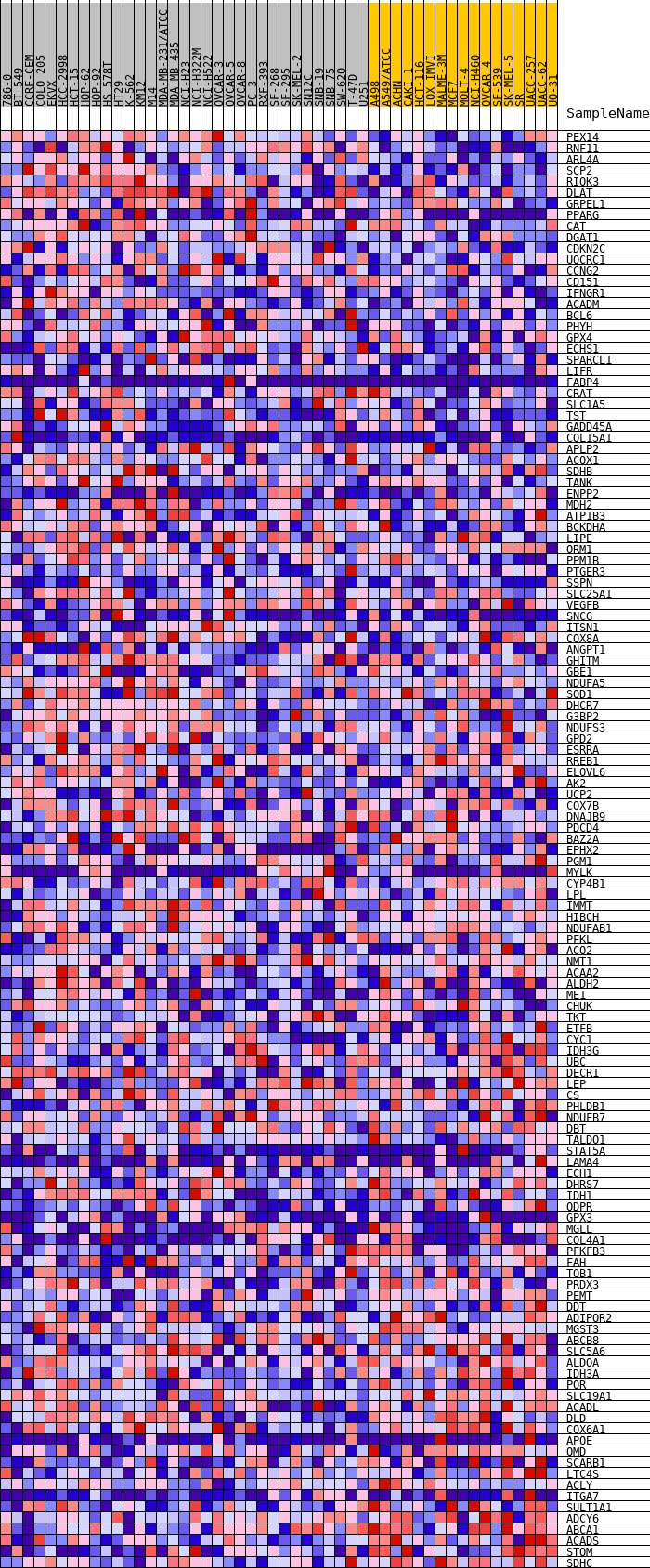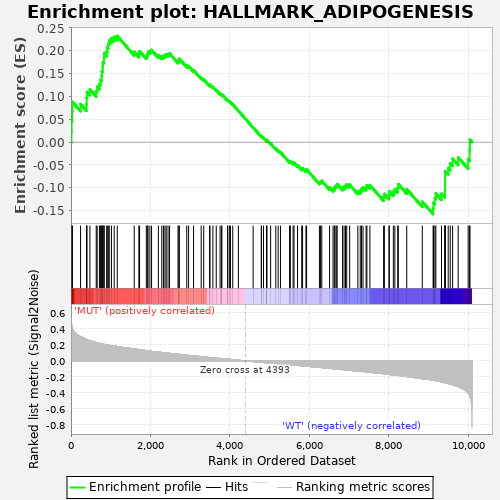
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | P53_collapsed_symbols.P53.cls#MUT_versus_WT |
| Phenotype | P53.cls#MUT_versus_WT |
| Upregulated in class | MUT |
| GeneSet | HALLMARK_ADIPOGENESIS |
| Enrichment Score (ES) | 0.2323492 |
| Normalized Enrichment Score (NES) | 0.87401533 |
| Nominal p-value | 0.5686275 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PEX14 | na | 11 | 0.457 | 0.0230 | Yes | ||
| 2 | RNF11 | na | 15 | 0.453 | 0.0465 | Yes | ||
| 3 | ARL4A | na | 29 | 0.414 | 0.0671 | Yes | ||
| 4 | SCP2 | na | 35 | 0.408 | 0.0880 | Yes | ||
| 5 | RIOK3 | na | 241 | 0.302 | 0.0834 | Yes | ||
| 6 | DLAT | na | 390 | 0.267 | 0.0827 | Yes | ||
| 7 | GRPEL1 | na | 393 | 0.266 | 0.0965 | Yes | ||
| 8 | PPARG | na | 402 | 0.264 | 0.1096 | Yes | ||
| 9 | CAT | na | 476 | 0.252 | 0.1156 | Yes | ||
| 10 | DGAT1 | na | 635 | 0.230 | 0.1118 | Yes | ||
| 11 | CDKN2C | na | 659 | 0.226 | 0.1214 | Yes | ||
| 12 | UQCRC1 | na | 716 | 0.219 | 0.1274 | Yes | ||
| 13 | CCNG2 | na | 745 | 0.216 | 0.1359 | Yes | ||
| 14 | CD151 | na | 774 | 0.212 | 0.1443 | Yes | ||
| 15 | IFNGR1 | na | 777 | 0.212 | 0.1553 | Yes | ||
| 16 | ACADM | na | 800 | 0.209 | 0.1641 | Yes | ||
| 17 | BCL6 | na | 804 | 0.209 | 0.1748 | Yes | ||
| 18 | PHYH | na | 831 | 0.207 | 0.1831 | Yes | ||
| 19 | GPX4 | na | 834 | 0.206 | 0.1938 | Yes | ||
| 20 | ECHS1 | na | 899 | 0.200 | 0.1979 | Yes | ||
| 21 | SPARCL1 | na | 912 | 0.200 | 0.2072 | Yes | ||
| 22 | LIFR | na | 939 | 0.197 | 0.2150 | Yes | ||
| 23 | FABP4 | na | 964 | 0.196 | 0.2229 | Yes | ||
| 24 | CRAT | na | 1019 | 0.191 | 0.2276 | Yes | ||
| 25 | SLC1A5 | na | 1088 | 0.186 | 0.2306 | Yes | ||
| 26 | TST | na | 1165 | 0.178 | 0.2323 | Yes | ||
| 27 | GADD45A | na | 1592 | 0.149 | 0.1975 | No | ||
