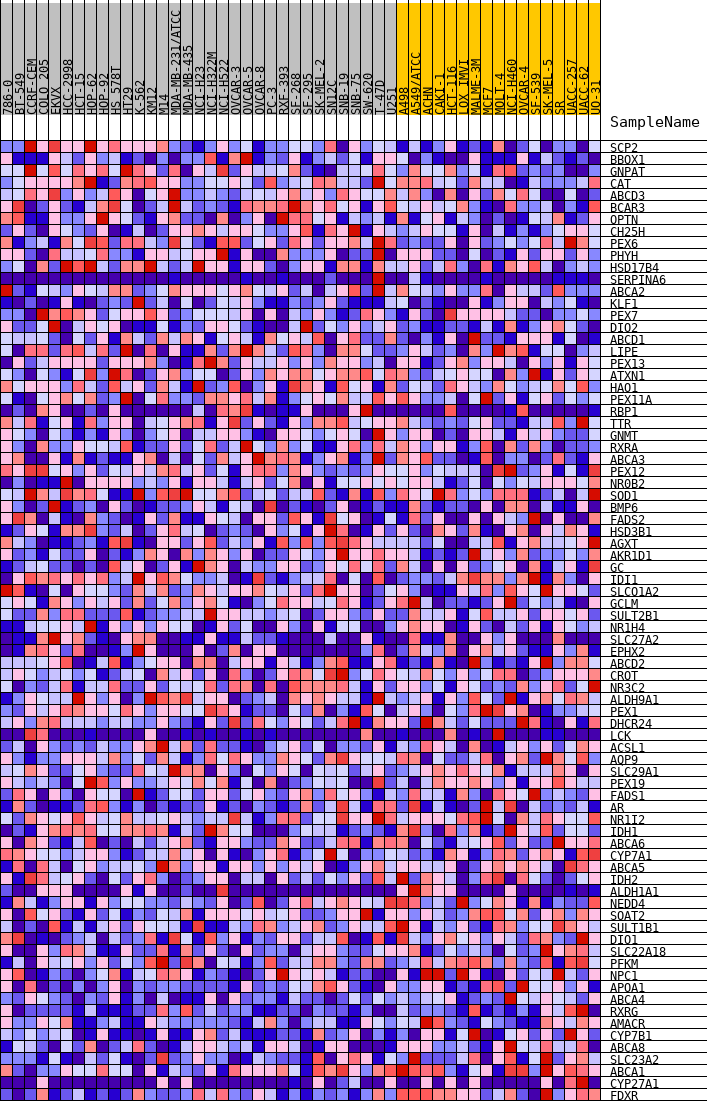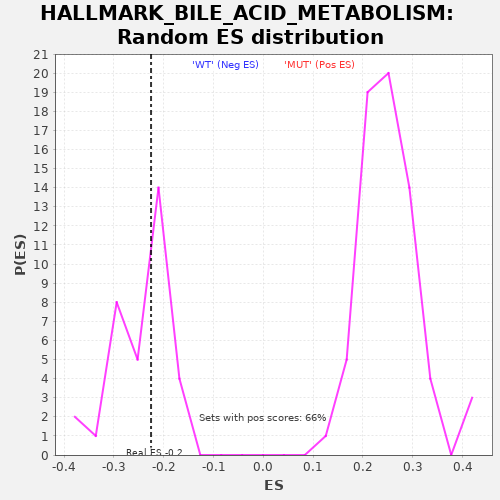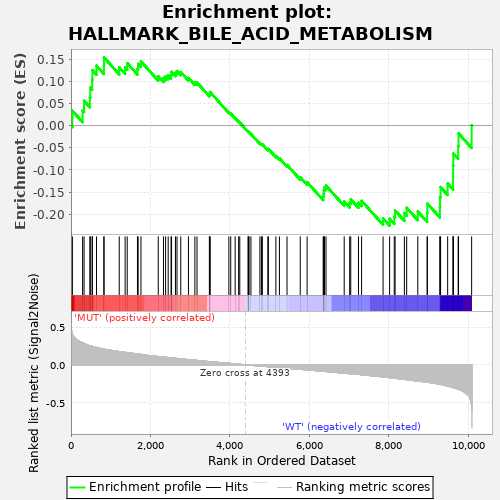
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_BILE_ACID_METABOLISM |
| Enrichment Score (ES) | -0.22543517 |
| Normalized Enrichment Score (NES) | -0.9233428 |
| Nominal p-value | 0.5588235 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SCP2 | na | 35 | 0.408 | 0.0333 | No | ||
| 2 | BBOX1 | na | 292 | 0.291 | 0.0340 | No | ||
| 3 | GNPAT | na | 329 | 0.282 | 0.0558 | No | ||
| 4 | CAT | na | 476 | 0.252 | 0.0640 | No | ||
| 5 | ABCD3 | na | 486 | 0.251 | 0.0857 | No | ||
| 6 | BCAR3 | na | 532 | 0.245 | 0.1033 | No | ||
| 7 | OPTN | na | 539 | 0.243 | 0.1246 | No | ||
| 8 | CH25H | na | 643 | 0.228 | 0.1349 | No | ||
| 9 | PEX6 | na | 829 | 0.207 | 0.1351 | No | ||
| 10 | PHYH | na | 831 | 0.207 | 0.1536 | No | ||
| 11 | HSD17B4 | na | 1215 | 0.175 | 0.1311 | No | ||
| 12 | SERPINA6 | na | 1363 | 0.165 | 0.1313 | No | ||
| 13 | ABCA2 | na | 1416 | 0.161 | 0.1407 | No | ||
| 14 | KLF1 | na | 1674 | 0.143 | 0.1279 | No | ||
| 15 | PEX7 | na | 1691 | 0.142 | 0.1392 | No | ||
| 16 | DIO2 | na | 1762 | 0.137 | 0.1446 | No | ||
| 17 | ABCD1 | na | 2198 | 0.109 | 0.1110 | No | ||
| 18 | LIPE | na | 2329 | 0.102 | 0.1072 | No | ||
| 19 | PEX13 | na | 2383 | 0.099 | 0.1108 | No | ||
| 20 | ATXN1 | na | 2448 | 0.095 | 0.1131 | No | ||
| 21 | HAO1 | na | 2520 | 0.091 | 0.1142 | No | ||
| 22 | PEX11A | na | 2533 | 0.091 | 0.1211 | No | ||
| 23 | RBP1 | na | 2631 | 0.086 | 0.1192 | No | ||
