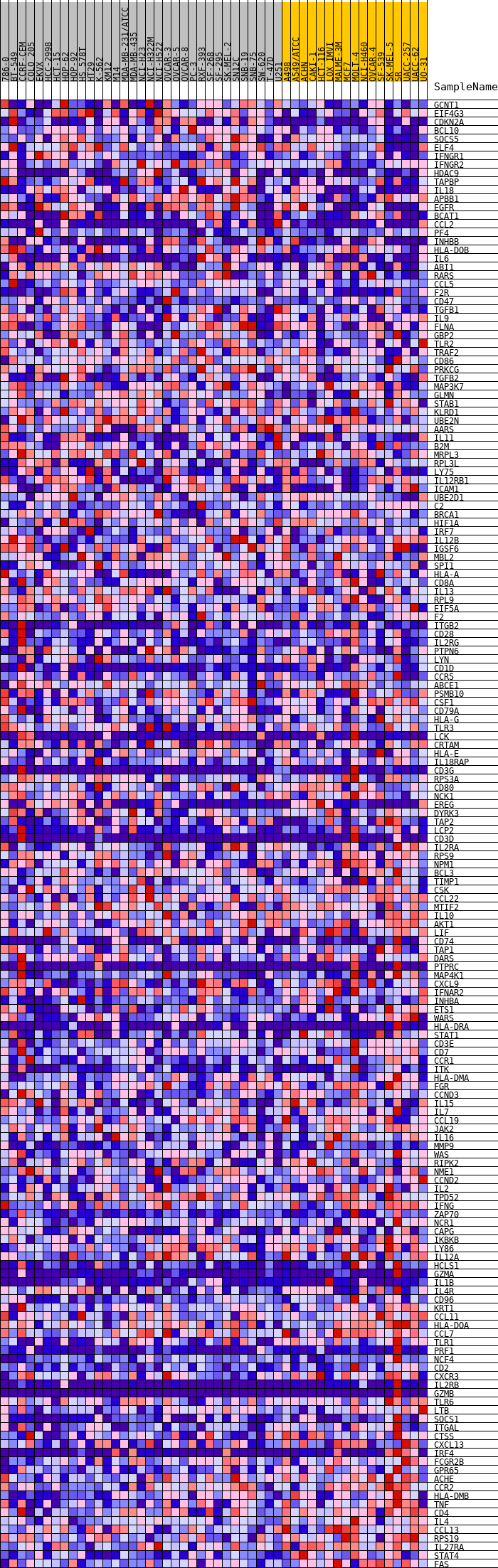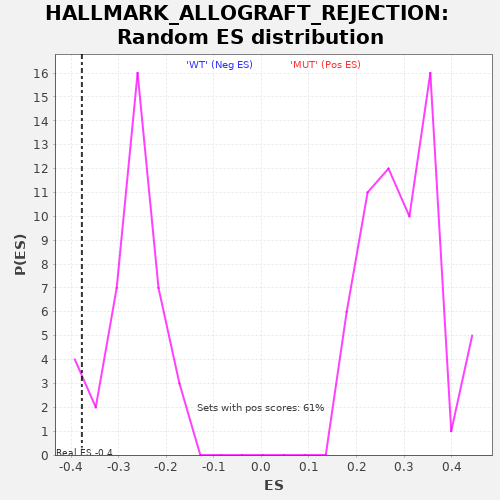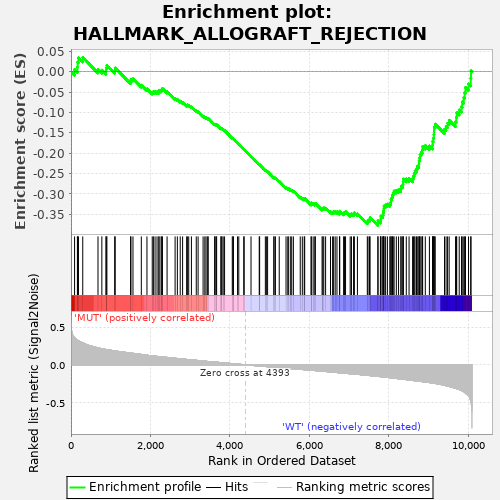
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | WT |
| GeneSet | HALLMARK_ALLOGRAFT_REJECTION |
| Enrichment Score (ES) | -0.37779245 |
| Normalized Enrichment Score (NES) | -1.381798 |
| Nominal p-value | 0.07692308 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.72 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GCNT1 | na | 90 | 0.360 | 0.0052 | No | ||
| 2 | EIF4G3 | na | 161 | 0.326 | 0.0111 | No | ||
| 3 | CDKN2A | na | 168 | 0.324 | 0.0234 | No | ||
| 4 | BCL10 | na | 188 | 0.320 | 0.0342 | No | ||
| 5 | SOCS5 | na | 296 | 0.290 | 0.0349 | No | ||
| 6 | ELF4 | na | 680 | 0.224 | 0.0052 | No | ||
| 7 | IFNGR1 | na | 777 | 0.212 | 0.0039 | No | ||
| 8 | IFNGR2 | na | 883 | 0.202 | 0.0014 | No | ||
| 9 | HDAC9 | na | 885 | 0.202 | 0.0093 | No | ||
| 10 | TAPBP | na | 904 | 0.200 | 0.0154 | No | ||
| 11 | IL18 | na | 1100 | 0.185 | 0.0031 | No | ||
| 12 | APBB1 | na | 1113 | 0.184 | 0.0092 | No | ||
| 13 | EGFR | na | 1500 | 0.156 | -0.0235 | No | ||
| 14 | BCAT1 | na | 1510 | 0.155 | -0.0182 | No | ||
| 15 | CCL2 | na | 1561 | 0.151 | -0.0173 | No | ||
| 16 | PF4 | na | 1771 | 0.137 | -0.0329 | No | ||
| 17 | INHBB | na | 1913 | 0.127 | -0.0421 | No | ||
| 18 | HLA-DOB | na | 2046 | 0.117 | -0.0507 | No | ||
| 19 | IL6 | na | 2075 | 0.115 | -0.0489 | No | ||
| 20 | ABI1 | na | 2117 | 0.113 | -0.0486 | No | ||
| 21 | RARS | na | 2167 | 0.111 | -0.0491 | No | ||
| 22 | CCL5 | na | 2213 | 0.109 | -0.0493 | No | ||
| 23 | F2R | na | 2218 | 0.108 | -0.0455 | No | ||
| 24 | CD47 | na | 2268 | 0.106 | -0.0462 | No | ||
| 25 | TGFB1 | na | 2285 | 0.105 | -0.0437 | No | ||
| 26 | IL9 | na | 2303 | 0.104 | -0.0412 | No | ||
| 27 | FLNA | na | 2420 | 0.097 | -0.0491 | No | ||
| 28 | GBP2 | na | 2622 | 0.086 | -0.0659 | No | ||
| 29 | TLR2 | na | 2679 | 0.083 | -0.0682 | No | ||
