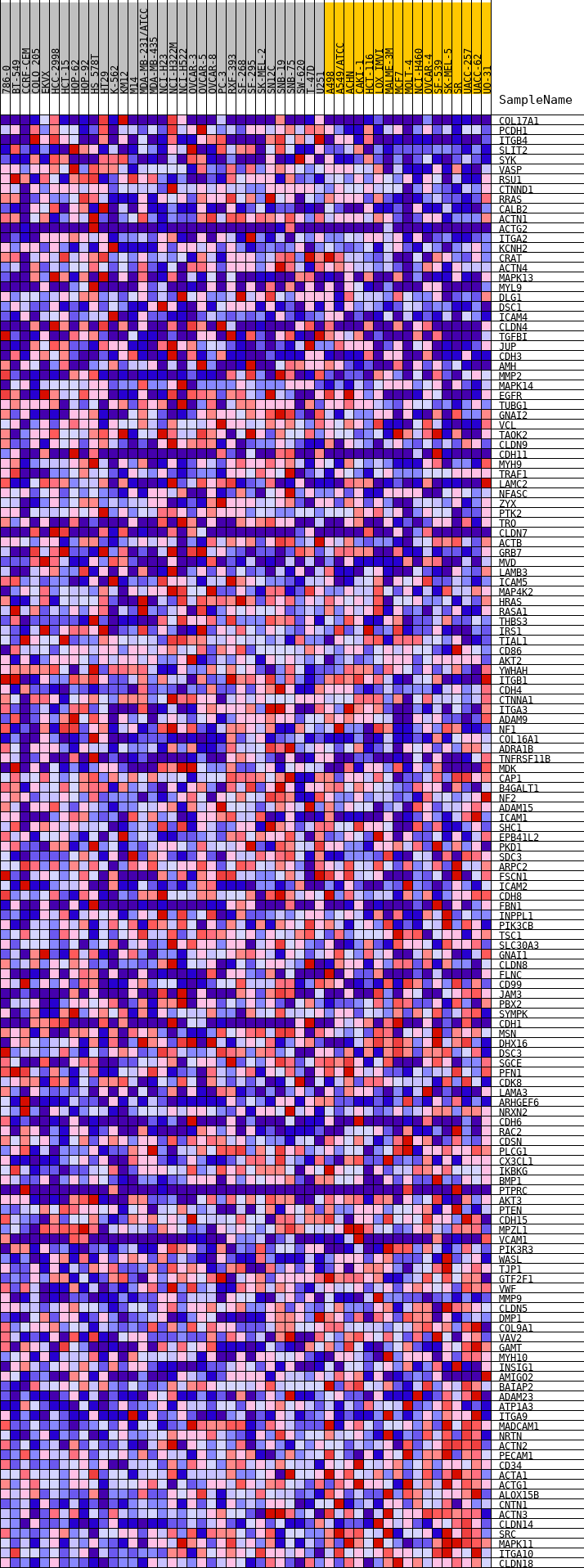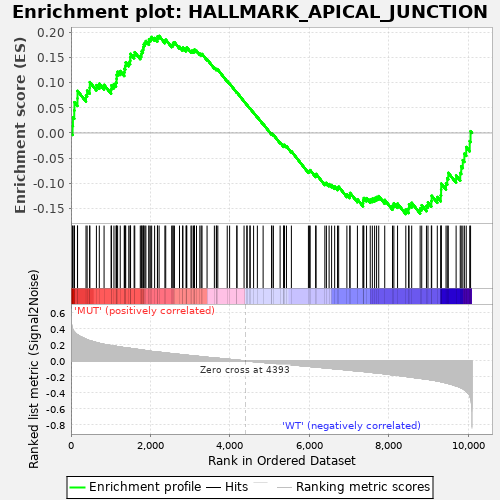
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_APICAL_JUNCTION |
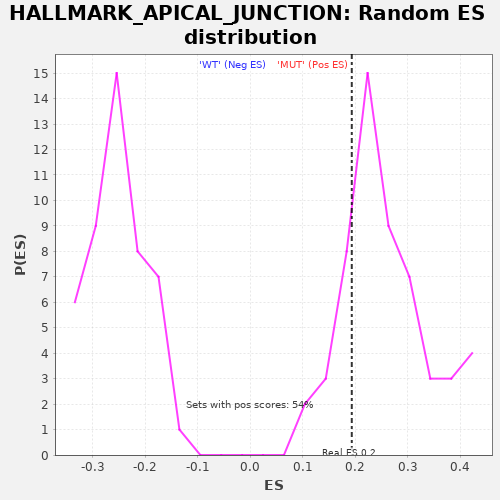
| Enrichment Score (ES) | 0.19308642 |
| Normalized Enrichment Score (NES) | 0.74810517 |
| Nominal p-value | 0.8333333 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COL17A1 | na | 40 | 0.398 | 0.0144 | Yes | ||
| 2 | PCDH1 | na | 49 | 0.386 | 0.0314 | Yes | ||
| 3 | ITGB4 | na | 82 | 0.363 | 0.0450 | Yes | ||
| 4 | SLIT2 | na | 87 | 0.360 | 0.0613 | Yes | ||
| 5 | SYK | na | 163 | 0.325 | 0.0688 | Yes | ||
| 6 | VASP | na | 165 | 0.325 | 0.0837 | Yes | ||
| 7 | RSU1 | na | 378 | 0.271 | 0.0750 | Yes | ||
| 8 | CTNND1 | na | 406 | 0.264 | 0.0845 | Yes | ||
| 9 | RRAS | na | 470 | 0.253 | 0.0898 | Yes | ||
| 10 | CALB2 | na | 474 | 0.252 | 0.1012 | Yes | ||
| 11 | ACTN1 | na | 640 | 0.228 | 0.0952 | Yes | ||
| 12 | ACTG2 | na | 713 | 0.220 | 0.0981 | Yes | ||
| 13 | ITGA2 | na | 833 | 0.206 | 0.0957 | Yes | ||
| 14 | KCNH2 | na | 1010 | 0.192 | 0.0869 | Yes | ||
| 15 | CRAT | na | 1019 | 0.191 | 0.0949 | Yes | ||
| 16 | ACTN4 | na | 1082 | 0.186 | 0.0973 | Yes | ||
| 17 | MAPK13 | na | 1134 | 0.182 | 0.1006 | Yes | ||
| 18 | MYL9 | na | 1148 | 0.180 | 0.1076 | Yes | ||
| 19 | DLG1 | na | 1151 | 0.180 | 0.1158 | Yes | ||
| 20 | DSC1 | na | 1174 | 0.178 | 0.1218 | Yes | ||
| 21 | ICAM4 | na | 1240 | 0.173 | 0.1233 | Yes | ||
| 22 | CLDN4 | na | 1341 | 0.166 | 0.1209 | Yes | ||
| 23 | TGFBI | na | 1348 | 0.165 | 0.1279 | Yes | ||
| 24 | JUP | na | 1368 | 0.164 | 0.1336 | Yes | ||
| 25 | CDH3 | na | 1378 | 0.163 | 0.1403 | Yes | ||
| 26 | AMH | na | 1455 | 0.159 | 0.1400 | Yes | ||
| 27 | MMP2 | na | 1489 | 0.156 | 0.1439 | Yes | ||
| 28 | MAPK14 | na | 1494 | 0.156 | 0.1507 | Yes | ||
| 29 | EGFR | na | 1500 | 0.156 | 0.1574 | Yes | ||
| 30 | TUBG1 | na | 1590 | 0.149 | 0.1554 | Yes | ||
| 31 | GNAI2 | na | 1606 | 0.148 | 0.1607 | Yes | ||
| 32 | VCL | na | 1745 | 0.138 | 0.1533 | Yes | ||
| 33 | TAOK2 | na | 1769 | 0.137 | 0.1573 | Yes | ||