| 24 | TTR | na | 2670 | 0.084 | 0.1230 | No | ||
| 25 | GNMT | na | 2765 | 0.078 | 0.1206 | No | ||
| 26 | RXRA | na | 2958 | 0.068 | 0.1076 | No | ||
| 27 | ABCA3 | na | 3120 | 0.060 | 0.0970 | No | ||
| 28 | PEX12 | na | 3174 | 0.058 | 0.0969 | No | ||
| 29 | NR0B2 | na | 3485 | 0.042 | 0.0697 | No | ||
| 30 | SOD1 | na | 3500 | 0.041 | 0.0720 | No | ||
| 31 | BMP6 | na | 3501 | 0.041 | 0.0757 | No | ||
| 32 | FADS2 | na | 3976 | 0.019 | 0.0301 | No | ||
| 33 | HSD3B1 | na | 4023 | 0.017 | 0.0271 | No | ||
| 34 | AGXT | na | 4135 | 0.011 | 0.0170 | No | ||
| 35 | AKR1D1 | na | 4222 | 0.008 | 0.0091 | No | ||
| 36 | GC | na | 4251 | 0.006 | 0.0068 | No | ||
| 37 | IDI1 | na | 4458 | -0.003 | -0.0135 | No | ||
| 38 | SLCO1A2 | na | 4471 | -0.003 | -0.0143 | No | ||
| 39 | GCLM | na | 4477 | -0.004 | -0.0145 | No | ||
| 40 | SULT2B1 | na | 4482 | -0.004 | -0.0146 | No | ||
| 41 | NR1H4 | na | 4532 | -0.006 | -0.0189 | No | ||
| 42 | SLC27A2 | na | 4758 | -0.017 | -0.0398 | No | ||
| 43 | EPHX2 | na | 4796 | -0.019 | -0.0418 | No | ||
| 44 | ABCD2 | na | 4820 | -0.020 | -0.0423 | No | ||
| 45 | CROT | na | 4958 | -0.025 | -0.0538 | No | ||
| 46 | NR3C2 | na | 4971 | -0.025 | -0.0527 | No | ||
| 47 | ALDH9A1 | na | 5160 | -0.033 | -0.0684 | No | ||
| 48 | PEX1 | na | 5250 | -0.037 | -0.0739 | No | ||
| 49 | DHCR24 | na | 5441 | -0.046 | -0.0888 | No | ||
| 50 | LCK | na | 5774 | -0.060 | -0.1165 | No | ||
| 51 | ACSL1 | na | 5947 | -0.067 | -0.1276 | No | ||
| 52 | AQP9 | na | 6352 | -0.086 | -0.1602 | No | ||