| 30 | TRAF2 | na | 2751 | 0.079 | -0.0722 | No | ||
| 31 | CD86 | na | 2811 | 0.076 | -0.0752 | No | ||
| 32 | PRKCG | na | 2913 | 0.071 | -0.0825 | No | ||
| 33 | TGFB2 | na | 2931 | 0.069 | -0.0815 | No | ||
| 34 | MAP3K7 | na | 2965 | 0.068 | -0.0821 | No | ||
| 35 | GLMN | na | 3033 | 0.065 | -0.0863 | No | ||
| 36 | STAB1 | na | 3155 | 0.058 | -0.0962 | No | ||
| 37 | KLRD1 | na | 3201 | 0.056 | -0.0985 | No | ||
| 38 | UBE2N | na | 3327 | 0.050 | -0.1091 | No | ||
| 39 | AARS | na | 3371 | 0.047 | -0.1116 | No | ||
| 40 | IL11 | na | 3402 | 0.046 | -0.1128 | No | ||
| 41 | B2M | na | 3442 | 0.043 | -0.1150 | No | ||
| 42 | MRPL3 | na | 3456 | 0.043 | -0.1146 | No | ||
| 43 | RPL3L | na | 3618 | 0.036 | -0.1294 | No | ||
| 44 | LY75 | na | 3628 | 0.035 | -0.1289 | No | ||
| 45 | IL12RB1 | na | 3655 | 0.034 | -0.1301 | No | ||
| 46 | ICAM1 | na | 3668 | 0.034 | -0.1300 | No | ||
| 47 | UBE2D1 | na | 3775 | 0.029 | -0.1396 | No | ||
| 48 | C2 | na | 3783 | 0.028 | -0.1391 | No | ||
| 49 | BRCA1 | na | 3802 | 0.027 | -0.1399 | No | ||
| 50 | HIF1A | na | 3833 | 0.026 | -0.1419 | No | ||
| 51 | IRF7 | na | 3868 | 0.024 | -0.1443 | No | ||
| 52 | IL12B | na | 4062 | 0.015 | -0.1632 | No | ||
| 53 | IGSF6 | na | 4081 | 0.013 | -0.1645 | No | ||
| 54 | MBL2 | na | 4091 | 0.013 | -0.1649 | No | ||
| 55 | SPI1 | na | 4195 | 0.009 | -0.1749 | No | ||
| 56 | HLA-A | na | 4226 | 0.007 | -0.1776 | No | ||
| 57 | CD8A | na | 4352 | 0.002 | -0.1901 | No | ||
| 58 | IL13 | na | 4361 | 0.001 | -0.1909 | No | ||
| 59 | RPL9 | na | 4534 | -0.007 | -0.2080 | No | ||
| 60 | EIF5A | na | 4741 | -0.016 | -0.2281 | No | ||
| 61 | F2 | na | 4751 | -0.017 | -0.2283 | No | ||
| 62 | ITGB2 | na | 4895 | -0.022 | -0.2418 | No | ||
| 63 | CD28 | na | 4921 | -0.023 | -0.2434 | No | ||
| 64 | IL2RG | na | 4952 | -0.025 | -0.2454 | No | ||
| 65 | PTPN6 | na | 5108 | -0.031 | -0.2598 | No | ||
| 66 | LYN | na | 5115 | -0.031 | -0.2592 | No | ||
