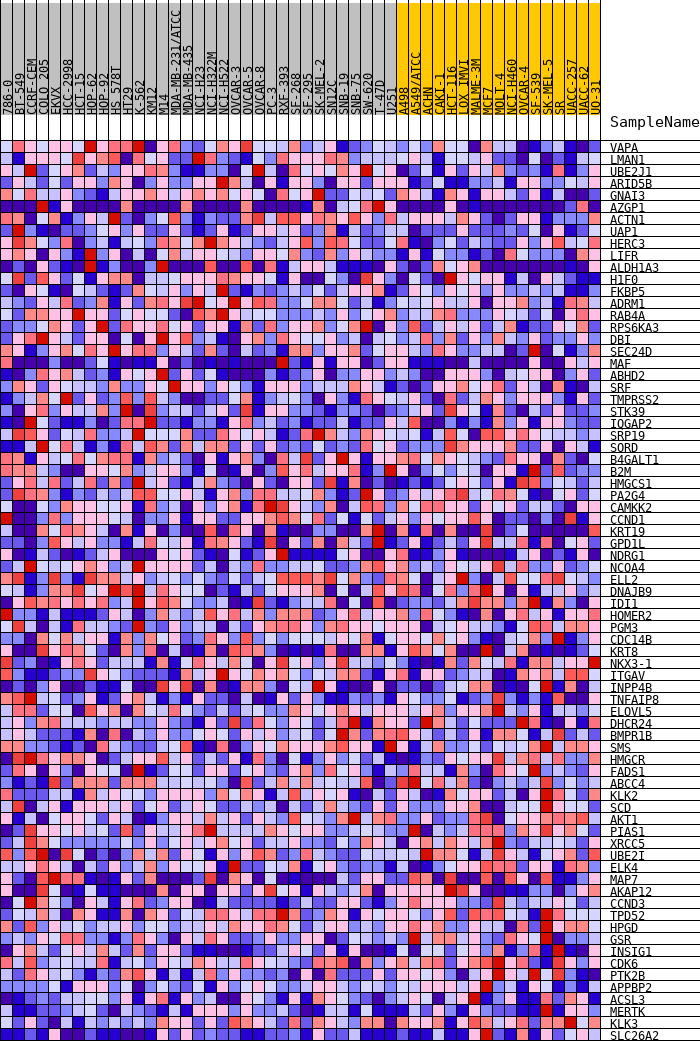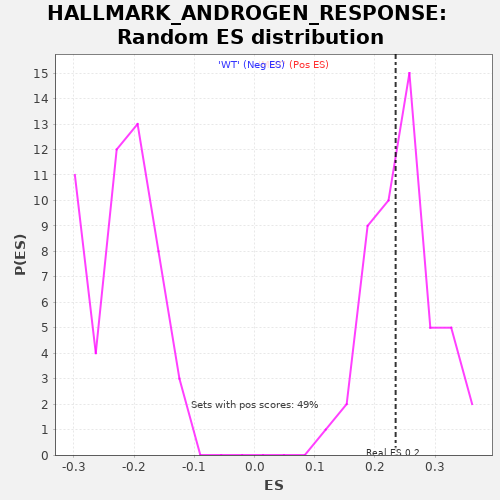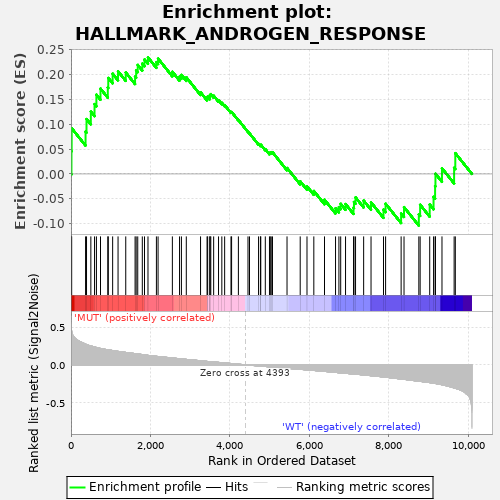
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_ANDROGEN_RESPONSE |
| Enrichment Score (ES) | 0.2345215 |
| Normalized Enrichment Score (NES) | 0.9532872 |
| Nominal p-value | 0.5714286 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | VAPA | na | 12 | 0.455 | 0.0458 | Yes | ||
| 2 | LMAN1 | na | 17 | 0.451 | 0.0919 | Yes | ||
| 3 | UBE2J1 | na | 367 | 0.272 | 0.0852 | Yes | ||
| 4 | ARID5B | na | 391 | 0.267 | 0.1105 | Yes | ||
| 5 | GNAI3 | na | 500 | 0.249 | 0.1255 | Yes | ||
| 6 | AZGP1 | na | 593 | 0.235 | 0.1406 | Yes | ||
| 7 | ACTN1 | na | 640 | 0.228 | 0.1595 | Yes | ||
| 8 | UAP1 | na | 743 | 0.216 | 0.1717 | Yes | ||
| 9 | HERC3 | na | 927 | 0.198 | 0.1739 | Yes | ||
| 10 | LIFR | na | 939 | 0.197 | 0.1932 | Yes | ||
| 11 | ALDH1A3 | na | 1049 | 0.188 | 0.2018 | Yes | ||
| 12 | H1F0 | na | 1185 | 0.177 | 0.2066 | Yes | ||
| 13 | FKBP5 | na | 1379 | 0.163 | 0.2042 | Yes | ||
| 14 | ADRM1 | na | 1614 | 0.148 | 0.1961 | Yes | ||
| 15 | RAB4A | na | 1641 | 0.146 | 0.2086 | Yes | ||
| 16 | RPS6KA3 | na | 1681 | 0.143 | 0.2195 | Yes | ||
| 17 | DBI | na | 1796 | 0.136 | 0.2221 | Yes | ||
| 18 | SEC24D | na | 1851 | 0.132 | 0.2303 | Yes | ||
| 19 | MAF | na | 1939 | 0.125 | 0.2345 | Yes | ||
| 20 | ABHD2 | na | 2151 | 0.111 | 0.2250 | No | ||
| 21 | SRF | na | 2193 | 0.109 | 0.2322 | No | ||
| 22 | TMPRSS2 | na | 2553 | 0.090 | 0.2056 | No | ||
| 23 | STK39 | na | 2734 | 0.080 | 0.1960 | No | ||
| 24 | IQGAP2 | na | 2781 | 0.077 | 0.1993 | No | ||
| 25 | SRP19 | na | 2904 | 0.071 | 0.1945 | No | ||
| 26 | SORD | na | 3263 | 0.053 | 0.1643 | No | ||
| 27 | B4GALT1 | na | 3427 | 0.044 | 0.1526 | No | ||
| 28 | B2M | na | 3442 | 0.043 | 0.1557 | No | ||
| 29 | HMGCS1 | na | 3498 | 0.041 | 0.1544 | No | ||
| 30 | PA2G4 | na | 3499 | 0.041 | 0.1586 | No | ||
| 31 | CAMKK2 | na | 3521 | 0.040 | 0.1607 | No | ||
| 32 | CCND1 | na | 3593 | 0.037 | 0.1574 | No | ||
| 33 | KRT19 | na | 3716 | 0.031 | 0.1485 | No | ||
| 34 | GPD1L | na | 3799 | 0.027 | 0.1431 | No | ||
| 35 | NDRG1 | na | 3870 | 0.024 | 0.1387 | No | ||