| 53 | SLC29A1 | na | 6362 | -0.086 | -0.1533 | No | ||
| 54 | PEX19 | na | 6369 | -0.086 | -0.1461 | No | ||
| 55 | FADS1 | na | 6385 | -0.088 | -0.1397 | No | ||
| 56 | AR | na | 6423 | -0.089 | -0.1354 | No | ||
| 57 | NR1I2 | na | 6881 | -0.111 | -0.1710 | No | ||
| 58 | IDH1 | na | 7019 | -0.117 | -0.1741 | No | ||
| 59 | ABCA6 | na | 7046 | -0.119 | -0.1659 | No | ||
| 60 | CYP7A1 | na | 7241 | -0.128 | -0.1738 | No | ||
| 61 | ABCA5 | na | 7319 | -0.132 | -0.1696 | No | ||
| 62 | IDH2 | na | 7860 | -0.160 | -0.2091 | No | ||
| 63 | ALDH1A1 | na | 8025 | -0.171 | -0.2100 | Yes | ||
| 64 | NEDD4 | na | 8142 | -0.177 | -0.2056 | Yes | ||
| 65 | SOAT2 | na | 8163 | -0.179 | -0.1915 | Yes | ||
| 66 | SULT1B1 | na | 8397 | -0.193 | -0.1974 | Yes | ||
| 67 | DIO1 | na | 8455 | -0.197 | -0.1853 | Yes | ||
| 68 | SLC22A18 | na | 8735 | -0.216 | -0.1937 | Yes | ||
| 69 | PFKM | na | 8970 | -0.231 | -0.1962 | Yes | ||
| 70 | NPC1 | na | 8976 | -0.232 | -0.1758 | Yes | ||
| 71 | APOA1 | na | 9291 | -0.258 | -0.1839 | Yes | ||
| 72 | ABCA4 | na | 9296 | -0.259 | -0.1609 | Yes | ||
| 73 | RXRG | na | 9309 | -0.261 | -0.1386 | Yes | ||
| 74 | AMACR | na | 9487 | -0.283 | -0.1308 | Yes | ||
| 75 | CYP7B1 | na | 9624 | -0.302 | -0.1172 | Yes | ||
| 76 | ABCA8 | na | 9625 | -0.302 | -0.0899 | Yes | ||
| 77 | SLC23A2 | na | 9630 | -0.303 | -0.0630 | Yes | ||
| 78 | ABCA1 | na | 9752 | -0.322 | -0.0460 | Yes | ||
| 79 | CYP27A1 | na | 9761 | -0.324 | -0.0176 | Yes | ||
| 80 | FDXR | na | 10091 | -0.568 | 0.0008 | Yes |