| 28 | COL15A1 | na | 1709 | 0.141 | 0.1933 | No | ||
| 29 | APLP2 | na | 1727 | 0.140 | 0.1989 | No | ||
| 30 | ACOX1 | na | 1897 | 0.128 | 0.1887 | No | ||
| 31 | SDHB | na | 1922 | 0.126 | 0.1930 | No | ||
| 32 | TANK | na | 1941 | 0.124 | 0.1977 | No | ||
| 33 | ENPP2 | na | 1987 | 0.121 | 0.1996 | No | ||
| 34 | MDH2 | na | 2029 | 0.118 | 0.2017 | No | ||
| 35 | ATP1B3 | na | 2201 | 0.109 | 0.1903 | No | ||
| 36 | BCKDHA | na | 2287 | 0.105 | 0.1873 | No | ||
| 37 | LIPE | na | 2329 | 0.102 | 0.1885 | No | ||
| 38 | ORM1 | na | 2367 | 0.100 | 0.1901 | No | ||
| 39 | PPM1B | na | 2397 | 0.098 | 0.1924 | No | ||
| 40 | PTGER3 | na | 2444 | 0.096 | 0.1928 | No | ||
| 41 | SSPN | na | 2483 | 0.093 | 0.1939 | No | ||
| 42 | SLC25A1 | na | 2696 | 0.082 | 0.1769 | No | ||
| 43 | VEGFB | na | 2718 | 0.081 | 0.1791 | No | ||
| 44 | SNCG | na | 2727 | 0.080 | 0.1825 | No | ||
| 45 | ITSN1 | na | 2915 | 0.070 | 0.1675 | No | ||
| 46 | COX8A | na | 2962 | 0.068 | 0.1664 | No | ||
| 47 | ANGPT1 | na | 3087 | 0.062 | 0.1573 | No | ||
| 48 | GHITM | na | 3279 | 0.052 | 0.1409 | No | ||
| 49 | GBE1 | na | 3343 | 0.049 | 0.1371 | No | ||
| 50 | NDUFA5 | na | 3491 | 0.041 | 0.1245 | No | ||
| 51 | SOD1 | na | 3500 | 0.041 | 0.1259 | No | ||
| 52 | DHCR7 | na | 3574 | 0.038 | 0.1206 | No | ||
| 53 | G3BP2 | na | 3658 | 0.034 | 0.1140 | No | ||
| 54 | NDUFS3 | na | 3755 | 0.029 | 0.1060 | No | ||
| 55 | GPD2 | na | 3794 | 0.028 | 0.1036 | No | ||
| 56 | ESRRA | na | 3798 | 0.028 | 0.1048 | No | ||
| 57 | RREB1 | na | 3943 | 0.021 | 0.0914 | No | ||
| 58 | ELOVL6 | na | 3947 | 0.021 | 0.0922 | No | ||
| 59 | AK2 | na | 3989 | 0.019 | 0.0891 | No | ||
| 60 | UCP2 | na | 4016 | 0.017 | 0.0874 | No | ||
| 61 | COX7B | na | 4074 | 0.014 | 0.0824 | No | ||
| 62 | DNAJB9 | na | 4216 | 0.008 | 0.0687 | No | ||