| 34 | CLDN9 | na | 1781 | 0.136 | 0.1625 | Yes | ||
| 35 | CDH11 | na | 1811 | 0.135 | 0.1658 | Yes | ||
| 36 | MYH9 | na | 1820 | 0.134 | 0.1712 | Yes | ||
| 37 | TRAF1 | na | 1833 | 0.133 | 0.1761 | Yes | ||
| 38 | LAMC2 | na | 1861 | 0.130 | 0.1795 | Yes | ||
| 39 | NFASC | na | 1886 | 0.129 | 0.1830 | Yes | ||
| 40 | ZYX | na | 1962 | 0.122 | 0.1811 | Yes | ||
| 41 | PTK2 | na | 1965 | 0.122 | 0.1866 | Yes | ||
| 42 | TRO | na | 2008 | 0.119 | 0.1879 | Yes | ||
| 43 | CLDN7 | na | 2032 | 0.118 | 0.1910 | Yes | ||
| 44 | ACTB | na | 2106 | 0.114 | 0.1889 | Yes | ||
| 45 | GRB7 | na | 2177 | 0.110 | 0.1870 | Yes | ||
| 46 | MVD | na | 2179 | 0.110 | 0.1920 | Yes | ||
| 47 | LAMB3 | na | 2219 | 0.108 | 0.1931 | Yes | ||
| 48 | ICAM5 | na | 2365 | 0.100 | 0.1831 | No | ||
| 49 | MAP4K2 | na | 2386 | 0.098 | 0.1857 | No | ||
| 50 | HRAS | na | 2536 | 0.090 | 0.1749 | No | ||
| 51 | RASA1 | na | 2566 | 0.089 | 0.1761 | No | ||
| 52 | THBS3 | na | 2574 | 0.088 | 0.1795 | No | ||
| 53 | IRS1 | na | 2605 | 0.087 | 0.1805 | No | ||
| 54 | TIAL1 | na | 2733 | 0.080 | 0.1715 | No | ||
| 55 | CD86 | na | 2811 | 0.076 | 0.1672 | No | ||
| 56 | AKT2 | na | 2818 | 0.075 | 0.1701 | No | ||
| 57 | YWHAH | na | 2897 | 0.071 | 0.1655 | No | ||
| 58 | ITGB1 | na | 2909 | 0.071 | 0.1677 | No | ||
| 59 | CDH4 | na | 2917 | 0.070 | 0.1703 | No | ||
| 60 | CTNNA1 | na | 3022 | 0.065 | 0.1628 | No | ||
| 61 | ITGA3 | na | 3044 | 0.064 | 0.1637 | No | ||
| 62 | ADAM9 | na | 3084 | 0.062 | 0.1627 | No | ||
| 63 | NF1 | na | 3106 | 0.061 | 0.1634 | No | ||
| 64 | COL16A1 | na | 3109 | 0.061 | 0.1660 | No | ||
| 65 | ADRA1B | na | 3161 | 0.058 | 0.1635 | No | ||
| 66 | TNFRSF11B | na | 3245 | 0.054 | 0.1577 | No | ||
| 67 | MDK | na | 3278 | 0.052 | 0.1569 | No | ||
| 68 | CAP1 | na | 3300 | 0.051 | 0.1571 | No | ||
| 69 | B4GALT1 | na | 3427 | 0.044 | 0.1465 | No | ||
| 70 | NF2 | na | 3611 | 0.036 | 0.1298 | No | ||
| 71 | ADAM15 | na | 3660 | 0.034 | 0.1266 | No | ||
| 72 | ICAM1 | na | 3668 | 0.034 | 0.1274 | No | ||