| 67 | CD1D | na | 5149 | -0.033 | -0.2612 | No | ||
| 68 | CCR5 | na | 5245 | -0.037 | -0.2693 | No | ||
| 69 | ABCE1 | na | 5416 | -0.045 | -0.2847 | No | ||
| 70 | PSMB10 | na | 5462 | -0.047 | -0.2873 | No | ||
| 71 | CSF1 | na | 5473 | -0.047 | -0.2864 | No | ||
| 72 | CD79A | na | 5524 | -0.050 | -0.2895 | No | ||
| 73 | HLA-G | na | 5559 | -0.051 | -0.2909 | No | ||
| 74 | TLR3 | na | 5601 | -0.053 | -0.2929 | No | ||
| 75 | LCK | na | 5774 | -0.060 | -0.3079 | No | ||
| 76 | CRTAM | na | 5826 | -0.062 | -0.3105 | No | ||
| 77 | HLA-E | na | 5874 | -0.064 | -0.3127 | No | ||
| 78 | IL18RAP | na | 5886 | -0.065 | -0.3112 | No | ||
| 79 | CD3G | na | 6049 | -0.072 | -0.3247 | No | ||
| 80 | RPS3A | na | 6051 | -0.072 | -0.3219 | No | ||
| 81 | CD80 | na | 6099 | -0.074 | -0.3237 | No | ||
| 82 | NCK1 | na | 6140 | -0.076 | -0.3247 | No | ||
| 83 | EREG | na | 6155 | -0.076 | -0.3231 | No | ||
| 84 | DYRK3 | na | 6324 | -0.085 | -0.3366 | No | ||
| 85 | TAP2 | na | 6344 | -0.085 | -0.3352 | No | ||
| 86 | LCP2 | na | 6367 | -0.086 | -0.3340 | No | ||
| 87 | CD3D | na | 6413 | -0.089 | -0.3350 | No | ||
| 88 | IL2RA | na | 6536 | -0.095 | -0.3435 | No | ||
| 89 | RPS9 | na | 6589 | -0.097 | -0.3449 | No | ||
| 90 | NPM1 | na | 6615 | -0.099 | -0.3435 | No | ||
| 91 | BCL3 | na | 6654 | -0.100 | -0.3433 | No | ||
| 92 | TIMP1 | na | 6699 | -0.102 | -0.3437 | No | ||
| 93 | CSK | na | 6763 | -0.106 | -0.3458 | No | ||
| 94 | CCL22 | na | 6769 | -0.106 | -0.3421 | No | ||
| 95 | MTIF2 | na | 6866 | -0.110 | -0.3475 | No | ||
| 96 | IL10 | na | 6888 | -0.111 | -0.3452 | No | ||
| 97 | AKT1 | na | 6915 | -0.112 | -0.3433 | No | ||
| 98 | LIF | na | 7032 | -0.118 | -0.3503 | No | ||
| 99 | CD74 | na | 7064 | -0.120 | -0.3487 | No | ||
| 100 | TAP1 | na | 7117 | -0.122 | -0.3490 | No | ||
| 101 | DARS | na | 7138 | -0.123 | -0.3462 | No | ||
| 102 | PTPRC | na | 7214 | -0.126 | -0.3487 | No | ||