| 36 | NCOA4 | na | 4030 | 0.017 | 0.1245 | No | ||
| 37 | ELL2 | na | 4045 | 0.015 | 0.1247 | No | ||
| 38 | DNAJB9 | na | 4216 | 0.008 | 0.1086 | No | ||
| 39 | IDI1 | na | 4458 | -0.003 | 0.0848 | No | ||
| 40 | HOMER2 | na | 4496 | -0.005 | 0.0816 | No | ||
| 41 | PGM3 | na | 4724 | -0.015 | 0.0606 | No | ||
| 42 | CDC14B | na | 4771 | -0.018 | 0.0578 | No | ||
| 43 | KRT8 | na | 4780 | -0.018 | 0.0589 | No | ||
| 44 | NKX3-1 | na | 4894 | -0.022 | 0.0499 | No | ||
| 45 | ITGAV | na | 4999 | -0.026 | 0.0423 | No | ||
| 46 | INPP4B | na | 5019 | -0.027 | 0.0432 | No | ||
| 47 | TNFAIP8 | na | 5043 | -0.028 | 0.0438 | No | ||
| 48 | ELOVL5 | na | 5075 | -0.029 | 0.0437 | No | ||
| 49 | DHCR24 | na | 5441 | -0.046 | 0.0120 | No | ||
| 50 | BMPR1B | na | 5773 | -0.060 | -0.0148 | No | ||
| 51 | SMS | na | 5944 | -0.067 | -0.0249 | No | ||
| 52 | HMGCR | na | 6116 | -0.075 | -0.0342 | No | ||
| 53 | FADS1 | na | 6385 | -0.088 | -0.0519 | No | ||
| 54 | ABCC4 | na | 6663 | -0.100 | -0.0691 | No | ||
| 55 | KLK2 | na | 6748 | -0.105 | -0.0667 | No | ||
| 56 | SCD | na | 6793 | -0.107 | -0.0601 | No | ||
| 57 | AKT1 | na | 6915 | -0.112 | -0.0605 | No | ||
| 58 | PIAS1 | na | 7116 | -0.122 | -0.0679 | No | ||
| 59 | XRCC5 | na | 7128 | -0.123 | -0.0563 | No | ||
| 60 | UBE2I | na | 7167 | -0.124 | -0.0472 | No | ||
| 61 | ELK4 | na | 7371 | -0.134 | -0.0536 | No | ||
| 62 | MAP7 | na | 7557 | -0.145 | -0.0571 | No | ||
| 63 | AKAP12 | na | 7871 | -0.161 | -0.0717 | No | ||
| 64 | CCND3 | na | 7924 | -0.164 | -0.0599 | No | ||
| 65 | TPD52 | na | 8315 | -0.188 | -0.0795 | No | ||
| 66 | HPGD | na | 8388 | -0.192 | -0.0668 | No | ||
| 67 | GSR | na | 8761 | -0.217 | -0.0815 | No | ||
| 68 | INSIG1 | na | 8794 | -0.220 | -0.0620 | No | ||
| 69 | CDK6 | na | 9035 | -0.236 | -0.0616 | No | ||
| 70 | PTK2B | na | 9132 | -0.243 | -0.0460 | No | ||
| 71 | APPBP2 | na | 9171 | -0.247 | -0.0244 | No | ||
| 72 | ACSL3 | na | 9180 | -0.248 | 0.0004 | No | ||
| 73 | MERTK | na | 9344 | -0.264 | 0.0114 | No | ||
| 74 | KLK3 | na | 9650 | -0.306 | 0.0126 | No | ||
| 75 | SLC26A2 | na | 9680 | -0.311 | 0.0418 | No |