| 73 | SHC1 | na | 3698 | 0.032 | 0.1260 | No | ||
| 74 | EPB41L2 | na | 3935 | 0.021 | 0.1032 | No | ||
| 75 | PKD1 | na | 3995 | 0.018 | 0.0982 | No | ||
| 76 | SDC3 | na | 4177 | 0.009 | 0.0804 | No | ||
| 77 | ARPC2 | na | 4179 | 0.009 | 0.0807 | No | ||
| 78 | FSCN1 | na | 4359 | 0.002 | 0.0628 | No | ||
| 79 | ICAM2 | na | 4428 | -0.002 | 0.0561 | No | ||
| 80 | CDH8 | na | 4437 | -0.002 | 0.0554 | No | ||
| 81 | FBN1 | na | 4506 | -0.005 | 0.0488 | No | ||
| 82 | INPPL1 | na | 4513 | -0.006 | 0.0484 | No | ||
| 83 | PIK3CB | na | 4599 | -0.010 | 0.0404 | No | ||
| 84 | TSC1 | na | 4694 | -0.014 | 0.0315 | No | ||
| 85 | SLC30A3 | na | 4839 | -0.020 | 0.0180 | No | ||
| 86 | GNAI1 | na | 5051 | -0.028 | -0.0019 | No | ||
| 87 | CLDN8 | na | 5060 | -0.029 | -0.0013 | No | ||
| 88 | FLNC | na | 5098 | -0.030 | -0.0037 | No | ||
| 89 | CD99 | na | 5267 | -0.038 | -0.0188 | No | ||
| 90 | JAM3 | na | 5357 | -0.042 | -0.0258 | No | ||
| 91 | PBX2 | na | 5358 | -0.042 | -0.0238 | No | ||
| 92 | SYMPK | na | 5378 | -0.043 | -0.0237 | No | ||
| 93 | CDH1 | na | 5427 | -0.045 | -0.0264 | No | ||
| 94 | MSN | na | 5551 | -0.051 | -0.0364 | No | ||
| 95 | DHX16 | na | 5980 | -0.069 | -0.0763 | No | ||
| 96 | DSC3 | na | 6005 | -0.070 | -0.0754 | No | ||
| 97 | SGCE | na | 6024 | -0.071 | -0.0739 | No | ||
| 98 | PFN1 | na | 6162 | -0.077 | -0.0842 | No | ||
| 99 | CDK8 | na | 6172 | -0.078 | -0.0815 | No | ||
| 100 | LAMA3 | na | 6394 | -0.088 | -0.0996 | No | ||
| 101 | ARHGEF6 | na | 6433 | -0.089 | -0.0993 | No | ||
| 102 | NRXN2 | na | 6501 | -0.093 | -0.1017 | No | ||
| 103 | CDH6 | na | 6564 | -0.096 | -0.1035 | No | ||
| 104 | RAC2 | na | 6641 | -0.100 | -0.1065 | No | ||
| 105 | CDSN | na | 6713 | -0.103 | -0.1089 | No | ||
| 106 | PLCG1 | na | 6742 | -0.105 | -0.1069 | No | ||
| 107 | CX3CL1 | na | 6949 | -0.114 | -0.1223 | No | ||
| 108 | IKBKG | na | 7022 | -0.118 | -0.1241 | No | ||
| 109 | BMP1 | na | 7029 | -0.118 | -0.1192 | No | ||
| 110 | PTPRC | na | 7214 | -0.126 | -0.1319 | No | ||
| 111 | AKT3 | na | 7356 | -0.133 | -0.1399 | No | ||