| 103 | MAP4K1 | na | 7463 | -0.139 | -0.3682 | No | ||
| 104 | CXCL9 | na | 7481 | -0.140 | -0.3643 | No | ||
| 105 | IFNAR2 | na | 7519 | -0.142 | -0.3624 | No | ||
| 106 | INHBA | na | 7533 | -0.143 | -0.3581 | No | ||
| 107 | ETS1 | na | 7730 | -0.153 | -0.3717 | Yes | ||
| 108 | WARS | na | 7732 | -0.153 | -0.3657 | Yes | ||
| 109 | HLA-DRA | na | 7793 | -0.157 | -0.3655 | Yes | ||
| 110 | STAT1 | na | 7803 | -0.157 | -0.3602 | Yes | ||
| 111 | CD3E | na | 7805 | -0.157 | -0.3541 | Yes | ||
| 112 | CD7 | na | 7852 | -0.160 | -0.3524 | Yes | ||
| 113 | CCR1 | na | 7864 | -0.161 | -0.3471 | Yes | ||
| 114 | ITK | na | 7873 | -0.161 | -0.3415 | Yes | ||
| 115 | HLA-DMA | na | 7880 | -0.162 | -0.3357 | Yes | ||
| 116 | FGR | na | 7883 | -0.162 | -0.3295 | Yes | ||
| 117 | CCND3 | na | 7924 | -0.164 | -0.3270 | Yes | ||
| 118 | IL15 | na | 7969 | -0.167 | -0.3248 | Yes | ||
| 119 | IL7 | na | 8030 | -0.171 | -0.3240 | Yes | ||
| 120 | CCL19 | na | 8057 | -0.173 | -0.3198 | Yes | ||
| 121 | JAK2 | na | 8060 | -0.173 | -0.3131 | Yes | ||
| 122 | IL16 | na | 8084 | -0.174 | -0.3085 | Yes | ||
| 123 | MMP9 | na | 8098 | -0.175 | -0.3029 | Yes | ||
| 124 | WAS | na | 8114 | -0.176 | -0.2974 | Yes | ||
| 125 | RIPK2 | na | 8138 | -0.177 | -0.2927 | Yes | ||
| 126 | NME1 | na | 8194 | -0.181 | -0.2911 | Yes | ||
| 127 | CCND2 | na | 8250 | -0.184 | -0.2893 | Yes | ||
| 128 | IL2 | na | 8308 | -0.187 | -0.2876 | Yes | ||
| 129 | TPD52 | na | 8315 | -0.188 | -0.2808 | Yes | ||
| 130 | IFNG | na | 8358 | -0.191 | -0.2774 | Yes | ||
| 131 | ZAP70 | na | 8364 | -0.191 | -0.2704 | Yes | ||
| 132 | NCR1 | na | 8370 | -0.191 | -0.2633 | Yes | ||
| 133 | CAPG | na | 8445 | -0.196 | -0.2629 | Yes | ||
| 134 | IKBKB | na | 8515 | -0.200 | -0.2619 | Yes | ||
| 135 | LY86 | na | 8604 | -0.206 | -0.2626 | Yes | ||
| 136 | IL12A | na | 8625 | -0.208 | -0.2563 | Yes | ||
| 137 | HCLS1 | na | 8644 | -0.210 | -0.2498 | Yes | ||