| 63 | PDCD4 | na | 4587 | -0.009 | 0.0320 | No | ||
| 64 | BAZ2A | na | 4794 | -0.019 | 0.0124 | No | ||
| 65 | EPHX2 | na | 4796 | -0.019 | 0.0132 | No | ||
| 66 | PGM1 | na | 4849 | -0.021 | 0.0091 | No | ||
| 67 | MYLK | na | 4925 | -0.023 | 0.0028 | No | ||
| 68 | CYP4B1 | na | 4930 | -0.024 | 0.0037 | No | ||
| 69 | LPL | na | 4932 | -0.024 | 0.0048 | No | ||
| 70 | IMMT | na | 5028 | -0.028 | -0.0032 | No | ||
| 71 | HIBCH | na | 5158 | -0.033 | -0.0144 | No | ||
| 72 | NDUFAB1 | na | 5218 | -0.036 | -0.0184 | No | ||
| 73 | PFKL | na | 5276 | -0.039 | -0.0221 | No | ||
| 74 | ACO2 | na | 5505 | -0.049 | -0.0424 | No | ||
| 75 | NMT1 | na | 5525 | -0.050 | -0.0417 | No | ||
| 76 | ACAA2 | na | 5594 | -0.053 | -0.0457 | No | ||
| 77 | ALDH2 | na | 5617 | -0.054 | -0.0451 | No | ||
| 78 | ME1 | na | 5706 | -0.057 | -0.0509 | No | ||
| 79 | CHUK | na | 5811 | -0.062 | -0.0581 | No | ||
| 80 | TKT | na | 5833 | -0.063 | -0.0569 | No | ||
| 81 | ETFB | na | 5918 | -0.066 | -0.0618 | No | ||
| 82 | CYC1 | na | 5934 | -0.067 | -0.0598 | No | ||
| 83 | IDH3G | na | 6257 | -0.082 | -0.0878 | No | ||
| 84 | UBC | na | 6286 | -0.083 | -0.0862 | No | ||
| 85 | DECR1 | na | 6317 | -0.084 | -0.0848 | No | ||
| 86 | LEP | na | 6513 | -0.093 | -0.0994 | No | ||
| 87 | CS | na | 6600 | -0.098 | -0.1029 | No | ||
| 88 | PHLDB1 | na | 6632 | -0.099 | -0.1007 | No | ||
| 89 | NDUFB7 | na | 6650 | -0.100 | -0.0972 | No | ||
| 90 | DBT | na | 6681 | -0.101 | -0.0949 | No | ||
| 91 | TALDO1 | na | 6705 | -0.103 | -0.0918 | No | ||
| 92 | STAT5A | na | 6840 | -0.109 | -0.0995 | No | ||
| 93 | LAMA4 | na | 6878 | -0.110 | -0.0974 | No | ||
| 94 | ECH1 | na | 6914 | -0.112 | -0.0950 | No | ||
| 95 | DHRS7 | na | 6945 | -0.114 | -0.0920 | No | ||
| 96 | IDH1 | na | 7019 | -0.117 | -0.0931 | No | ||