| 112 | PTEN | na | 7359 | -0.133 | -0.1339 | No | ||
| 113 | CDH15 | na | 7376 | -0.134 | -0.1293 | No | ||
| 114 | MPZL1 | na | 7444 | -0.138 | -0.1297 | No | ||
| 115 | VCAM1 | na | 7537 | -0.143 | -0.1323 | No | ||
| 116 | PIK3R3 | na | 7590 | -0.146 | -0.1307 | No | ||
| 117 | WASL | na | 7647 | -0.149 | -0.1295 | No | ||
| 118 | TJP1 | na | 7697 | -0.151 | -0.1274 | No | ||
| 119 | GTF2F1 | na | 7751 | -0.154 | -0.1256 | No | ||
| 120 | VWF | na | 7902 | -0.163 | -0.1331 | No | ||
| 121 | MMP9 | na | 8098 | -0.175 | -0.1446 | No | ||
| 122 | CLDN5 | na | 8132 | -0.177 | -0.1398 | No | ||
| 123 | DMP1 | na | 8225 | -0.183 | -0.1406 | No | ||
| 124 | COL9A1 | na | 8433 | -0.195 | -0.1523 | No | ||
| 125 | VAV2 | na | 8504 | -0.200 | -0.1501 | No | ||
| 126 | GAMT | na | 8514 | -0.200 | -0.1418 | No | ||
| 127 | MYH10 | na | 8581 | -0.205 | -0.1389 | No | ||
| 128 | INSIG1 | na | 8794 | -0.220 | -0.1500 | No | ||
| 129 | AMIGO2 | na | 8834 | -0.222 | -0.1437 | No | ||
| 130 | BAIAP2 | na | 8953 | -0.230 | -0.1449 | No | ||
| 131 | ADAM23 | na | 8991 | -0.232 | -0.1379 | No | ||
| 132 | ATP1A3 | na | 9076 | -0.240 | -0.1352 | No | ||
| 133 | ITGA9 | na | 9085 | -0.240 | -0.1249 | No | ||
| 134 | MADCAM1 | na | 9226 | -0.251 | -0.1273 | No | ||
| 135 | NRTN | na | 9314 | -0.261 | -0.1240 | No | ||
| 136 | ACTN2 | na | 9322 | -0.262 | -0.1126 | No | ||
| 137 | PECAM1 | na | 9325 | -0.262 | -0.1006 | No | ||
| 138 | CD34 | na | 9446 | -0.276 | -0.0999 | No | ||
| 139 | ACTA1 | na | 9477 | -0.281 | -0.0899 | No | ||
| 140 | ACTG1 | na | 9503 | -0.284 | -0.0793 | No | ||
| 141 | ALOX15B | na | 9700 | -0.315 | -0.0844 | No | ||
| 142 | CNTN1 | na | 9804 | -0.333 | -0.0794 | No | ||
| 143 | ACTN3 | na | 9829 | -0.338 | -0.0661 | No | ||
| 144 | CLDN14 | na | 9869 | -0.348 | -0.0539 | No | ||
| 145 | SRC | na | 9908 | -0.362 | -0.0410 | No | ||
| 146 | MAPK11 | na | 9955 | -0.384 | -0.0279 | No | ||
| 147 | ITGA10 | na | 10043 | -0.441 | -0.0162 | No | ||
| 148 | CLDN18 | na | 10064 | -0.470 | 0.0035 | No |