| 138 | GZMA | na | 8665 | -0.211 | -0.2434 | Yes | ||
| 139 | IL1B | na | 8703 | -0.214 | -0.2387 | Yes | ||
| 140 | IL4R | na | 8721 | -0.215 | -0.2318 | Yes | ||
| 141 | CD96 | na | 8763 | -0.218 | -0.2273 | Yes | ||
| 142 | KRT1 | na | 8766 | -0.218 | -0.2189 | Yes | ||
| 143 | CCL11 | na | 8781 | -0.219 | -0.2116 | Yes | ||
| 144 | HLA-DOA | na | 8790 | -0.219 | -0.2037 | Yes | ||
| 145 | CCL7 | na | 8819 | -0.221 | -0.1978 | Yes | ||
| 146 | TLR1 | na | 8850 | -0.223 | -0.1919 | Yes | ||
| 147 | PRF1 | na | 8857 | -0.224 | -0.1836 | Yes | ||
| 148 | NCF4 | na | 8924 | -0.228 | -0.1812 | Yes | ||
| 149 | CD2 | na | 9027 | -0.235 | -0.1822 | Yes | ||
| 150 | CXCR3 | na | 9107 | -0.242 | -0.1805 | Yes | ||
| 151 | IL2RB | na | 9109 | -0.242 | -0.1710 | Yes | ||
| 152 | GZMB | na | 9126 | -0.243 | -0.1630 | Yes | ||
| 153 | TLR6 | na | 9139 | -0.244 | -0.1545 | Yes | ||
| 154 | LTB | na | 9145 | -0.244 | -0.1453 | Yes | ||
| 155 | SOCS1 | na | 9147 | -0.244 | -0.1357 | Yes | ||
| 156 | ITGAL | na | 9174 | -0.247 | -0.1285 | Yes | ||
| 157 | CTSS | na | 9408 | -0.272 | -0.1412 | Yes | ||
| 158 | CXCL13 | na | 9447 | -0.276 | -0.1341 | Yes | ||
| 159 | IRF4 | na | 9485 | -0.282 | -0.1266 | Yes | ||
| 160 | FCGR2B | na | 9525 | -0.286 | -0.1191 | Yes | ||
| 161 | GPR65 | na | 9686 | -0.312 | -0.1229 | Yes | ||
| 162 | ACHE | na | 9706 | -0.315 | -0.1123 | Yes | ||
| 163 | CCR2 | na | 9715 | -0.316 | -0.1005 | Yes | ||
| 164 | HLA-DMB | na | 9778 | -0.327 | -0.0938 | Yes | ||
| 165 | TNF | na | 9835 | -0.340 | -0.0859 | Yes | ||
| 166 | CD4 | na | 9854 | -0.345 | -0.0741 | Yes | ||
| 167 | IL4 | na | 9893 | -0.354 | -0.0638 | Yes | ||
| 168 | CCL13 | na | 9915 | -0.364 | -0.0515 | Yes | ||
| 169 | RPS19 | na | 9936 | -0.375 | -0.0386 | Yes | ||
| 170 | IL27RA | na | 10013 | -0.409 | -0.0300 | Yes | ||
| 171 | STAT4 | na | 10068 | -0.480 | -0.0164 | Yes | ||
| 172 | FAS | na | 10075 | -0.489 | 0.0024 | Yes |