| 97 | QDPR | na | 7225 | -0.127 | -0.1070 | No | ||
| 98 | GPX3 | na | 7285 | -0.130 | -0.1060 | No | ||
| 99 | MGLL | na | 7316 | -0.132 | -0.1021 | No | ||
| 100 | COL4A1 | na | 7355 | -0.133 | -0.0989 | No | ||
| 101 | PFKFB3 | na | 7433 | -0.137 | -0.0994 | No | ||
| 102 | FAH | na | 7453 | -0.138 | -0.0940 | No | ||
| 103 | TOB1 | na | 7528 | -0.143 | -0.0939 | No | ||
| 104 | PRDX3 | na | 7874 | -0.161 | -0.1200 | No | ||
| 105 | PEMT | na | 7893 | -0.162 | -0.1132 | No | ||
| 106 | DDT | na | 8010 | -0.169 | -0.1159 | No | ||
| 107 | ADIPOR2 | na | 8015 | -0.170 | -0.1074 | No | ||
| 108 | MGST3 | na | 8119 | -0.176 | -0.1084 | No | ||
| 109 | ABCB8 | na | 8157 | -0.178 | -0.1028 | No | ||
| 110 | SLC5A6 | na | 8230 | -0.183 | -0.1003 | No | ||
| 111 | ALDOA | na | 8243 | -0.184 | -0.0919 | No | ||
| 112 | IDH3A | na | 8456 | -0.197 | -0.1027 | No | ||
| 113 | POR | na | 8848 | -0.223 | -0.1302 | No | ||
| 114 | SLC19A1 | na | 9121 | -0.243 | -0.1447 | No | ||
| 115 | ACADL | na | 9130 | -0.243 | -0.1327 | No | ||
| 116 | DLD | na | 9165 | -0.246 | -0.1231 | No | ||
| 117 | COX6A1 | na | 9189 | -0.248 | -0.1123 | No | ||
| 118 | APOE | na | 9332 | -0.263 | -0.1127 | No | ||
| 119 | OMD | na | 9418 | -0.273 | -0.1068 | No | ||
| 120 | SCARB1 | na | 9421 | -0.274 | -0.0926 | No | ||
| 121 | LTC4S | na | 9422 | -0.274 | -0.0782 | No | ||
| 122 | ACLY | na | 9423 | -0.274 | -0.0638 | No | ||
| 123 | ITGA7 | na | 9501 | -0.284 | -0.0565 | No | ||
| 124 | SULT1A1 | na | 9552 | -0.290 | -0.0462 | No | ||
| 125 | ADCY6 | na | 9612 | -0.299 | -0.0364 | No | ||
| 126 | ABCA1 | na | 9752 | -0.322 | -0.0333 | No | ||
| 127 | ACADS | na | 10004 | -0.403 | -0.0372 | No | ||
| 128 | STOM | na | 10041 | -0.439 | -0.0177 | No | ||
| 129 | SDHC | na | 10047 | -0.444 | 0.0052 | No |